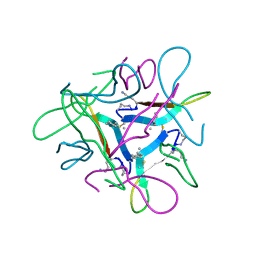
7JXO
 
 | |
5HOY
 
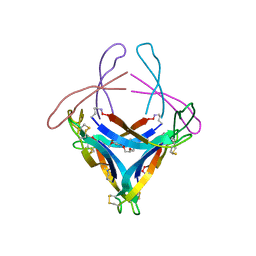 | | X-ray crystallographic structure of an A-beta 17_36 beta-hairpin. X-ray diffractometer data set. (LVFFAEDCGSNKCAII(SAR)LMV). | | Descriptor: | Amyloid beta A4 protein, O-(O-(2-AMINOPROPYL)-O'-(2-METHOXYETHYL)POLYPROPYLENE GLYCOL 500) | | Authors: | Kreutzer, A.G, Spencer, R.K, Nowick, J.S. | | Deposit date: | 2016-01-19 | | Release date: | 2016-03-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.295 Å) | | Cite: | X-ray Crystallographic Structures of a Trimer, Dodecamer, and Annular Pore Formed by an A beta 17-36 beta-Hairpin.
J.Am.Chem.Soc., 138, 2016
|
|
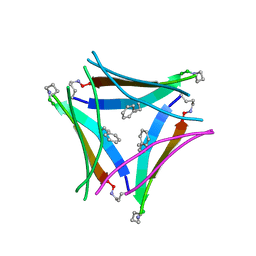
5W4H
 | |
5W4I
 
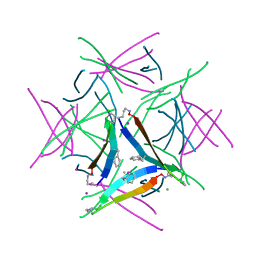 | |
5W4J
 
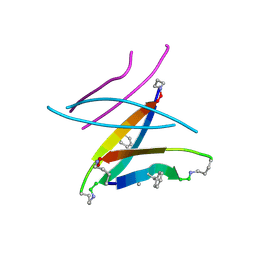 | |
7U4P
 
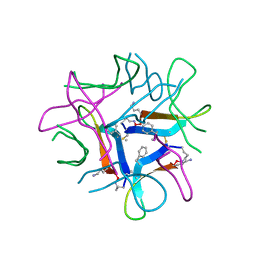 | |
8GJC
 
 | | X-ray crystallographic structure of a beta-hairpin peptide derived from Abeta 17-35. (ORN)LVFFAED(ORN)GAI(N-Me-Ile)GLM | | Descriptor: | DI(HYDROXYETHYL)ETHER, METHIONINE, beta-hairpin peptide derived from Abeta 17-35, ... | | Authors: | Kreutzer, A.G, Ruttenberg, S.M, Nowick, J.S. | | Deposit date: | 2023-03-15 | | Release date: | 2024-01-17 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.431 Å) | | Cite: | beta-Hairpin Alignment Alters Oligomer Formation in A beta-Derived Peptides.
Biochemistry, 63, 2024
|
|
8GJD
 
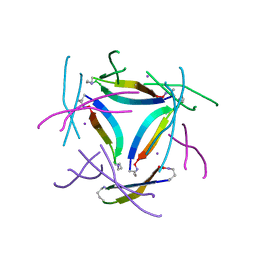 | | X-ray crystallographic structure of a beta-hairpin peptide derived from Abeta 17-36. (ORN)LVFFAED(ORN)AII(N-Me-Gly)LMV | | Descriptor: | Beta-hairpin peptide derived from Abeta 17-36, IODIDE ION | | Authors: | Kreutzer, A.G, Ruttenberg, S.M, Nowick, J.S, Truex, N.L. | | Deposit date: | 2023-03-15 | | Release date: | 2024-01-17 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | beta-Hairpin Alignment Alters Oligomer Formation in A beta-Derived Peptides.
Biochemistry, 63, 2024
|
|
8CUG
 
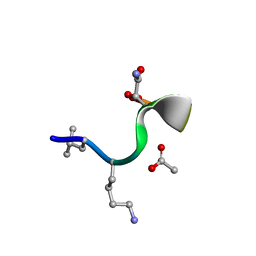 | | Synthetic epi-Novo29 (2R,3S), synchrotron structure | | Descriptor: | ACETATE ION, Synthetic epi-Novo29 (2R,3S) | | Authors: | Kreutzer, A.G, Li, X, Krumberger, M, Nowick, J.S. | | Deposit date: | 2022-05-17 | | Release date: | 2023-01-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.131 Å) | | Cite: | Synthesis and Stereochemical Determination of the Peptide Antibiotic Novo29.
J.Org.Chem., 88, 2023
|
|
8CUF
 
