8EOP
 
 | | Cryo-EM Structure of Nanodisc reconstituted human ABCA7 EQ mutant in ATP bound closed state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Alam, A, Le, L.T.M, Thompson, J.R. | | Deposit date: | 2022-10-04 | | Release date: | 2022-12-21 | | Last modified: | 2023-02-15 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Cryo-EM structures of human ABCA7 provide insights into its phospholipid translocation mechanisms.
Embo J., 42, 2023
|
|
8F8Y
 
 | | PHF2 (PHD+JMJ) in Complex with VRK1 N-Terminal Peptide | | Descriptor: | 1,2-ETHANEDIOL, Lysine-specific demethylase PHF2, SULFATE ION, ... | | Authors: | Horton, J.R, Cheng, X. | | Deposit date: | 2022-11-22 | | Release date: | 2023-01-18 | | Last modified: | 2023-02-08 | | Method: | X-RAY DIFFRACTION (3.06 Å) | | Cite: | A complete methyl-lysine binding aromatic cage constructed by two domains of PHF2.
J.Biol.Chem., 299, 2022
|
|
8F8Z
 
 | |
1SHK
 
 | |
1GU0
 
 | | CRYSTAL STRUCTURE OF TYPE II DEHYDROQUINASE FROM STREPTOMYCES COELICOLOR | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 3-DEHYDROQUINATE DEHYDRATASE | | Authors: | Roszak, A.W, Krell, T, Robinson, D, Hunter, I.S, Coggins, J.R, Lapthorn, A.J. | | Deposit date: | 2002-01-22 | | Release date: | 2002-04-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Structure and Mechanism of the Type II Dehydroquinase from Streptomyces Coelicolor
Structure, 10, 2002
|
|
8F94
 
 | |
1H3D
 
 | | STRUCTURE OF THE E.COLI ATP-PHOSPHORIBOSYLTRANSFERASE | | Descriptor: | ADENOSINE MONOPHOSPHATE, ATP-PHOSPHORIBOSYLTRANSFERASE, L(+)-TARTARIC ACID | | Authors: | Lohkamp, B, McDermott, G, Coggins, J.R, Lapthorn, A.J. | | Deposit date: | 2002-08-27 | | Release date: | 2003-10-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The Structure of Escherichia Coli ATP-Phosphoribosyltransferase: Identification of Substrate Binding Sites and Mode of AMP Inhibition
J.Mol.Biol., 336, 2004
|
|
3DL2
 
 | | Hexagonal structure of the LDH domain of Human Ubiquitin-conjugating Enzyme E2-like Isoform A | | Descriptor: | PHOSPHATE ION, SODIUM ION, Ubiquitin-conjugating enzyme E2 variant 3 | | Authors: | Walker, J.R, Avvakumov, G.V, Xue, S, Newman, E.M, Finerty Jr, P.J, Butler-Cole, C, Bountra, C, Wolkstrom, M, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2008-06-26 | | Release date: | 2008-07-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Investigation Into the L-Lactate Dehydrogenase Domain of Human Ubiquitin-Conjugating Enzyme E2-Like Isoform A.
To be Published
|
|
1HA3
 
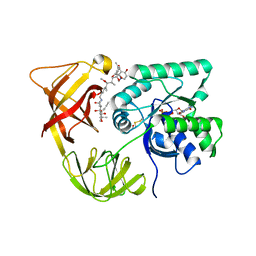 | | ELONGATION FACTOR TU IN COMPLEX WITH aurodox | | Descriptor: | BETA-MERCAPTOETHANOL, ELONGATION FACTOR TU, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Vogeley, L, Palm, G.J, Mesters, J.R, Hilgenfeld, R. | | Deposit date: | 2001-03-26 | | Release date: | 2001-05-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Conformational Change of Elongation Factor TU Induced by Antibiotic Binding: Crystal Structure of the Complex between EF-TU:Gdp and Aurodox
J.Biol.Chem., 276, 2001
|
|
2C4G
 
