4Q4M
 
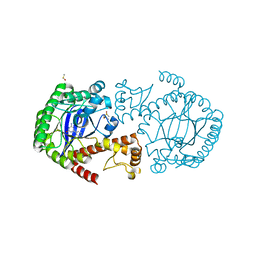 | | tRNA-Guanine Transglycosylase (TGT) in Complex with 6-Amino-4-phenyl-1,2-dihydro-1,3,5-triazin-2-one | | Descriptor: | 6-amino-4-phenyl-1,3,5-triazin-2(1H)-one, DIMETHYL SULFOXIDE, Queuine tRNA-ribosyltransferase, ... | | Authors: | Neeb, M, Heine, A, Klebe, G. | | Deposit date: | 2014-04-15 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.621 Å) | | Cite: | 5-Azacytosines as a Novel Scaffold to Inhibit Z. mobilis TGT with Expected Improved Bioavailability and Synthetic Accessibility
To be Published
|
|
5JT7
 
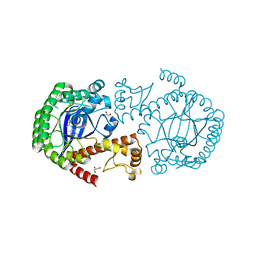 | |
3EGK
 
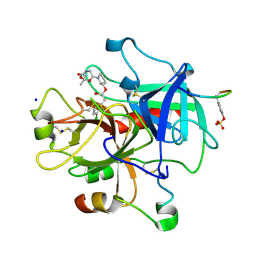 | | KNOBLE Inhibitor | | Descriptor: | Hirudin variant-1, SODIUM ION, Thrombin heavy chain, ... | | Authors: | Baum, B, Heine, A, Klebe, G, Muenzel, M. | | Deposit date: | 2008-09-10 | | Release date: | 2008-09-30 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | KNOBLE: a knowledge-based approach for the design and synthesis of readily accessible small-molecule chemical probes to test protein binding
Angew.Chem.Int.Ed.Engl., 46, 2007
|
|
6Y2O
 
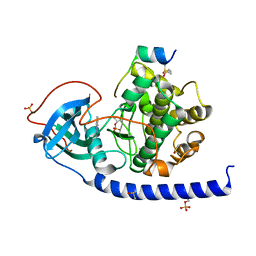 | | Crystal structure of the cAMP-dependent protein kinase A cocrystallized with 1,7-Naphthyridin-8-amine and PKI (5-24) | | Descriptor: | 1,7-naphthyridin-8-amine, DIMETHYL SULFOXIDE, cAMP-dependent protein kinase catalytic subunit alpha, ... | | Authors: | Oebbeke, M, Heine, A, Klebe, G. | | Deposit date: | 2020-02-17 | | Release date: | 2020-09-30 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Fragment based drug design - Small chemical changes of fragments effecting big changes in binding
To Be Published
|
|
3EQ0
 
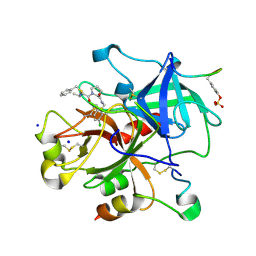 | | Thrombin Inhibitor | | Descriptor: | (2S)-N-[[2-(aminomethyl)-5-chloro-phenyl]methyl]-1-[(2R)-5-carbamimidamido-2-(phenylmethylsulfonylamino)pentanoyl]pyrrolidine-2-carboxamide, Hirudin variant-1, SODIUM ION, ... | | Authors: | Baum, B, Heine, A, Klebe, G, Steinmetzer, T. | | Deposit date: | 2008-09-30 | | Release date: | 2009-08-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Thrombin Inhibition
To be Published
|
|
3F68
 
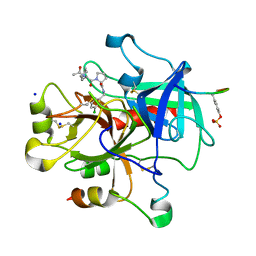 | | Thrombin Inhibition | | Descriptor: | Hirudin variant-2, N-acetyl-3-cyclohexyl-D-alanyl-N-(3-chlorobenzyl)-L-prolinamide, Prothrombin, ... | | Authors: | Baum, B, Heine, A, Klebe, G. | | Deposit date: | 2008-11-05 | | Release date: | 2009-10-20 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Think twice: understanding the high potency of bis(phenyl)methane inhibitors of thrombin.
J.Mol.Biol., 391, 2009
|
|
6YNT
 
