4ZTT
 
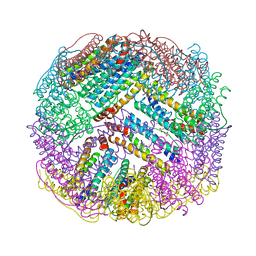 | | Crystal structures of ferritin mutants reveal diferric-peroxo intermediates | | Descriptor: | Bacterial non-heme ferritin, FE (II) ION, FE (III) ION, ... | | Authors: | Kim, S, Park, Y.H, Jung, S.W, Seok, J.H, Chung, Y.B, Lee, D.B, Gowda, G, Lee, J.H, Han, H.R, Cho, A.E, Lee, C, Chung, M.S, Kim, K.H. | | Deposit date: | 2015-05-15 | | Release date: | 2016-06-15 | | Last modified: | 2020-02-19 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Structural Basis of Novel Iron-Uptake Route and Reaction Intermediates in Ferritins from Gram-Negative Bacteria.
J. Mol. Biol., 428, 2016
|
|
7DNZ
 
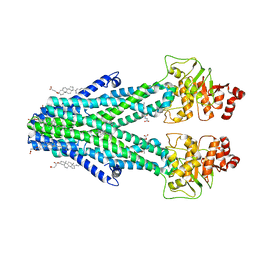 | | Cryo-EM structure of the human ABCB6 (Hemin and GSH-bound) | | Descriptor: | ATP-binding cassette sub-family B member 6, mitochondrial, CHOLESTEROL HEMISUCCINATE, ... | | Authors: | Kim, S, Lee, S.S, Park, J.G, Kim, J.W, Kim, S, Kim, N.J, Hong, S, Kang, J.Y, Jin, M.S. | | Deposit date: | 2020-12-11 | | Release date: | 2022-06-22 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structural Insights into Porphyrin Recognition by the Human ATP-Binding Cassette Transporter ABCB6.
Mol.Cells, 45, 2022
|
|
7DNY
 
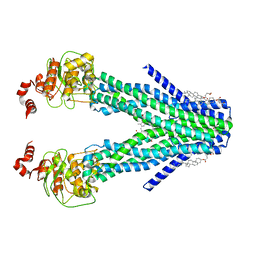 | | Cryo-EM structure of the human ABCB6 (coproporphyrin III-bound) | | Descriptor: | ATP-binding cassette sub-family B member 6, mitochondrial, CHOLESTEROL HEMISUCCINATE, ... | | Authors: | Kim, S, Lee, S.S, Park, J.G, Kim, J.W, Kim, S, Kim, N.J, Hong, S, Kang, J.Y, Jin, M.S. | | Deposit date: | 2020-12-11 | | Release date: | 2022-06-22 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structural Insights into Porphyrin Recognition by the Human ATP-Binding Cassette Transporter ABCB6.
Mol.Cells, 45, 2022
|
|
7C1Y
 
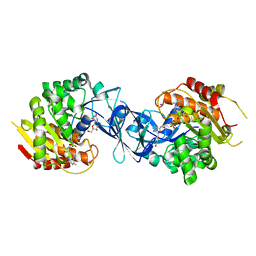 | | Pseudouridine and ADP bound structure of Pseudouridine kinase (PUKI) from Arabidopsis thaliana | | Descriptor: | 5-[(2~{S},3~{R},4~{S},5~{R})-5-(hydroxymethyl)-3,4-bis(oxidanyl)oxolan-2-yl]-1~{H}-pyrimidine-2,4-dione, ADENOSINE-5'-DIPHOSPHATE, PfkB-like carbohydrate kinase family protein, ... | | Authors: | Kim, S.H, Rhee, S. | | Deposit date: | 2020-05-06 | | Release date: | 2020-11-18 | | Last modified: | 2021-06-02 | | Method: | X-RAY DIFFRACTION (2.083426 Å) | | Cite: | Structural basis for the substrate specificity and catalytic features of pseudouridine kinase from Arabidopsis thaliana.
Nucleic Acids Res., 49, 2021
|
|
6KD7
 
 | | Crystal structure of geranylgeranyl pyrophosphate synthase | | Descriptor: | GLYCEROL, MAGNESIUM ION, PYROPHOSPHATE, ... | | Authors: | Kim, S, Kim, K.-J. | | Deposit date: | 2019-07-01 | | Release date: | 2019-09-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of geranylgeranyl pyrophosphate synthase (crtE) from Nonlabens dokdonensis DSW-6.
Biochem.Biophys.Res.Commun., 518, 2019
|
|
1OPY
 
