5X3R
 
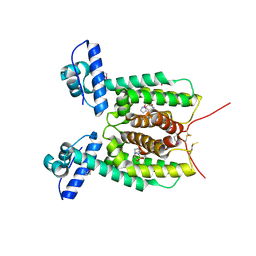 | | Crystal structure of the SmcR complexed with QStatin | | Descriptor: | 1-(5-bromanylthiophen-2-yl)sulfonylpyrazole, LuxR family transcriptional regulator, SULFATE ION | | Authors: | Jang, S.Y, Hwang, J, Kim, M.H. | | Deposit date: | 2017-02-07 | | Release date: | 2018-01-24 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | QStatin, a Selective Inhibitor of Quorum Sensing inVibrioSpecies
MBio, 9, 2018
|
|
402D
 
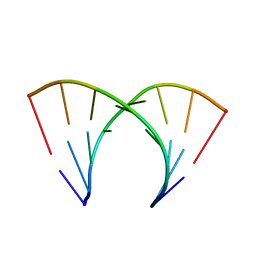 | | 5'-R(*CP*GP*CP*CP*AP*GP*CP*G)-3' | | Descriptor: | RNA(5'-R(*CP*GP*CP*CP*AP*GP*CP*G)-3') | | Authors: | Jang, S.B, Hung, L.-W, Chi, Y.-I, Holbrook, E.L, Carter, R, Holbrook, S.R. | | Deposit date: | 1998-06-04 | | Release date: | 1998-06-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of an RNA internal loop consisting of tandem C-A+ base pairs.
Biochemistry, 37, 1998
|
|
1H3Q
 
 | |
1SAQ
 
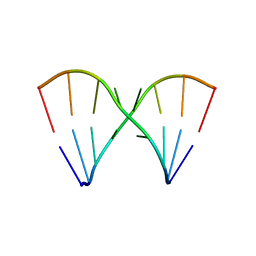 | | Crystal Structure of the RNA octamer GIC(GA)GCC | | Descriptor: | 5'-R(*GP*IP*CP*GP*AP*GP*CP*C)-3' | | Authors: | Jang, S.B, Baeyens, K, Jeong, M.S, SantaLucia Jr, J, Turner, D, Holbrook, S.R. | | Deposit date: | 2004-02-09 | | Release date: | 2004-05-18 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structures of two RNA octamers containing tandem G.A base pairs.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1SA9
 
 | | Crystal Structure of the RNA octamer GGCGAGCC | | Descriptor: | 5'-R(*GP*GP*CP*GP*AP*GP*CP*C)-3' | | Authors: | Jang, S.B, Baeyens, K, Jeong, M.S, SantaLucia Jr, J, Turner, D, Holbrook, S.R. | | Deposit date: | 2004-02-09 | | Release date: | 2004-05-18 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.86 Å) | | Cite: | Structures of two RNA octamers containing tandem G.A base pairs.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
5GLU
 
 | | Tl-gal with SiaLac | | Descriptor: | GALECTIN, N-acetyl-alpha-neuraminic acid-(2-3)-beta-D-galactopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Jang, S.B, Hwang, E.Y. | | Deposit date: | 2016-07-12 | | Release date: | 2016-11-09 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
5GLZ
 
 | | Tl-gal with Glucose | | Descriptor: | beta-D-glucopyranose, galectin | | Authors: | Jang, S.B, Hwang, E.Y. | | Deposit date: | 2016-07-12 | | Release date: | 2016-11-09 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
5GLT
 
 | | Tl-gal with LNT | | Descriptor: | Galectin, beta-D-galactopyranose-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Jang, S.B, Hwang, E.Y. | | Deposit date: | 2016-07-12 | | Release date: | 2016-11-09 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
5GLV
 
 | | Tl-gal | | Descriptor: | galectin | | Authors: | Jang, S.B, Hwang, E.Y. | | Deposit date: | 2016-07-12 | | Release date: | 2016-11-09 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
5GLW
 
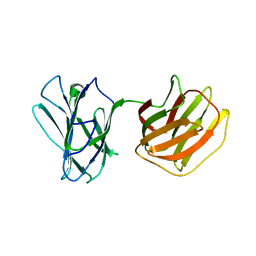 | | Tl-gal with LacNAc | | Descriptor: | beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, galectin | | Authors: | Jang, S.B, Hwang, E.Y. | | Deposit date: | 2016-07-12 | | Release date: | 2016-11-09 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
2G3S
 