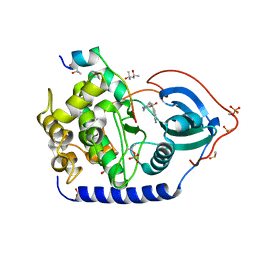 | | Crystal structure of the cAMP-dependent protein kinase A in complex with aminofasudil and PKI (5-24) | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, 5-(1,4-diazepan-1-ylsulfonyl)isoquinolin-1-amine, ... | | Authors: | Oebbeke, M, Gerber, H.-D, Heine, A, Klebe, G. | | Deposit date: | 2020-04-14 | | Release date: | 2020-10-14 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Two Methods, One Goal: Structural Differences between Cocrystallization and Crystal Soaking to Discover Ligand Binding Poses.
Chemmedchem, 16, 2021
|
|
6Y8C
 | |
3CKT
 | | HIV-1 protease in complex with a dimethylallyl decorated pyrrolidine based inhibitor (orthorombic space group) | | Descriptor: | (3S,4S),-3,4-Bis-[(4-carbamoyl-benzensulfonyl)-(3-methyl-but-2-enyl)-amino]-pyrrolidine, CHLORIDE ION, Protease | | Authors: | Boettcher, J, Blum, A, Heine, A, Diederich, W.E, Klebe, G. | | Deposit date: | 2008-03-17 | | Release date: | 2009-03-24 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Two Solutions for the Same Problem: Multiple Binding Modes of Pyrrolidine-Based HIV-1 Protease Inhibitors
J.Mol.Biol., 410, 2011
|
|
6YNR
 
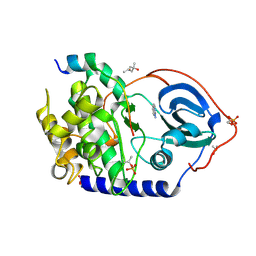 | | Crystal structure of the cAMP-dependent protein kinase A in complex with 1,7-Naphthyridin-8-amine (soaked) and PKI (5-24) | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 1,7-naphthyridin-8-amine, DIMETHYL SULFOXIDE, ... | | Authors: | Oebbeke, M, Heine, A, Klebe, G. | | Deposit date: | 2020-04-14 | | Release date: | 2020-10-14 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Two Methods, One Goal: Structural Differences between Cocrystallization and Crystal Soaking to Discover Ligand Binding Poses.
Chemmedchem, 16, 2021
|
|
6Y05
 
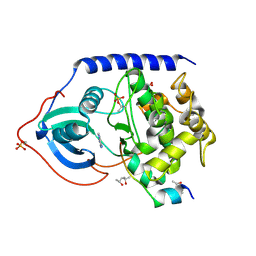 | | Crystal structure of the cAMP-dependent protein kinase A cocrystallized with adenine and PKI (5-24) | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, ADENINE, ... | | Authors: | Oebbeke, M, Wienen-Schmidt, B, Heine, A, Klebe, G. | | Deposit date: | 2020-02-06 | | Release date: | 2021-02-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Fragment based drug design - Small chemical changes of fragments effecting big changes in binding
To Be Published
|
|
6Y89
 | | Crystal structure of the cAMP-dependent protein kinase A cocrystallized with Methyl 5-isoquinolinecarboxylate and PKI (5-24) | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, METHANOL, ... | | Authors: | Oebbeke, M, Gerber, H.-D, Heine, A, Klebe, G. | | Deposit date: | 2020-03-04 | | Release date: | 2021-03-31 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Fragment based drug design - Small chemical changes of fragments effecting big changes in binding
To Be Published
|
|
3DUX
 
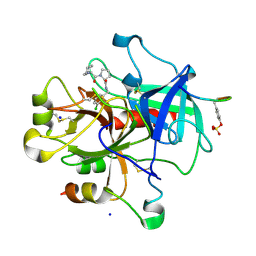 | | Understanding Thrombin Inhibition | | Descriptor: | 3-cyclohexyl-D-alanyl-N-(3-chlorobenzyl)-L-prolinamide, Hirudin variant-1, SODIUM ION, ... | | Authors: | Baum, B, Heine, A, Klebe, G. | | Deposit date: | 2008-07-18 | | Release date: | 2009-06-23 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Think twice: understanding the high potency of bis(phenyl)methane inhibitors of thrombin
J.Mol.Biol., 391, 2009
|
|
3DHK
 
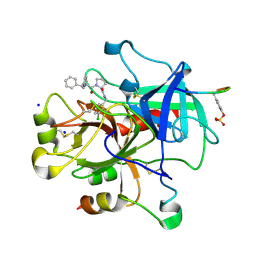 | | Bisphenylic Thrombin Inhibitors | | Descriptor: | Hirudin variant-1, SODIUM ION, Thrombin heavy chain, ... | | Authors: | Baum, B, Heine, A, Klebe, G. | | Deposit date: | 2008-06-18 | | Release date: | 2009-05-19 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Think twice: understanding the high potency of bis(phenyl)methane inhibitors of thrombin
J.Mol.Biol., 391, 2009
|
|
4Y4B
 