 | | KSI | | Descriptor: | DELTA5-3-KETOSTEROID IOSMERASE | | Authors: | Kim, S.-W, Cha, S.-S, Cho, H.-S, Kim, J.-S, Ha, N.-C, Cho, M.-J, Choi, K.-Y, Oh, B.-H. | | Deposit date: | 1997-05-23 | | Release date: | 1998-04-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | High-resolution crystal structures of delta5-3-ketosteroid isomerase with and without a reaction intermediate analogue.
Biochemistry, 36, 1997
|
|
1AIN
 
 | |
7C1Z
 
 | | ATP bound structure of Pseudouridine kinase (PUKI) from Arabidopsis thaliana | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, PfkB-like carbohydrate kinase family protein, ... | | Authors: | Kim, S.H, Rhee, S. | | Deposit date: | 2020-05-06 | | Release date: | 2020-11-18 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.09617043 Å) | | Cite: | Structural basis for the substrate specificity and catalytic features of pseudouridine kinase from Arabidopsis thaliana.
Nucleic Acids Res., 49, 2021
|
|
6KIE
 
 | | Crystal structure of human leucyl-tRNA synthetase, Leu-AMS-bound form | | Descriptor: | 5'-O-(L-leucylsulfamoyl)adenosine, LEUCINE, Leucine--tRNA ligase, ... | | Authors: | Kim, S, Son, J, Kim, S, Hwang, K.Y. | | Deposit date: | 2019-07-18 | | Release date: | 2021-01-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Leucine-sensing mechanism of leucyl-tRNA synthetase 1 for mTORC1 activation.
Cell Rep, 35, 2021
|
|
132D
 
 | |
7C1X
 
 | |
7Y1H
 | | Controlling fibrosis using compound with novel binding mode to prolyl-tRNA synthetase 1 | | Descriptor: | 1-(5-chloranyl-4-methyl-benzimidazol-1-yl)-3-[(2R,3S)-3-oxidanylpiperidin-2-yl]propan-2-one, ADENOSINE-5'-TRIPHOSPHATE, Bifunctional glutamate/proline--tRNA ligase, ... | | Authors: | Kim, S, Yoon, I, Son, J, Park, S, Hwang, K.Y. | | Deposit date: | 2022-06-08 | | Release date: | 2023-07-05 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Control of fibrosis with enhanced safety via asymmetric inhibition of prolyl-tRNA synthetase 1.
Embo Mol Med, 15, 2023
|
|
7Y1W
 
 | | Controlling fibrosis using compound with novel binding mode to prolyl-tRNA synthetase 1 | | Descriptor: | (2R,3S)-2-[3-[4,5-bis(chloranyl)benzimidazol-1-yl]propyl]piperidin-3-ol, ADENOSINE-5'-TRIPHOSPHATE, Bifunctional glutamate/proline--tRNA ligase, ... | | Authors: | Kim, S, Yoon, I, Son, J, Park, S, Hwang, K.Y. | | Deposit date: | 2022-06-09 | | Release date: | 2023-07-05 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Control of fibrosis with enhanced safety via asymmetric inhibition of prolyl-tRNA synthetase 1.
Embo Mol Med, 15, 2023
|
|
7Y28
 
 | | Controlling fibrosis using compound with novel binding mode to prolyl-tRNA synthetase 1 | | Descriptor: | 1-[6-(3-fluorophenyl)benzimidazol-1-yl]-3-[(2R,3S)-3-oxidanylpiperidin-2-yl]propan-2-one, ADENOSINE-5'-TRIPHOSPHATE, Bifunctional glutamate/proline--tRNA ligase, ... | | Authors: | Kim, S, Yoon, I, Son, J, Park, S, Hwang, K.Y. | | Deposit date: | 2022-06-09 | | Release date: | 2023-07-05 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Control of fibrosis with enhanced safety via asymmetric inhibition of prolyl-tRNA synthetase 1.
Embo Mol Med, 15, 2023
|
|
7Y3S
 
 | | Controlling fibrosis using compound with novel binding mode to prolyl-tRNA synthetase 1 | | Descriptor: | 1-(6-bromanyl-7-methyl-imidazo[4,5-b]pyridin-3-yl)-3-[(2R,3S)-3-oxidanylpiperidin-2-yl]propan-2-one, ADENOSINE-5'-TRIPHOSPHATE, Bifunctional glutamate/proline--tRNA ligase, ... | | Authors: | Kim, S, Yoon, I, Son, J, Park, S, Hwang, K.Y. | | Deposit date: | 2022-06-12 | | Release date: | 2023-07-05 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Control of fibrosis with enhanced safety via asymmetric inhibition of prolyl-tRNA synthetase 1.
Embo Mol Med, 15, 2023
|
|
6K8U
 