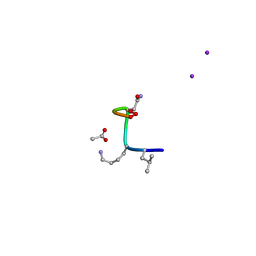 | | Synthetic epi-Novo29 (2R,3S), X-ray diffractometer structure | | Descriptor: | ACETATE ION, IODIDE ION, Synthetic epi-Novo29 (2R,3S) | | Authors: | Kreutzer, A.G, Li, X, Krumberger, M, Nowick, J.S. | | Deposit date: | 2022-05-17 | | Release date: | 2023-01-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Synthesis and Stereochemical Determination of the Peptide Antibiotic Novo29.
J.Org.Chem., 88, 2023
|
|
5SUS
 
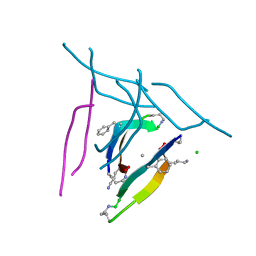 | | X-ray crystallographic structure of a covalent trimer derived from A-beta 17_36. X-ray diffractometer data set. (ORN)CVF(MEA)CED(ORN)AIIGL(ORN)V. | | Descriptor: | 16mer A-beta peptide: ORN-CYS-VAL-PHE-MEA-CYS-GLU-ASP-ORN-ALA-ILE-ILE-GLY-LEU-ORN-VAL, CHLORIDE ION, SODIUM ION | | Authors: | Kreutzer, A.G, Yoo, S, Nowick, J.S. | | Deposit date: | 2016-08-03 | | Release date: | 2017-01-11 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.349 Å) | | Cite: | Stabilization, Assembly, and Toxicity of Trimers Derived from A beta.
J.Am.Chem.Soc., 139, 2017
|
|
5SUT
 
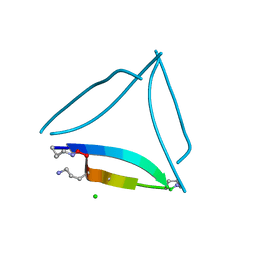 | | X-ray crystallographic structure of a covalent trimer derived from A-beta 17_36. Synchrotron data set. (ORN)CVFFCED(ORN)AII(SAR)L(ORN)V. | | Descriptor: | 16mer A-beta peptide: ORN-CYS-VAL-PHE-PHE-CYS-GLU-ASP-ORN-ALA-ILE-ILE-SAR-LEU-ORN-VAL, CHLORIDE ION | | Authors: | Kreutzer, A.G, Spencer, R.K, Nowick, J.S. | | Deposit date: | 2016-08-03 | | Release date: | 2017-01-11 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.902 Å) | | Cite: | Stabilization, Assembly, and Toxicity of Trimers Derived from A beta.
J.Am.Chem.Soc., 139, 2017
|
|
5SUR
 
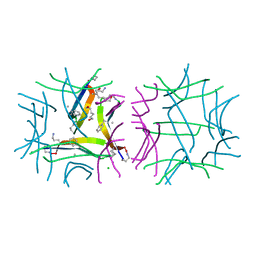 | | X-ray crystallographic structure of a covalent trimer derived from A-beta 17_36. Synchrotron data set. (ORN)CVF(MEA)CED(ORN)AIIGL(ORN)V. | | Descriptor: | 16mer A-beta peptide: ORN-CYS-VAL-PHE-MEA-CYS-GLU-ASP-ORN-ALA-ILE-ILE-GLY-LEU-ORN-VAL, CHLORIDE ION, HEXANE-1,6-DIOL, ... | | Authors: | Kreutzer, A.G, Yoo, S, Nowick, J.S. | | Deposit date: | 2016-08-03 | | Release date: | 2017-01-11 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.801 Å) | | Cite: | Stabilization, Assembly, and Toxicity of Trimers Derived from A beta.
J.Am.Chem.Soc., 139, 2017
|
|
7JQT
 
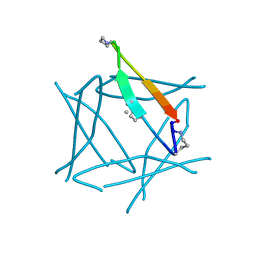 | |
7JQU
 
 | |
7JQS
 
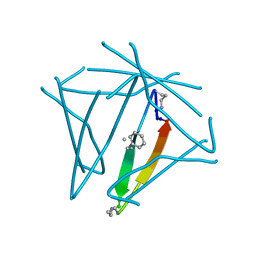 | |
7JQR
 
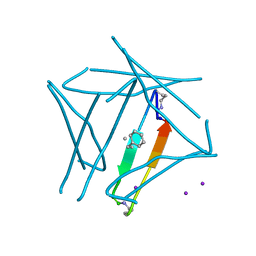 | | Abeta 16-36 beta-hairpin mimic with E22G Arctic mutation | | Descriptor: | Abeta 16-36 beta-hairpin mimic VAL-ORN-LYS-LEU-VAL-MEA-PHE-ALA-GLY-ORN-ALA-ILE-ILE-GLY-LEU-MET, IODIDE ION | | Authors: | Kreutzer, A.G, McKnelly, K.J, Nowick, J.S. | | Deposit date: | 2020-08-11 | | Release date: | 2021-09-08 | | Last modified: | 2022-03-30 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Effects of Familial Alzheimer's Disease Mutations on the Assembly of a beta-Hairpin Peptide Derived from A beta 16-36 .
Biochemistry, 61, 2022
|
|
7JXN
 