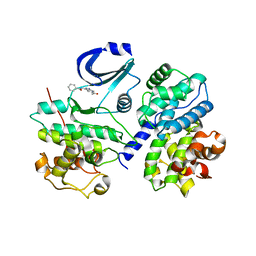 | | STRUCTURE OF CDK2-CYCLIN A WITH PHA-533514 | | Descriptor: | (3Z)-5-ACETYL-3-(BENZOYLIMINO)-3,6-DIHYDROPYRROLO[3,4-C]PYRAZOL-5-IUM, CELL DIVISION PROTEIN KINASE 2, CYCLIN A2, ... | | Authors: | Cameron, A, Fogliatto, G, Pevarello, P, Fancelli, D, Vulpetti, A, Amici, R, Villa, M, Pittala, V, Ciomei, M, Mercurio, C, Bischoff, J.R, Roletto, F, Varasi, M, Brasca, M.G. | | Deposit date: | 2005-10-19 | | Release date: | 2005-11-23 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | 3-Amino-1,4,5,6-Tetrahydropyrrolo[3,4-C]Pyrazoles: A New Class of Cdk2 Inhibitors.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
3MHU
 
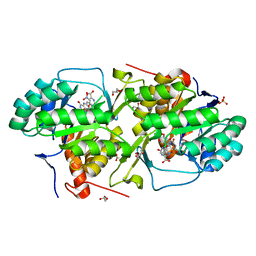 | | Crystal structure of dihydroorotate dehydrogenase from Leishmania major in complex with 5-Nitroorotic acid | | Descriptor: | 5-nitro-2,6-dioxo-1,2,3,6-tetrahydropyrimidine-4-carboxylic acid, Dihydroorotate dehydrogenase, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Pinheiro, M.P, Rocha, J.R, Cheleski, J, Montanari, C.A, Nonato, M.C. | | Deposit date: | 2010-04-08 | | Release date: | 2011-02-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Novel insights for dihydroorotate dehydrogenase class 1A inhibitors discovery.
Eur.J.Med.Chem., 45, 2010
|
|
3DKO
 
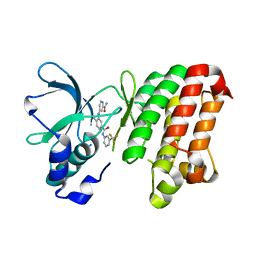 | | Complex between the kinase domain of human ephrin type-a receptor 7 (epha7) and inhibitor alw-ii-49-7 | | Descriptor: | 5-[(2-methyl-5-{[3-(trifluoromethyl)phenyl]carbamoyl}phenyl)amino]pyridine-3-carboxamide, Ephrin type-A receptor 7 | | Authors: | Walker, J.R, Syeda, F, Gray, N, Butler-Cole, C, Bountra, C, Wolkstrom, M, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2008-06-25 | | Release date: | 2008-08-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Kinase domain of human ephrin type-a receptor 7 (epha7) in complex with ALW-II-49-7
To be Published
|
|
3MJY
 
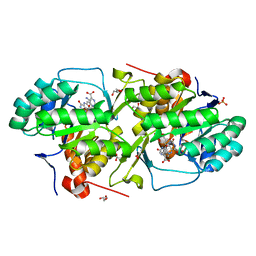 | | Crystal structure of dihydroorotate dehydrogenase from Leishmania major in complex with 5-Aminoorotic acid | | Descriptor: | 5-amino-2,6-dioxo-1,2,3,6-tetrahydropyrimidine-4-carboxylic acid, Dihydroorotate dehydrogenase, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Pinheiro, M.P, Rocha, J.R, Cheleski, J, Montanari, C.A, Nonato, M.C. | | Deposit date: | 2010-04-13 | | Release date: | 2011-02-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Novel insights for dihydroorotate dehydrogenase class 1A inhibitors discovery.
Eur.J.Med.Chem., 45, 2010
|
|
2C4C
 
