2RVC
 
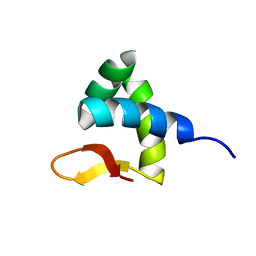 | | Solution structure of Zalpha domain of goldfish ZBP-containing protein kinase | | Descriptor: | Interferon-inducible and double-stranded-dependent eIF-2kinase | | Authors: | Lee, A, Park, C, Park, J, Kwon, M, Choi, Y, Kim, K, Choi, B, Lee, J. | | Deposit date: | 2015-07-08 | | Release date: | 2016-02-03 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the Z-DNA binding domain of PKR-like protein kinase from Carassius auratus and quantitative analyses of the intermediate complex during B-Z transition.
Nucleic Acids Res., 44, 2016
|
|
1SXL
 
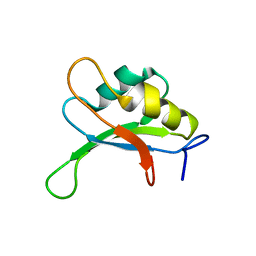 | |
1TYN
 
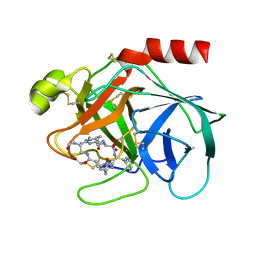 | | ATOMIC STRUCTURE OF THE TRYPSIN-CYCLOTHEONAMIDE A COMPLEX: LESSONS FOR THE DESIGN OF SERINE PROTEASE INHIBITORS | | Descriptor: | BETA-TRYPSIN, CYCLOTHEONAMIDE A | | Authors: | Lee, A.Y, Hagihara, M, Karmacharya, R, Albers, M.W, Schreiber, S.L, Clardy, J. | | Deposit date: | 1994-09-19 | | Release date: | 1995-01-26 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Atomic Structure of the Trypsin-Cyclotheonamide a Complex: Lessons for the Design of Serine Protease Inhibitors
J.Am.Chem.Soc., 115, 1993
|
|
1BJ2
 
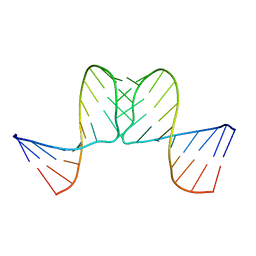 | |
1TPS
 
 | |
1LYW
 
 | | CATHEPSIN D AT PH 7.5 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CATHEPSIN D | | Authors: | Lee, A.Y, Gulnik, S.V, Erickson, J.W. | | Deposit date: | 1998-06-30 | | Release date: | 1999-07-22 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Conformational switching in an aspartic proteinase.
Nat.Struct.Biol., 5, 1998
|
|
2BJ2
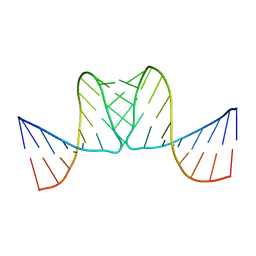 
 | |
8XTH
 
 | |
8IL2
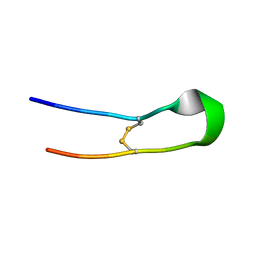 
 | |
8IL6
 
 | |
8IL1
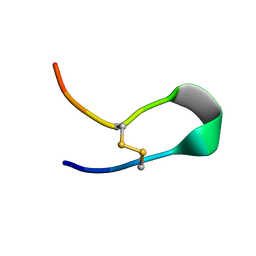 
 | |
7TL3
 
 | | Crystal Structure of Yeast p58C Multi-Tyrosine Mutant 5YF431 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, DNA primase large subunit, IRON/SULFUR CLUSTER | | Authors: | Blee, A.M, Salay, L.E, Chazin, W.J. | | Deposit date: | 2022-01-18 | | Release date: | 2022-06-29 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.066 Å) | | Cite: | Modification of the 4Fe-4S Cluster Charge Transport Pathway Alters RNA Synthesis by Yeast DNA Primase.
Biochemistry, 61, 2022
|
|
7TL4
 
 | | Crystal Structure of Yeast p58C Multi-Tyrosine Mutant 6YF | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, DNA primase large subunit, IRON/SULFUR CLUSTER | | Authors: | Blee, A.M, Salay, L.E, Chazin, W.J. | | Deposit date: | 2022-01-18 | | Release date: | 2022-06-29 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.805 Å) | | Cite: | Modification of the 4Fe-4S Cluster Charge Transport Pathway Alters RNA Synthesis by Yeast DNA Primase.
Biochemistry, 61, 2022
|
|
7TL2
 
