2CKA
 
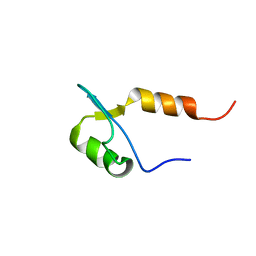 | | Solution structures of the BRK domains of the human Chromo Helicase Domain 7 and 8, reveals structural similarity with GYF domain suggesting a role in protein interaction | | Descriptor: | CHROMODOMAIN-HELICASE-DNA-BINDING PROTEIN 8 | | Authors: | Ab, E, de Jong, R.N, Diercks, T, Xiaoyun, J, Daniels, M, Kaptein, R, Folkers, G.E. | | Deposit date: | 2006-04-14 | | Release date: | 2007-05-08 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Solution Structures of the Brk Domains of the Human Chromo Helicase Domain 7 and 8, Reveals Structural Similarity with Gyf Domain Suggesting a Role in Protein Interaction
To be Published
|
|
2YMB
 
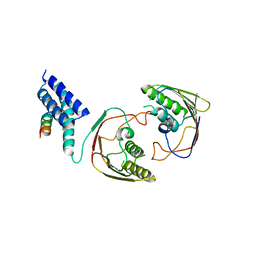 | | Structures of MITD1 | | Descriptor: | CHARGED MULTIVESICULAR BODY PROTEIN 1A, MIT DOMAIN-CONTAINING PROTEIN 1 | | Authors: | Hadders, M.A, Agromayor, M, Obita, T, Perisic, O, Caballe, A, Kloc, M, Lamers, M.H, Williams, R.L, Martin-Serrano, J. | | Deposit date: | 2012-10-08 | | Release date: | 2012-10-24 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.404 Å) | | Cite: | Escrt-III Binding Protein Mitd1 is Involved in Cytokinesis and Has an Unanticipated Pld Fold that Binds Membranes.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
6N60
 
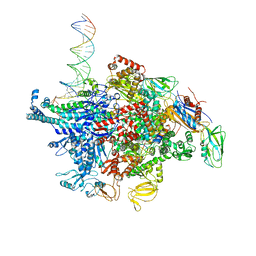 | | Escherichia coli RNA polymerase sigma70-holoenzyme bound to upstream fork promoter DNA and Microcin J25 (MccJ25) | | Descriptor: | DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, DNA-directed RNA polymerase subunit beta', ... | | Authors: | Braffman, N, Hauver, J, Campbell, E.A, Darst, S.A. | | Deposit date: | 2018-11-23 | | Release date: | 2019-01-09 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.68 Å) | | Cite: | Structural mechanism of transcription inhibition by lasso peptides microcin J25 and capistruin.
Proc. Natl. Acad. Sci. U.S.A., 116, 2019
|
|
6N7Z
 
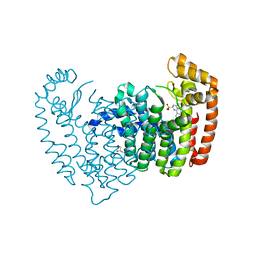 | |
5UCS
 
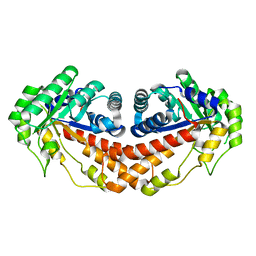 | |
5UD2
 
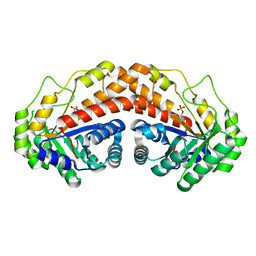 | | Class II fructose-1,6-bisphosphate aldolase H180Q variant of Helicobacter pylori with DHAP | | Descriptor: | 1,3-DIHYDROXYACETONEPHOSPHATE, Fructose-bisphosphate aldolase, SODIUM ION, ... | | Authors: | Jacques, B, Sygusch, J. | | Deposit date: | 2016-12-23 | | Release date: | 2018-04-11 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.775 Å) | | Cite: | Active site remodeling during the catalytic cycle in metal-dependent fructose-1,6-bisphosphate aldolases.
J. Biol. Chem., 293, 2018
|
|
2YMX
 
