2V7Y
 
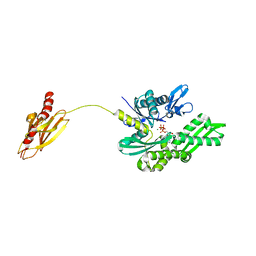 | | Crystal structure of the molecular chaperone DnaK from Geobacillus kaustophilus HTA426 in post-ATP hydrolysis state | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, CHAPERONE PROTEIN DNAK, MAGNESIUM ION, ... | | Authors: | Chang, Y.-W, Sun, Y.-J, Wang, C, Hsiao, C.-D. | | Deposit date: | 2007-08-02 | | Release date: | 2008-04-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Crystal Structures of the 70-kDa Heat Shock Proteins in Domain Disjoining Conformation.
J.Biol.Chem., 283, 2008
|
|
7X4Z
 | |
7X4V
 
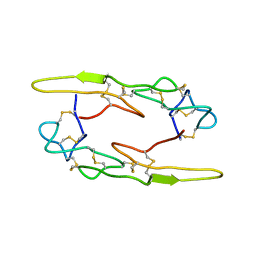 | |
7X4S
 
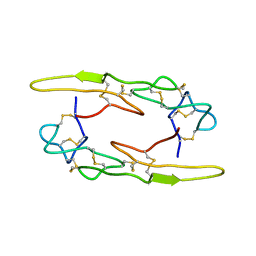 | |
6IPJ
 
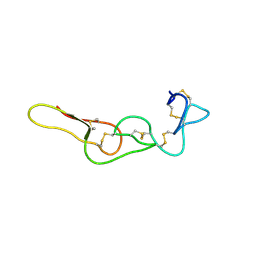
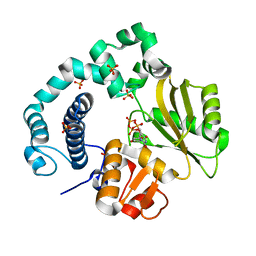 | | Binary Complex of Human DNA Polymerase Mu with MndTTP | | Descriptor: | DNA-directed DNA/RNA polymerase mu,DNA-directed DNA/RNA polymerase mu, MANGANESE (II) ION, SULFATE ION, ... | | Authors: | Chang, Y.K, Wu, W.J, Tsai, M.D. | | Deposit date: | 2018-11-03 | | Release date: | 2019-05-29 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Human DNA Polymerase mu Can Use a Noncanonical Mechanism for Multiple Mn2+-Mediated Functions.
J.Am.Chem.Soc., 141, 2019
|
|
2V7Z
 
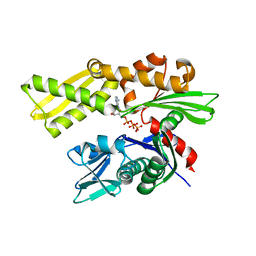 | | Crystal structure of the 70-kDa heat shock cognate protein from Rattus norvegicus in post-ATP hydrolysis state | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, HEAT SHOCK COGNATE 71 KDA PROTEIN, PHOSPHATE ION | | Authors: | Chang, Y.-W, Sun, Y.-J, Wang, C, Hsiao, C.-D. | | Deposit date: | 2007-08-02 | | Release date: | 2008-04-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Crystal Structures of the 70-kDa Heat Shock Proteins in Domain Disjoining Conformation.
J.Biol.Chem., 283, 2008
|
|
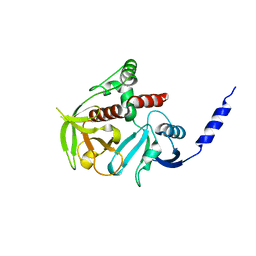
4GMV
 
 | |
4GN1
 
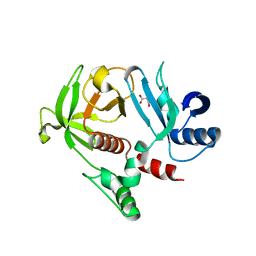 | | Crystal Structure of the RA and PH domains of Lamellipodin | | Descriptor: | MALONATE ION, Ras-associated and pleckstrin homology domains-containing protein 1 | | Authors: | Chang, Y.C.E, Wu, J. | | Deposit date: | 2012-08-16 | | Release date: | 2012-11-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of Lamellipodin implicates diverse functions in actin polymerization and Ras signaling.
Protein Cell, 4, 2013
|
|
5URW
 
 | |
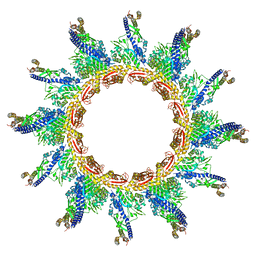
5URX
 | |
2ZUD
 
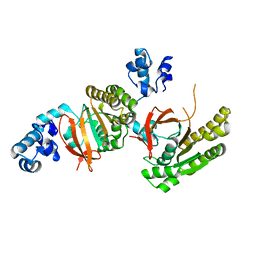 | |
2ZUC
 
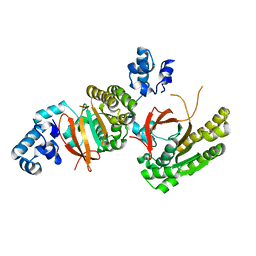 | |
2ZUB
 
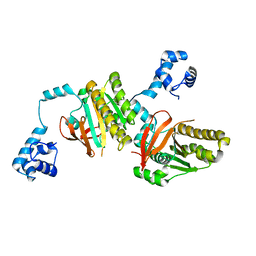 | | Left handed RadA | | Descriptor: | DNA repair and recombination protein radA | | Authors: | Chang, Y.W, Ko, T.P, Wang, T.F, Wang, A.H.J. | | Deposit date: | 2008-10-15 | | Release date: | 2009-04-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Three new structures of left-handed RADA helical filaments: structural flexibility of N-terminal domain is critical for recombinase activity
Plos One, 4, 2009
|
|
3JC9
 