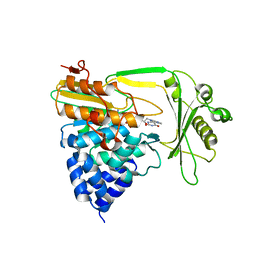 | | Crystal structure of the NADPH-treated monooxygenase domain of MICAL | | Descriptor: | CHLORIDE ION, FLAVIN-ADENINE DINUCLEOTIDE, NEDD9-INTERACTING PROTEIN WITH CALPONIN HOMOLOGY AND LIM DOMAINS | | Authors: | Siebold, C, Berrow, N, Walter, T.S, Harlos, K, Owens, R.J, Terman, J.R, Stuart, D.I, Kolodkin, A.L, Pasterkamp, R.J, Jones, E.Y. | | Deposit date: | 2005-10-18 | | Release date: | 2005-10-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | High-Resolution Structure of the Catalytic Region of Mical (Molecule Interacting with Casl), a Multidomain Flavoenzyme-Signaling Molecule.
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
3BUA
 
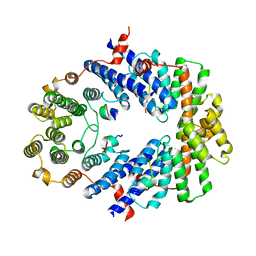 | | Crystal Structure of TRF2 TRFH domain and APOLLO peptide complex | | Descriptor: | DNA cross-link repair 1B protein, Telomeric repeat-binding factor 2 | | Authors: | Chen, Y, Yang, Y, van Overbeek, M, Donigian, J.R, Baciu, P, de Lange, T, Lei, M. | | Deposit date: | 2008-01-02 | | Release date: | 2008-02-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A shared docking motif in TRF1 and TRF2 used for differential recruitment of telomeric proteins.
Science, 319, 2008
|
|
8F4O
 
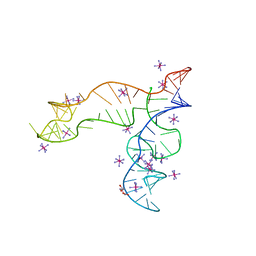 | | Apo structure of the TPP riboswitch aptamer domain | | Descriptor: | IRIDIUM HEXAMMINE ION, TETRAETHYLENE GLYCOL, TPP riboswitch aptamer domain, ... | | Authors: | Lee, H.-K, Wang, Y.-X, Stagno, J.R. | | Deposit date: | 2022-11-11 | | Release date: | 2023-05-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal structure of Escherichia coli thiamine pyrophosphate-sensing riboswitch in the apo state.
Structure, 31, 2023
|
|
3MBW
 
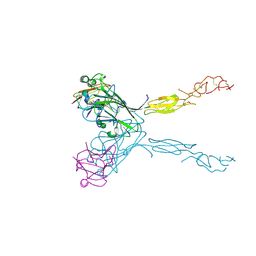 | | Crystal structure of the human ephrin A2 LBD and CRD domains in complex with ephrin A1 | | Descriptor: | Ephrin type-A receptor 2, Ephrin-A1, UNKNOWN ATOM OR ION, ... | | Authors: | Walker, J.R, Yermekbayeva, L, Seitova, A, Butler-Cole, C, Bountra, C, Weigelt, J, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-03-26 | | Release date: | 2010-06-09 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Architecture of Eph receptor clusters.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
1JA8
 
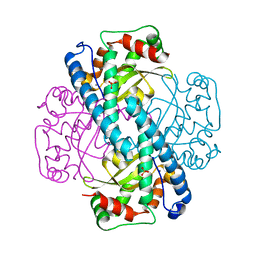 | | Kinetic Analysis of Product Inhibition in Human Manganese Superoxide Dismutase | | Descriptor: | MANGANESE (II) ION, Manganese Superoxide Dismutase, SULFATE ION | | Authors: | Hearn, A.S, Stroupe, M.E, Cabelli, D.E, Lepock, J.R, Tainer, J.A, Nick, H.S, Silverman, D.S. | | Deposit date: | 2001-05-29 | | Release date: | 2001-06-06 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Kinetic analysis of product inhibition in human manganese superoxide dismutase.
Biochemistry, 40, 2001
|
|
1ZV1
 