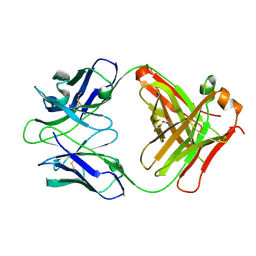 | | Crystal structure of inhibitory anti-AChE Fab408 | | Descriptor: | FAB ANTIBODY HEAVY CHAIN, FAB ANTIBODY LIGHT CHAIN, GLYCEROL | | Authors: | Bourne, Y, Renault, L, Essono, S, Mondielli, G, Lamourette, P, Bocquet, D, Grassi, J, Marchot, P. | | Deposit date: | 2012-10-10 | | Release date: | 2013-10-23 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular Characterization of Monoclonal Antibodies that Inhibit Acetylcholinesterase by Targeting the Peripheral Site and Backdoor Region
Plos One, 8, 2013
|
|
2C77
 
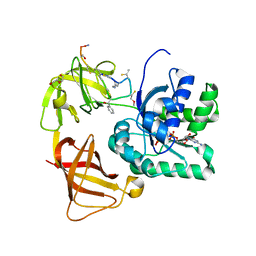 | | EF-Tu complexed with a GTP analog and the antibiotic GE2270 A | | Descriptor: | DI(HYDROXYETHYL)ETHER, ELONGATION FACTOR TU-B, MAGNESIUM ION, ... | | Authors: | Parmeggiani, A, Krab, I.M, Okamura, S, Nielsen, R.C, Nyborg, J, Nissen, P. | | Deposit date: | 2005-11-18 | | Release date: | 2006-03-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural basis of the action of pulvomycin and GE2270 A on elongation factor Tu.
Biochemistry, 45, 2006
|
|
2CNW
 
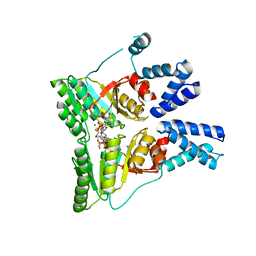 | | GDPALF4 complex of the SRP GTPases Ffh and FtsY | | Descriptor: | CELL DIVISION PROTEIN FTSY, GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-MONOPHOSPHATE, ... | | Authors: | Focia, P.J, Gawronski-Salerno, J, Coon V, J.S, Freymann, D.M. | | Deposit date: | 2006-05-24 | | Release date: | 2006-10-11 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Structure of a Gdp:Alf(4) Complex of the Srp Gtpases Ffh and Ftsy, and Identification of a Peripheral Nucleotide Interaction Site.
J.Mol.Biol., 360, 2006
|
|
6MJV
 
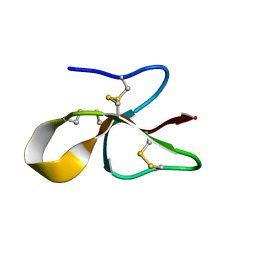 | | A consensus human beta defensin | | Descriptor: | Human beta-defensin | | Authors: | Amero, C, Villegas-Moreno, J, Rodriguez, A, Norton, R.S, Corzo, G. | | Deposit date: | 2018-09-22 | | Release date: | 2019-08-28 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Antimicrobial activity and structure of a consensus human beta-defensin and its comparison to a novel putative hBD10.
Proteins, 88, 2020
|
|
2CDE
 
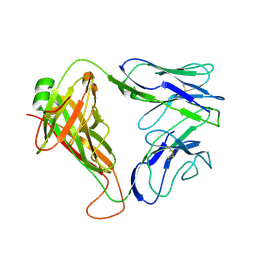 | | Structure and binding kinetics of three different human CD1d-alpha- Galactosylceramide specific T cell receptors - iNKT-TCR | | Descriptor: | INKT-TCR | | Authors: | Gadola, S.D, Koch, M, Marles-Wright, J, Lissin, N.M, Sheperd, D, Matulis, G, Harlos, K, Villiger, P.M, Stuart, D.I, Jakobsen, B.K, Cerundolo, V, Jones, E.Y. | | Deposit date: | 2006-01-23 | | Release date: | 2006-03-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structrue and Binding Kinetics of Three Different Human Cd1D-Alpha-Galactosylceramide-Specific T Cell Receptors
J.Exp.Med., 203, 2006
|
|
2CDY
 