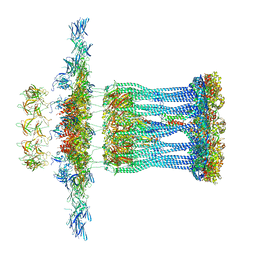 | | Architectural model of the type IVa pilus machine in a non-piliated state | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, PilA, ... | | Authors: | Chang, Y.-W, Rettberg, L.A, Jensen, G.J. | | Deposit date: | 2015-11-24 | | Release date: | 2016-03-16 | | Last modified: | 2018-07-18 | | Method: | ELECTRON MICROSCOPY | | Cite: | Architecture of the type IVa pilus machine.
Science, 351, 2016
|
|
3JC8
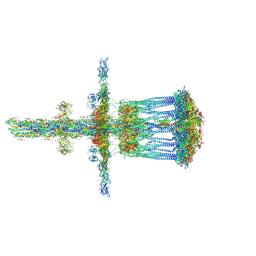 
 | | Architectural model of the type IVa pilus machine in a piliated state | | Descriptor: | LysM domain protein, PilA, PilN, ... | | Authors: | Chang, Y.-W, Rettberg, L.A, Jensen, G.J. | | Deposit date: | 2015-11-24 | | Release date: | 2016-03-16 | | Last modified: | 2018-07-18 | | Method: | ELECTRON MICROSCOPY | | Cite: | Architecture of the type IVa pilus machine.
Science, 351, 2016
|
|
4Z4X
 
 | |
4EJW
 
 | |
4EJU
 
 | |
4EJV
 
 | | Staphylococcus epidermidis TcaR in complex with chloramphenicol | | Descriptor: | 2,2-dichloro-N-[(1S,2S)-1,3-dihydroxy-1-(4-nitrophenyl)propan-2-yl]acetamide, Transcriptional regulator TcaR | | Authors: | Chang, Y.M, Chen, C.K.M, Wang, A.H.J. | | Deposit date: | 2012-04-07 | | Release date: | 2013-05-08 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural analysis of the antibiotic recognition mechanism of MarR family proteins
Acta Crystallogr.,Sect.D, 2013
|
|
4M4C
 
 | | Crystal structure of Rhodostomin ARGDP mutant | | Descriptor: | SULFATE ION, Zinc metalloproteinase/disintegrin | | Authors: | Chang, Y.T, Jeng, W.Y, Shiu, J.H, Chen, C.Y, Chuang, W.J. | | Deposit date: | 2013-08-07 | | Release date: | 2014-08-13 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Effect of C-terminal proline residue adjacent to the RGD motif in rhodostomin on its activity and structure
To be Published
|
|
4GDF
 
 | | A Crystal Structure of SV40 Large T Antigen | | Descriptor: | DNA (32-MER), Large T antigen, ZINC ION | | Authors: | Chang, Y.P, Xu, M, Chen, X.S. | | Deposit date: | 2012-07-31 | | Release date: | 2013-04-10 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Mechanism of Origin DNA Recognition and Assembly of an Initiator-Helicase Complex by SV40 Large Tumor Antigen.
Cell Rep, 3, 2013
|
|
3MO2
 
 | | human G9a-like (GLP, also known as EHMT1) in complex with inhibitor E67 | | Descriptor: | 7-[(5-aminopentyl)oxy]-N~4~-(1-benzylpiperidin-4-yl)-N~2~-[3-(dimethylamino)propyl]-6-methoxyquinazoline-2,4-diamine, Histone-lysine N-methyltransferase, H3 lysine-9 specific 5, ... | | Authors: | Chang, Y, Horton, J.R, Cheng, X. | | Deposit date: | 2010-04-22 | | Release date: | 2010-06-30 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Adding a lysine mimic in the design of potent inhibitors of histone lysine methyltransferases.
J.Mol.Biol., 400, 2010
|
|
3MO5
 
 | | Human G9a-like (GLP, also known as EHMT1) in complex with inhibitor E72 | | Descriptor: | 7-[(5-aminopentyl)oxy]-N~4~-[1-(5-aminopentyl)piperidin-4-yl]-N~2~-[3-(dimethylamino)propyl]-6-methoxyquinazoline-2,4-diamine, Histone-lysine N-methyltransferase, H3 lysine-9 specific 5, ... | | Authors: | Chang, Y, Horton, J.R, Cheng, X. | | Deposit date: | 2010-04-22 | | Release date: | 2010-06-30 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Adding a lysine mimic in the design of potent inhibitors of histone lysine methyltransferases.
J.Mol.Biol., 400, 2010
|
|
7XAO
 
 | | Crystal structure of thioredoxin 1 | | Descriptor: | Thioredoxin | | Authors: | Chang, Y.J, Park, H.H. | | Deposit date: | 2022-03-18 | | Release date: | 2022-06-01 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | High-resolution crystal structure of Acinetobacter baumannii thioredoxin 1.
Biochem.Biophys.Res.Commun., 608, 2022
|
|
7X44
 
 | |