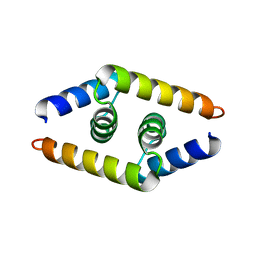 | | Crystal structure of the dimerization domain of doublesex protein from D. melanogaster | | Descriptor: | Doublesex protein | | Authors: | Weiss, M.A, Bayrer, J.R, Wan, Z, Li, B, Phillips, N.B. | | Deposit date: | 2005-06-01 | | Release date: | 2005-08-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Dimerization of doublesex is mediated by a cryptic ubiquitin-associated domain fold: implications for sex-specific gene regulation
J.Biol.Chem., 280, 2005
|
|
1JEY
 
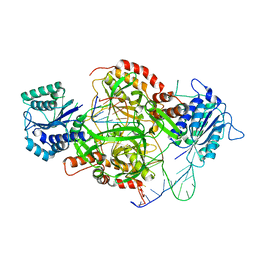 | | Crystal Structure of the Ku heterodimer bound to DNA | | Descriptor: | DNA (34-MER), DNA (5'-D(*GP*TP*TP*TP*TP*TP*AP*GP*TP*TP*TP*AP*TP*TP*GP*GP*GP*CP*GP*CP*G)-3'), Ku70, ... | | Authors: | Walker, J.R, Corpina, R.A, Goldberg, J. | | Deposit date: | 2001-06-19 | | Release date: | 2001-08-10 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the Ku heterodimer bound to DNA and its implications for double-strand break repair.
Nature, 412, 2001
|
|
8EQG
 
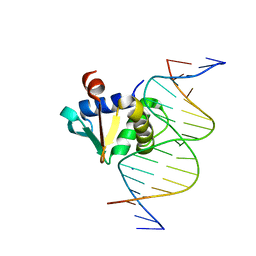 | |
8EKV
 
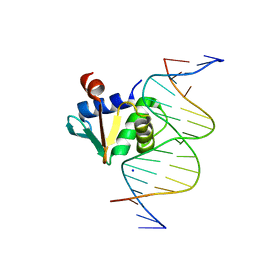 | | Human PU.1 ETS-Domain (165-270) Bound to d(AATAAGCGGATGTGGG) | | Descriptor: | DNA (5'-D(*AP*AP*TP*AP*AP*GP*CP*GP*GP*AP*TP*GP*TP*GP*GP*G)-3'), DNA (5'-D(*TP*CP*CP*CP*AP*CP*AP*TP*CP*CP*GP*CP*TP*TP*AP*T)-3'), SODIUM ION, ... | | Authors: | Terrell, J.R, Poon, G.M.K. | | Deposit date: | 2022-09-22 | | Release date: | 2023-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | DNA selection by the master transcription factor PU.1.
Cell Rep, 42, 2023
|
|
8EM9
 
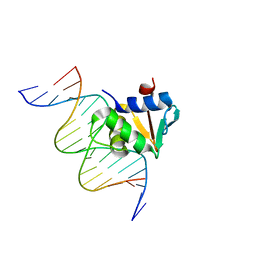 | | Human PU.1 ETS-Domain (165-270) Bound to d(AATAGGAGAAGTAGGG) | | Descriptor: | DNA (5'-D(*AP*AP*TP*AP*GP*GP*AP*GP*AP*AP*GP*TP*AP*GP*GP*G)-3'), DNA (5'-D(*TP*CP*CP*CP*TP*AP*CP*TP*TP*CP*TP*CP*CP*TP*A)-3'), Transcription factor PU.1 | | Authors: | Terrell, J.R, Poon, G.M.K. | | Deposit date: | 2022-09-27 | | Release date: | 2023-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.341 Å) | | Cite: | DNA selection by the master transcription factor PU.1.
Cell Rep, 42, 2023
|
|
8ENG
 
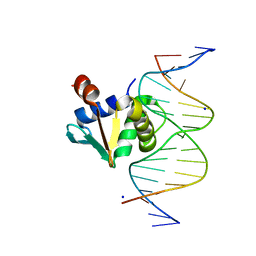 | |
8EK3
 
 | |