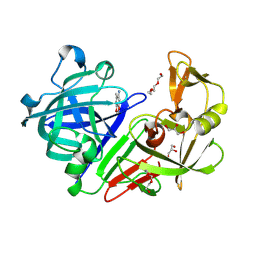 | | Endothiapepsin in complex with fragment 323 | | Descriptor: | 6,6,8-trimethyl-5,6-dihydro[1,3]dioxolo[4,5-g]quinoline, ACETATE ION, Endothiapepsin, ... | | Authors: | Fu, K, Heine, A, Klebe, G. | | Deposit date: | 2015-02-10 | | Release date: | 2016-02-17 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.108 Å) | | Cite: | Crystallographic Fragment Screening of an Entire Library
To Be Published
|
|
3F2P
 
 | | Thermolysin inhibition | | Descriptor: | 3-methyl-2-(propanoyloxy)benzoic acid, CALCIUM ION, DIMETHYL SULFOXIDE, ... | | Authors: | Englert, L, Heine, A, Klebe, G. | | Deposit date: | 2008-10-30 | | Release date: | 2009-11-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Fragment-Based Lead Discovery: Screening and Optimizing Fragments for Thermolysin Inhibition.
Chemmedchem, 5, 2010
|
|
3FCQ
 
 | | Thermolysin inhibition | | Descriptor: | 2-(acetyloxy)-3-methylbenzoic acid, CALCIUM ION, Thermolysin, ... | | Authors: | Steuber, H, Englert, L, Silber, K, Heine, A, Klebe, G. | | Deposit date: | 2008-11-22 | | Release date: | 2009-12-08 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Fragment-Based Lead Discovery: Screening and Optimizing Fragments for Thermolysin Inhibition.
Chemmedchem, 5, 2010
|
|
3D49
 
 | | Thrombin Inhibition | | Descriptor: | BENZAMIDINE, Hirudin variant-1, SODIUM ION, ... | | Authors: | Baum, B, Heine, A, Klebe, G. | | Deposit date: | 2008-05-14 | | Release date: | 2009-04-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Understanding Thrombin Inhibition
To be Published
|
|
6Z44
 
 | | Crystal structure of the cAMP-dependent protein kinase A in complex with phenol | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, DIMETHYL SULFOXIDE, PHENOL, ... | | Authors: | Oebbeke, M, Heine, A, Klebe, G. | | Deposit date: | 2020-05-22 | | Release date: | 2020-12-09 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Fragment Binding to Kinase Hinge: If Charge Distribution and Local pK a Shifts Mislead Popular Bioisosterism Concepts.
Angew.Chem.Int.Ed.Engl., 60, 2021
|
|
3F28
 
 | | Thermolysin inhibition | | Descriptor: | 2-[(cyclopropylcarbonyl)oxy]-3-methylbenzoic acid, CALCIUM ION, Thermolysin, ... | | Authors: | Englert, L, Heine, A, Klebe, G. | | Deposit date: | 2008-10-29 | | Release date: | 2009-11-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Fragment-Based Lead Discovery: Screening and Optimizing Fragments for Thermolysin Inhibition.
Chemmedchem, 5, 2010
|
|
3FB0
 
 | |
3FBO
 
 | |
3FLF
 
 | | Thermolysin inhibition | | Descriptor: | CALCIUM ION, GLYCEROL, N-[(S)-({[(benzyloxy)carbonyl]amino}methyl)(hydroxy)phosphoryl]-L-valyl-L-leucine, ... | | Authors: | Englert, L, Biela, A, Heine, A, Klebe, G. | | Deposit date: | 2008-12-18 | | Release date: | 2010-01-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Displacement of disordered water molecules from hydrophobic pocket creates enthalpic signature: binding of phosphonamidate to the S1'-pocket of thermolysin.
Biochim.Biophys.Acta, 1800, 2010
|
|
3FXP
 
 | | Thermolysin inhibition | | Descriptor: | CALCIUM ION, D-glucose, N~2~-[(2S)-2-{[1-(4-carboxybenzyl)-1H-1,2,3-triazol-4-yl]methyl}-3-methylbutanoyl]-L-lysine, ... | | Authors: | Englert, L, Heine, A, Klebe, G. | | Deposit date: | 2009-01-21 | | Release date: | 2010-02-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Thermolysin in complex with triazolic inhibitor
To be Published
|
|
2POT
 
 | | tRNA guanine transglycosylase (TGT) E235Q mutant in complex with guanine | | Descriptor: | GLYCEROL, GUANINE, Queuine tRNA-ribosyltransferase, ... | | Authors: | Tidten, N, Heine, A, Reuter, K, Klebe, G. | | Deposit date: | 2007-04-27 | | Release date: | 2007-10-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Glutamate versus Glutamine Exchange Swaps Substrate Selectivity in tRNA-Guanine Transglycosylase: Insight into the Regulation of Substrate Selectivity by Kinetic and Crystallographic Studies.
J.Mol.Biol., 374, 2007
|
|