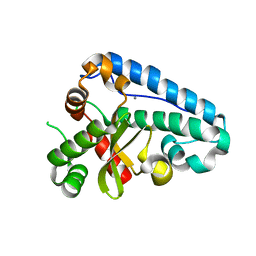 | | Manganese Superoxide Dismutase (Mn-SOD) from Deinococcus radiodurans | | Descriptor: | MANGANESE (II) ION, SUPEROXIDE DISMUTASE [MN] | | Authors: | Dennis, R, Micossi, E, McCarthy, J, Moe, E, Gordon, E, Leonard, G, McSweeney, S. | | Deposit date: | 2006-01-31 | | Release date: | 2006-02-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the manganese superoxide dismutase from Deinococcus radiodurans in two crystal forms.
Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun., 62, 2006
|
|
2Y5H
 
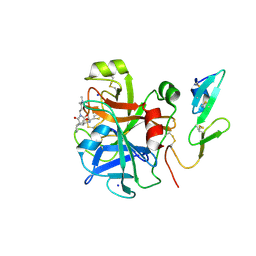 | | FACTOR XA - CATION INHIBITOR COMPLEX | | Descriptor: | 3-[(3AS,4R,5S,8AS,8BR)-4-[2-(5-CHLOROTHIOPHEN-2-YL)-1,3-OXAZOL-4-YL]-1,3-DIOXO-4,6,7,8,8A,8B-HEXAHYDRO-3AH-PYRROLO[3,4-A]PYRROLIZIN-2-YL]PROPYL-TRIMETHYL-AZANIUM, ACTIVATED FACTOR XA HEAVY CHAIN, FACTOR X LIGHT CHAIN, ... | | Authors: | Banner, D.W, Salonen, L.M, Holland, M.C, Haap, W, Benz, J, Diederich, F. | | Deposit date: | 2011-01-13 | | Release date: | 2011-12-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | Molecular Recognition at the Active Site of Factor Xa: Cation-Pi Interactions, Stacking on Planar Peptide Surfaces, and Replacement of Structural Water.
Chemistry, 18, 2012
|
|
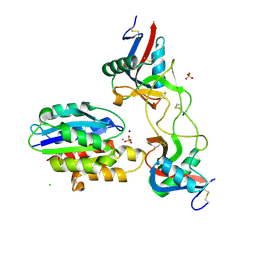
6NDC
 
 | |
2XD7
 
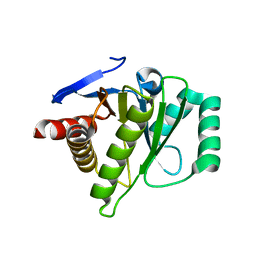 | | Crystal structure of the macro domain of human core histone H2A | | Descriptor: | CORE HISTONE MACRO-H2A.2 | | Authors: | Vollmar, M, Phillips, C, Carpenter, E.P, Muniz, J.R.C, Krojer, T, Ugochukwu, E, von Delft, F, Bountra, C, Arrowsmith, C.H, Weigelt, J, Edwards, A, Gileadi, O. | | Deposit date: | 2010-04-29 | | Release date: | 2010-05-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Crystal Structure of the Macro Domain of Human Core Histone H2A
To be Published
|
|
2XJ9
 
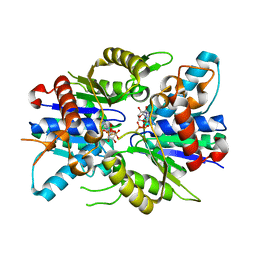 | | Dimer Structure of the bacterial cell division regulator MipZ | | Descriptor: | MAGNESIUM ION, MIPZ, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER | | Authors: | Michie, K.A, Lowe, J. | | Deposit date: | 2010-07-02 | | Release date: | 2011-07-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Localized Dimerization and Nucleoid Binding Drive Gradient Formation by the Bacterial Cell Division Inhibitor Mipz.
Mol.Cell, 46, 2012
|
|
2XIR
 