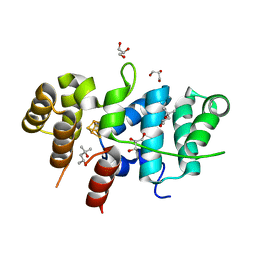 | | Crystal Structure of Yeast p58C Multi-Tyrosine Mutant 5YF412 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, DNA primase large subunit, GLYCEROL, ... | | Authors: | Blee, A.M, Salay, L.E, Chazin, W.J. | | Deposit date: | 2022-01-18 | | Release date: | 2022-06-29 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.529 Å) | | Cite: | Modification of the 4Fe-4S Cluster Charge Transport Pathway Alters RNA Synthesis by Yeast DNA Primase.
Biochemistry, 61, 2022
|
|
9ILX
 
 | | Thanatin VF16QK in complex with LPS | | Descriptor: | VAL-PRO-ILE-ILE-TYR-CYS-ASN-ARG-ARG-THR-DLY-LYS-CYS-LYS-ARG-PHE | | Authors: | Swaleeha, A, Bhattacharyya, S. | | Deposit date: | 2024-07-01 | | Release date: | 2025-05-07 | | Method: | SOLUTION NMR | | Cite: | Single Disulfide Bond in Host Defense Thanatin Analog Peptides: Antimicrobial Activity, Atomic-Resolution Structures and Target Interactions.
Int J Mol Sci, 26, 2024
|
|
9IKO
 
 | | VF16QK in free solution | | Descriptor: | VAL-PRO-ILE-ILE-TYR-CYS-ASN-ARG-ARG-THR-DLY-LYS-CYS-LYS-ARG-PHE | | Authors: | Swaleeha, A, Bhattacharyya, S. | | Deposit date: | 2024-06-28 | | Release date: | 2025-05-07 | | Method: | SOLUTION NMR | | Cite: | Single Disulfide Bond in Host Defense Thanatin Analog Peptides: Antimicrobial Activity, Atomic-Resolution Structures and Target Interactions.
Int J Mol Sci, 26, 2024
|
|
3PSA
 
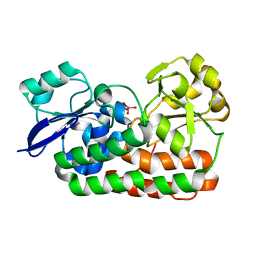 | | Classification of a Haemophilus influenzae ABC transporter HI1470/71 through its cognate molybdate periplasmic binding protein MolA (MolA bound to tungstate) | | Descriptor: | TUNGSTATE(VI)ION, protein HI_1472 | | Authors: | Tirado-Lee, L, Lee, A, Rees, D.C, Pinkett, H.W. | | Deposit date: | 2010-12-01 | | Release date: | 2011-11-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Classification of a Haemophilus influenzae ABC Transporter HI1470/71 through Its Cognate Molybdate Periplasmic Binding Protein, MolA.
Structure, 19, 2011
|
|
3PSH
 
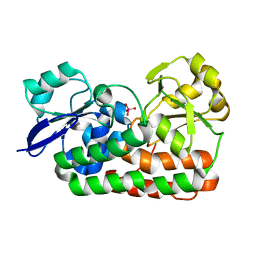 | | Classification of a Haemophilus influenzae ABC transporter HI1470/71 through its cognate molybdate periplasmic binding protein MolA (MolA bound to Molybdate) | | Descriptor: | MOLYBDATE ION, protein HI_1472 | | Authors: | Tirado-Lee, L, Lee, A, Rees, D.C, Pinkett, H.W. | | Deposit date: | 2010-12-01 | | Release date: | 2011-11-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Classification of a Haemophilus influenzae ABC Transporter HI1470/71 through Its Cognate Molybdate Periplasmic Binding Protein, MolA.
Structure, 19, 2011
|
|
4TXY
 
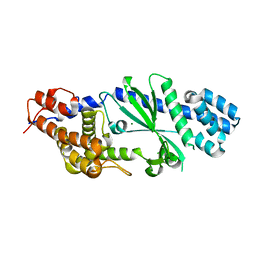 | | Crystal structure of Vibrio cholerae DncV cyclic AMP-GMP synthase, a prokaryotic cGAS homolog | | Descriptor: | Cyclic AMP-GMP synthase, MAGNESIUM ION | | Authors: | Kranzusch, P.J, Lee, A.S.Y, Wilson, S.C, Solovykh, M.S, Vance, R.E, Berger, J.M, Doudna, J.A. | | Deposit date: | 2014-07-07 | | Release date: | 2014-08-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.0001 Å) | | Cite: | Structure-Guided Reprogramming of Human cGAS Dinucleotide Linkage Specificity.
Cell, 158, 2014
|
|
4TXZ
 