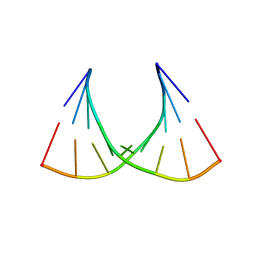 | | RNA structure containing GU base pairs | | Descriptor: | 5'-R(*GP*GP*CP*GP*UP*GP*CP*C)-3', MAGNESIUM ION | | Authors: | Jang, S.B, Hung, L.W, Jeong, M.S, Holbrook, E.L, Chen, X, Turner, D.H, Holbrook, S.R. | | Deposit date: | 2006-02-20 | | Release date: | 2007-01-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.499 Å) | | Cite: | The crystal structure at 1.5 angstroms resolution of an RNA octamer duplex containing tandem G.U basepairs
Biophys.J., 90, 2006
|
|
2GHY
 
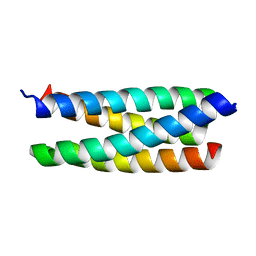 | | Novel Crystal Form of the ColE1 Rom Protein | | Descriptor: | Regulatory protein rop | | Authors: | Jang, S.B, Jeong, M.S, Carter, R.J, Holbrook, E.L, Comolli, L.R, Holbrook, S.R. | | Deposit date: | 2006-03-28 | | Release date: | 2006-05-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Novel crystal form of the ColE1 Rom protein.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
1DE0
 
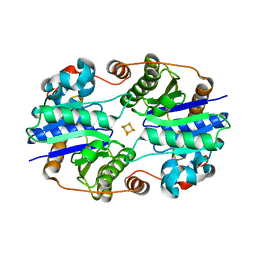 | |
1FP6
 
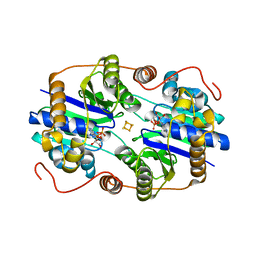 | | THE NITROGENASE FE PROTEIN FROM AZOTOBACTER VINELANDII COMPLEXED WITH MGADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, IRON/SULFUR CLUSTER, MAGNESIUM ION, ... | | Authors: | Jang, S.B, Seefeldt, L.C, Peters, J.W. | | Deposit date: | 2000-08-30 | | Release date: | 2001-01-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Insights into nucleotide signal transduction in nitrogenase: structure of an iron protein with MgADP bound.
Biochemistry, 39, 2000
|
|
7UD4
 
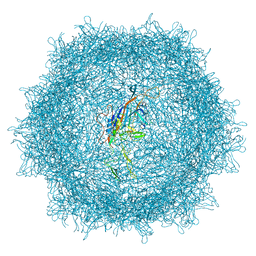 | | Cryo-EM structure of AAV-PHP.eB | | Descriptor: | Capsid protein VP1 | | Authors: | Jang, S, Shen, H.K, Ding, X, Miles, T.F, Gradinaru, V. | | Deposit date: | 2022-03-18 | | Release date: | 2022-09-21 | | Method: | ELECTRON MICROSCOPY (2.24 Å) | | Cite: | Structural basis of receptor usage by the engineered capsid AAV-PHP.eB.
Mol Ther Methods Clin Dev, 26, 2022
|
|
1XC4
 
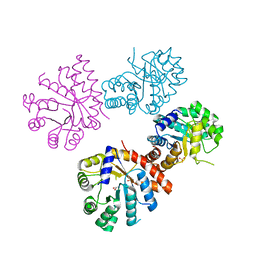 | |
1XD8
 
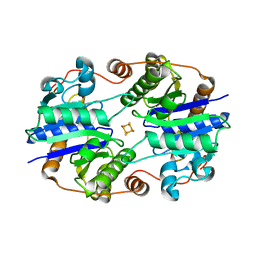 | | Crystal Structure of the Nitrogenase Fe protein Asp39Asn | | Descriptor: | IRON/SULFUR CLUSTER, Nitrogenase iron protein 1 | | Authors: | Jang, S.B, Jeong, M.S, Seefeldt, L.C, Peters, J.W. | | Deposit date: | 2004-09-05 | | Release date: | 2005-03-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural and biochemical implications of single amino acid substitutions in the nucleotide-dependent switch regions of the nitrogenase Fe protein from Azotobacter vinelandii
J.Biol.Inorg.Chem., 9, 2004
|
|
1XCF
 