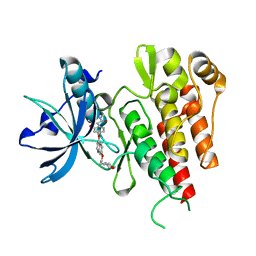 | | Crystal structure of the VEGFR2 kinase domain in complex with PF- 00337210 (N,2-dimethyl-6-(7-(2-morpholinoethoxy)quinolin-4-yloxy) benzofuran-3-carboxamide) | | Descriptor: | N,2-DIMETHYL-6-{[7-(2-MORPHOLIN-4-YLETHOXY)QUINOLIN-4-YL]OXY}-1-BENZOFURAN-3-CARBOXAMIDE, VASCULAR ENDOTHELIAL GROWTH FACTOR RECEPTOR 2 | | Authors: | McTigue, M, Wickersham, J, Pinko, C, Hong, Y, Marrone, T. | | Deposit date: | 2010-06-30 | | Release date: | 2011-04-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Discovery of the Selective Vegfr Inhibitor Pf- 00337210
To be Published
|
|
1NQD
 
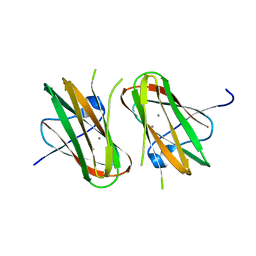 | | CRYSTAL STRUCTURE OF CLOSTRIDIUM HISTOLYTICUM COLG COLLAGENASE COLLAGEN-BINDING DOMAIN 3B AT 1.65 ANGSTROM RESOLUTION IN PRESENCE OF CALCIUM | | Descriptor: | CALCIUM ION, class 1 collagenase | | Authors: | Wilson, J.J, Matsushita, O, Okabe, A, Sakon, J. | | Deposit date: | 2003-01-21 | | Release date: | 2003-04-15 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | A Bacterial Collagen-Binding Domain with Novel Calcium-Binding Motif Controls Domain Orientation
Embo J., 22, 2003
|
|
2CFY
 
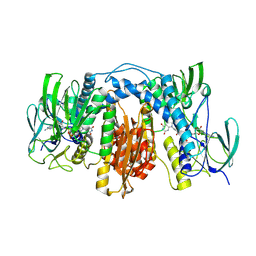 | | Crystal structure of human thioredoxin reductase 1 | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, THIOREDOXIN REDUCTASE 1 | | Authors: | Debreczeni, J.E, Johansson, C, Kavanagh, K, Savitsky, P, Sundstrom, M, Arrowsmith, C, Weigelt, J, Edwards, A, von Delft, F, Oppermann, U. | | Deposit date: | 2006-02-26 | | Release date: | 2006-03-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structure of Human Thioredoxin Reductase 1
To be Published
|
|
6NDF
 
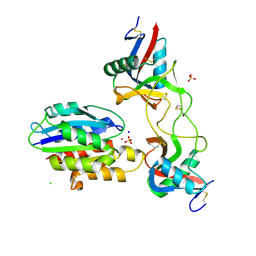 | |
2XMS
 
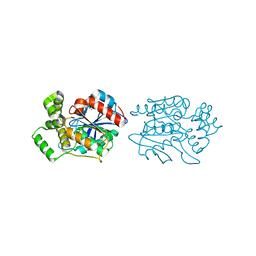 | | Crystal structure of human NDRG2 protein provides insight into its role as a tumor suppressor | | Descriptor: | CHLORIDE ION, IMIDAZOLE, PROTEIN NDRG2 | | Authors: | Hwang, J, Kim, Y, Lee, H, Kim, M.H. | | Deposit date: | 2010-07-29 | | Release date: | 2011-01-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal Structure of Human Ndrg2 Protein Provides Insight Into its Role as a Tumor Suppressor.
J.Biol.Chem., 286, 2011
|
|
2CE0
 