 | |
2OLE
 
 | | Crystal Structure Of Human Dipeptidyl Peptidase IV (DPPIV) Complex With Cyclic Hydrazine Derivatives | | Descriptor: | (2R)-4-(2-BENZOYL-1,2-DIAZEPAN-1-YL)-4-OXO-1-(2,4,5-TRIFLUOROPHENYL)BUTAN-2-AMINE, Dipeptidyl peptidase 4 | | Authors: | Kim, S.S, Ahn, J.H, Lee, J.O. | | Deposit date: | 2007-01-19 | | Release date: | 2008-01-22 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Synthesis, biological evaluation and structural determination of beta-aminoacyl-containing cyclic hydrazine derivatives as dipeptidyl peptidase IV (DPP-IV) inhibitors
Bioorg.Med.Chem.Lett., 17, 2007
|
|
6K8W
 
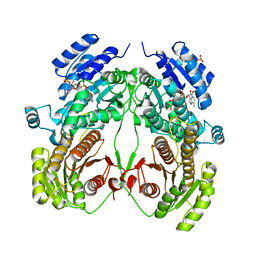 | | Crystal structure of N-domain with NADP of baterial malonyl-CoA reductase | | Descriptor: | NAD-dependent epimerase/dehydratase:Short-chain dehydrogenase/reductase SDR, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, SULFATE ION | | Authors: | Kim, S, Kim, K.-J. | | Deposit date: | 2019-06-13 | | Release date: | 2020-03-18 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.17 Å) | | Cite: | Structural insight into bi-functional malonyl-CoA reductase.
Environ.Microbiol., 22, 2020
|
|
6K8S
 
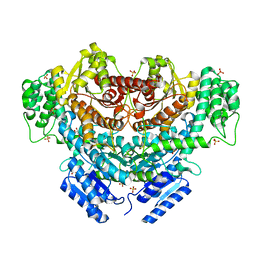 | |
2ESB
 
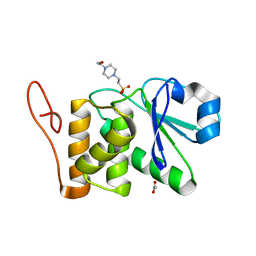 | | Crystal structure of human DUSP18 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ACETATE ION, Dual specificity protein phosphatase 18 | | Authors: | Kim, S.J, Ryu, S.E, Jeong, D.G, Cho, Y.H, Yoon, T.S, Kim, J.H. | | Deposit date: | 2005-10-25 | | Release date: | 2006-06-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of human DSP18, a member of the dual-specificity protein tyrosine phosphatase family.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
4KDL
 
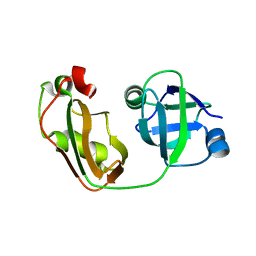 | | Crystal structure of p97/VCP N in complex with OTU1 UBXL | | Descriptor: | Transitional endoplasmic reticulum ATPase, Ubiquitin thioesterase OTU1 | | Authors: | Kim, S.J, Kim, E.E. | | Deposit date: | 2013-04-25 | | Release date: | 2014-03-19 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Structural Basis for Ovarian Tumor Domain-containing Protein 1 (OTU1) Binding to p97/Valosin-containing Protein (VCP).
J.Biol.Chem., 289, 2014
|
|
6L33
 
 | |
7VG4
 
 | | 10,5-methenyltetrahydrofolate cyclohydrolase from Methylobacterium extorquens AM1 strain | | Descriptor: | Methenyltetrahydrofolate cyclohydrolase | | Authors: | Kim, S, Lee, S, Kim, I.-K, Seo, H, Kim, K.-J. | | Deposit date: | 2021-09-14 | | Release date: | 2022-07-06 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.77 Å) | | Cite: | Structural insight into a molecular mechanism of methenyltetrahydrofolate cyclohydrolase from Methylobacterium extorquens AM1.
Int.J.Biol.Macromol., 202, 2022
|
|
8I70
 
 | |
8I6Z
 
 | |