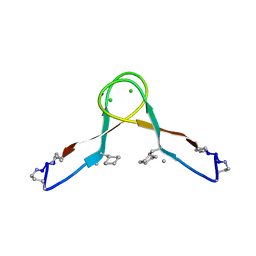 | |
5HOX
 
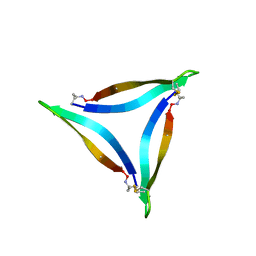 | | X-ray crystallographic structure of an A-beta 17_36 beta-hairpin. Synchrotron data set. (LVFFAEDCGSNKCAII(SAR)LMV). | | Descriptor: | Amyloid beta A4 protein, O-(O-(2-AMINOPROPYL)-O'-(2-METHOXYETHYL)POLYPROPYLENE GLYCOL 500) | | Authors: | Kreutzer, A.G, Nowick, J.S, Spencer, R.K. | | Deposit date: | 2016-01-19 | | Release date: | 2016-03-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | X-ray Crystallographic Structures of a Trimer, Dodecamer, and Annular Pore Formed by an A beta 17-36 beta-Hairpin.
J.Am.Chem.Soc., 138, 2016
|
|
5HOW
 
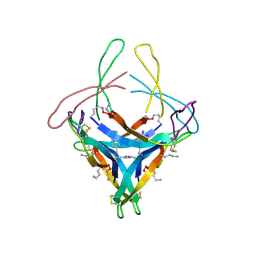 | |
8EC9
 
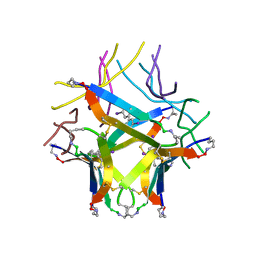 | |
8ECA
 
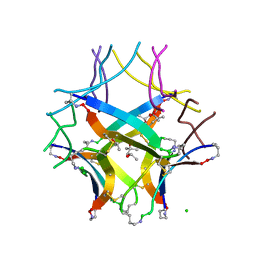 | | Covalently stabilized triangular trimer derived from Abeta16-36 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CHLORIDE ION, ORN-LYS-LEU-VAL-PHE-PHE-ALA-GLU-ORN-CYS-ILE-ILE-SAR-CYS-MET | | Authors: | Kreutzer, A.G, Yoo, S, Nowick, J.S. | | Deposit date: | 2022-09-01 | | Release date: | 2023-05-31 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Probing differences among A beta oligomers with two triangular trimers derived from A beta.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
5SUU
 
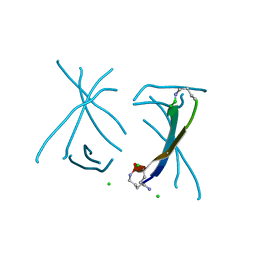 | | X-ray crystallographic structure of a covalent trimer derived from A-beta 17-36. X-ray diffractometer data set. (ORN)CVFFCED(ORN)AII(SAR)L(ORN)V. | | Descriptor: | 16mer A-beta peptide: ORN-CYS-VAL-PHE-PHE-CYS-GLU-ASP-ORN-ALA-ILE-ILE-SAR-LEU-ORN-VAL, CHLORIDE ION, IODIDE ION | | Authors: | Kreutzer, A.G, Spencer, R.K, Nowick, J.S. | | Deposit date: | 2016-08-03 | | Release date: | 2017-01-11 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.032 Å) | | Cite: | Stabilization, Assembly, and Toxicity of Trimers Derived from A beta.
J.Am.Chem.Soc., 139, 2017
|
|
5V65
 
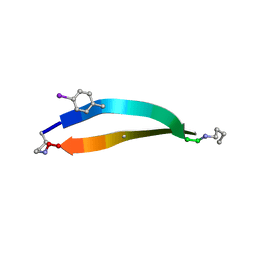 | |
8T0O
 
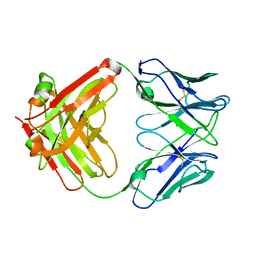 | | Fab from mAb RB2AT_87 | | Descriptor: | CHLORIDE ION, RB2AT_87 Fab Heavy chain, RB2AT_87 Fab Light chain | | Authors: | Kreutzer, A.G, Malonis, R.J, Lai, J.R, Nowick, J.S. | | Deposit date: | 2023-06-01 | | Release date: | 2024-03-27 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Generation and Study of Antibodies against Two Triangular Trimers Derived from A beta.
Pept Sci (Hoboken), 116, 2024
|
|