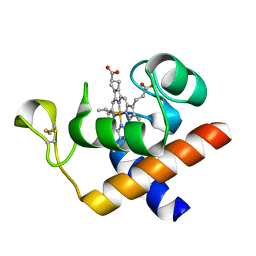 | | Structure of oxidized Arabidopsis thaliana cytochrome 6A | | Descriptor: | CYTOCHROME C6, HEME C | | Authors: | Marcaida, M.J, Schlarb-Ridley, B.G, Worrall, J.A.R, Wastl, J, Evans, T.J, Bendall, D.S, Luisi, B.F, Howe, C.J. | | Deposit date: | 2006-02-01 | | Release date: | 2006-07-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.24 Å) | | Cite: | Structure of Cytochrome C(6A), a Novel Dithio-Cytochrome of Arabidopsis Thaliana, and its Reactivity with Plastocyanin: Implications for Function.
J.Mol.Biol., 360, 2006
|
|
2XO5
 
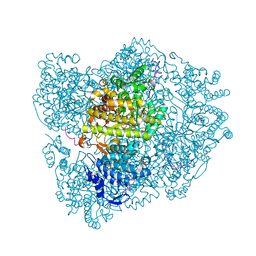 | | RIBONUCLEOTIDE REDUCTASE Y731NH2Y MODIFIED R1 SUBUNIT OF E. COLI | | Descriptor: | RIBONUCLEOSIDE-DIPHOSPHATE REDUCTASE 1 SUBUNIT ALPHA, RIBONUCLEOSIDE-DIPHOSPHATE REDUCTASE 1 SUBUNIT BETA | | Authors: | Minnihan, E.C, Seyedsayamdost, M.R, Uhlin, U, Stubbe, J. | | Deposit date: | 2010-08-09 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Kinetics of Radical Intermediate Formation and Deoxynucleotide Production in 3-Aminotyrosine- Substituted Escherichia Coli Ribonucleotide Reductases.
J.Am.Chem.Soc., 133, 2011
|
|
6NEZ
 
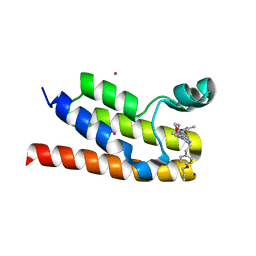 | | Trypanosoma brucei - BDF5, Tb427tmp.01.5000 A, solved with PF-CBP1 | | Descriptor: | 5-(3,5-dimethyl-1,2-oxazol-4-yl)-1-[2-(morpholin-4-yl)ethyl]-2-[2-(4-propoxyphenyl)ethyl]-1H-benzimidazole, UNKNOWN ATOM OR ION, Uncharacterized protein | | Authors: | Lin, Y.H, Dong, A, Tempel, W, McAuley, J, Loppnau, P, Bountra, C, Arrowsmith, C.H, Edwards, A.M, Hui, R, Vedadi, M, Harding, R.J, Structural Genomics Consortium (SGC) | | Deposit date: | 2018-12-18 | | Release date: | 2019-01-23 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Trypanosoma brucei - BDF5, Tb427tmp.01.5000 A, solved with PF-CBP1
to be published
|
|
1I78
 
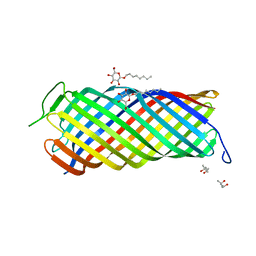 | | CRYSTAL STRUCTURE OF OUTER MEMBRANE PROTEASE OMPT FROM ESCHERICHIA COLI | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, PROTEASE VII, octyl beta-D-glucopyranoside | | Authors: | Vandeputte-Rutten, L, Kramer, R.A, Kroon, J, Dekker, N, Egmond, M.R, Gros, P. | | Deposit date: | 2001-03-08 | | Release date: | 2001-10-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of the outer membrane protease OmpT from Escherichia coli suggests a novel catalytic site.
EMBO J., 20, 2001
|
|