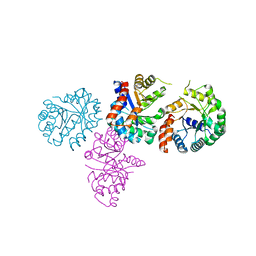 | |
1XDB
 
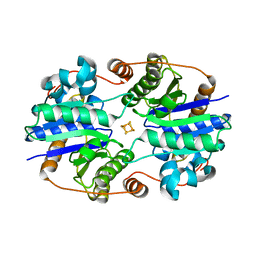 | | Crystal Structure of the Nitrogenase Fe protein Asp129Glu | | Descriptor: | IRON/SULFUR CLUSTER, Nitrogenase iron protein 1 | | Authors: | Jang, S.B, Jeong, M.S, Seefeldt, L.C, Peters, J.W. | | Deposit date: | 2004-09-05 | | Release date: | 2005-03-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural and biochemical implications of single amino acid substitutions in the nucleotide-dependent switch regions of the nitrogenase Fe protein from Azotobacter vinelandii
J.Biol.Inorg.Chem., 9, 2004
|
|
1XD9
 
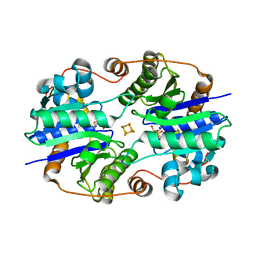 | | Crystal Structure of the Nitrogenase Fe protein Asp39Asn with MgADP bound | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, IRON/SULFUR CLUSTER, MAGNESIUM ION, ... | | Authors: | Jang, S.B, Jeong, M.S, Seefeldt, L.C, Peters, J.W. | | Deposit date: | 2004-09-05 | | Release date: | 2005-03-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural and biochemical implications of single amino acid substitutions in the nucleotide-dependent switch regions of the nitrogenase Fe protein from Azotobacter vinelandii
J.Biol.Inorg.Chem., 9, 2004
|
|
5ZC1
 
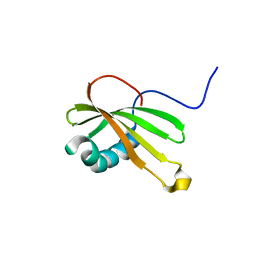 | |
6JM9
 
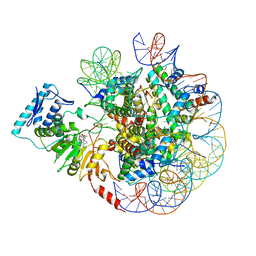 | | cryo-EM structure of DOT1L bound to unmodified nucleosome | | Descriptor: | DNA strand I, DNA strand J, Histone H2A, ... | | Authors: | Jang, S, Song, J.J. | | Deposit date: | 2019-03-07 | | Release date: | 2019-05-15 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (7.3 Å) | | Cite: | Structural basis of recognition and destabilization of the histone H2B ubiquitinated nucleosome by the DOT1L histone H3 Lys79 methyltransferase.
Genes Dev., 33, 2019
|
|
6JMA
 
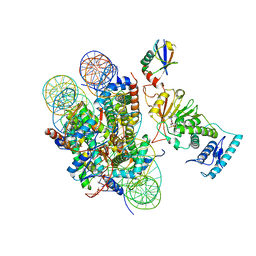 | | cryo-EM structure of DOT1L bound to H2B ubiquitinated nucleosome | | Descriptor: | DNA I&J, Histone H2A, Histone H2B 1.1, ... | | Authors: | Jang, S, Song, J.J. | | Deposit date: | 2019-03-07 | | Release date: | 2019-05-15 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (6.8 Å) | | Cite: | Structural basis of recognition and destabilization of the histone H2B ubiquitinated nucleosome by the DOT1L histone H3 Lys79 methyltransferase.
Genes Dev., 33, 2019
|
|
3GIO
 
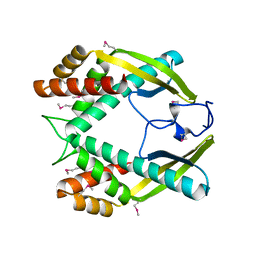 | | Crystal structure of the TNF-alpha inducing protein (Tip alpha) from Helicobacter pylori | | Descriptor: | Putative uncharacterized protein | | Authors: | Jang, J.Y, Yoon, H.J, Yoon, J.Y, Kim, H.S, Lee, S.J, Kim, K.H, Kim, D.J, Han, B.G, Lee, B.I, Jang, S, Suh, S.W. | | Deposit date: | 2009-03-05 | | Release date: | 2009-08-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of the TNF-alpha-Inducing Protein (Tipalpha) from Helicobacter pylori: Insights into Its DNA-Binding Activity.
J.Mol.Biol., 2009
|
|
2OTR
 | |