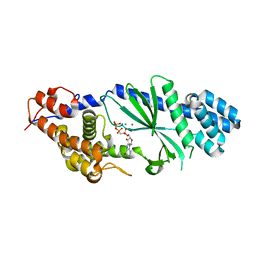 | | Crystal structure of Vibrio cholerae DncV cyclic AMP-GMP synthase in complex with nonhydrolyzable GTP | | Descriptor: | Cyclic AMP-GMP synthase, MAGNESIUM ION, PHOSPHOMETHYLPHOSPHONIC ACID GUANYLATE ESTER | | Authors: | Kranzusch, P.J, Lee, A.S.Y, Wilson, S.C, Solovykh, M.S, Vance, R.E, Berger, J.M, Doudna, J.A. | | Deposit date: | 2014-07-07 | | Release date: | 2014-08-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure-Guided Reprogramming of Human cGAS Dinucleotide Linkage Specificity.
Cell, 158, 2014
|
|
4TY0
 
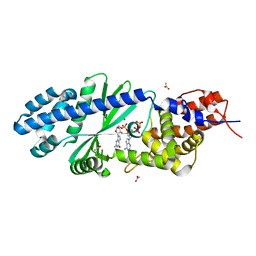 | | Crystal structure of Vibrio cholerae DncV cyclic AMP-GMP synthase in complex with linear intermediate 5' pppA(3',5')pG | | Descriptor: | ACETATE ION, Cyclic AMP-GMP synthase, MAGNESIUM ION, ... | | Authors: | Kranzusch, P.J, Lee, A.S.Y, Wilson, S.C, Solovykh, M.S, Vance, R.E, Berger, J.M, Doudna, J.A. | | Deposit date: | 2014-07-07 | | Release date: | 2014-08-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure-Guided Reprogramming of Human cGAS Dinucleotide Linkage Specificity.
Cell, 158, 2014
|
|
2A0L
 
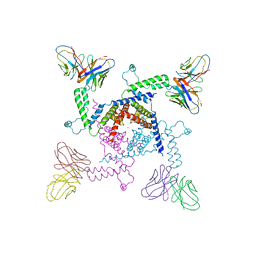 | | Crystal structure of KvAP-33H1 Fv complex | | Descriptor: | 33H1 Fv fragment, POTASSIUM ION, Voltage-gated potassium channel | | Authors: | Lee, S.Y, Lee, A, Chen, J, Mackinnon, R. | | Deposit date: | 2005-06-16 | | Release date: | 2005-11-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.9 Å) | | Cite: | Structure of the KvAP voltage-dependent K+ channel and its dependence on the lipid membrane.
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
7STU
 
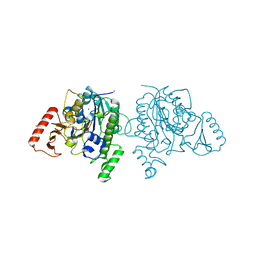 | | Crystal structure of sulfatase from Pedobacter yulinensis | | Descriptor: | BROMIDE ION, CALCIUM ION, N-acetylgalactosamine-6-sulfatase, ... | | Authors: | O'Malley, A, Schlachter, C.R, Grimes, L.L, Tomashek, J.J, Lee, A.L, Chruszcz, M. | | Deposit date: | 2021-11-15 | | Release date: | 2022-01-26 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Purification, Characterization, and Structural Studies of a Sulfatase from Pedobacter yulinensis .
Molecules, 27, 2021
|
|
7STT
 
 | | Crystal structure of sulfatase from Pedobacter yulinensis | | Descriptor: | CALCIUM ION, CHLORIDE ION, MALONATE ION, ... | | Authors: | O'Malley, A, Schlachter, C.R, Grimes, L.L, Tomashek, J.J, Lee, A.L, Chruszcz, M. | | Deposit date: | 2021-11-15 | | Release date: | 2022-01-26 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.603 Å) | | Cite: | Purification, Characterization, and Structural Studies of a Sulfatase from Pedobacter yulinensis .
Molecules, 27, 2021
|
|
7STV
 
 | | Crystal structure of sulfatase from Pedobacter yulinensis | | Descriptor: | CALCIUM ION, CHLORIDE ION, CITRIC ACID, ... | | Authors: | O'Malley, A, Schlachter, C.R, Grimes, L.L, Tomashek, J.J, Lee, A.L, Chruszcz, M. | | Deposit date: | 2021-11-15 | | Release date: | 2022-01-26 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Purification, Characterization, and Structural Studies of a Sulfatase from Pedobacter yulinensis .
Molecules, 27, 2021
|
|
