8ODT
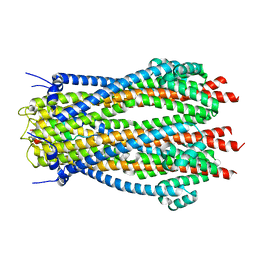 
 | | Structure of TolQR complex from E.coli | | Descriptor: | Tol-Pal system protein TolQ, Tol-Pal system protein TolR | | Authors: | Webby, M.N, Kleanthous, C, Press, C.E. | | Deposit date: | 2023-03-09 | | Release date: | 2023-11-01 | | Last modified: | 2023-11-29 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Tunable force transduction through the Escherichia coli cell envelope.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
1HK5
 
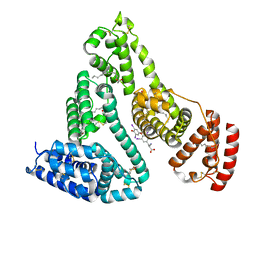 | | HUMAN SERUM ALBUMIN MUTANT R218H COMPLEXED WITH THYROXINE (3,3',5,5'-TETRAIODO-L-THYRONINE) and myristic acid (tetradecanoic acid) | | Descriptor: | 3,5,3',5'-TETRAIODO-L-THYRONINE, MYRISTIC ACID, SERUM ALBUMIN | | Authors: | Petitpas, I, Petersen, C.E, Ha, C.E, Bhattacharya, A.A, Zunszain, P.A, Ghuman, J, Bhagavan, N.V, Curry, S. | | Deposit date: | 2003-03-05 | | Release date: | 2003-05-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural Basis of Albumin-Thyroxine Interactions and Familial Dysalbuminemic Hyperthyroxinemia
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1HK3
 
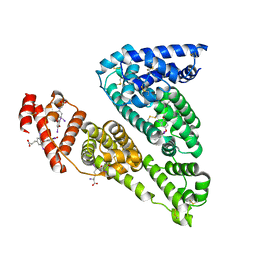 | | Human serum albumin mutant r218p complexed with thyroxine (3,3',5,5'-tetraiodo-l-thyronine) | | Descriptor: | 3,5,3',5'-TETRAIODO-L-THYRONINE, SERUM ALBUMIN | | Authors: | Petitpas, I, Petersen, C.E, Ha, C.E, Bhattacharya, A.A, Zunszain, P.A, Ghuman, J, Bhagavan, N.V, Curry, S. | | Deposit date: | 2003-03-05 | | Release date: | 2003-05-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural Basis of Albumin-Thyroxine Interactions and Familial Dysalbuminemic Hyperthyroxinemia
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1HK4
 
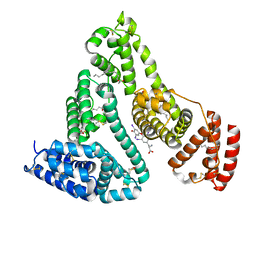 | | HUMAN SERUM ALBUMIN COMPLEXED WITH THYROXINE (3,3',5,5'-TETRAIODO-L-THYRONINE) and myristic acid (tetradecanoic acid) | | Descriptor: | 3,5,3',5'-TETRAIODO-L-THYRONINE, MYRISTIC ACID, SERUM ALBUMIN | | Authors: | Petitpas, I, Petersen, C.E, Ha, C.E, Bhattacharya, A.A, Zunszain, P.A, Ghuman, J, Bhagavan, N.V, Curry, S. | | Deposit date: | 2003-03-05 | | Release date: | 2003-05-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural Basis of Albumin-Thyroxine Interactions and Familial Dysalbuminemic Hyperthyroxinemia
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1HK2
 
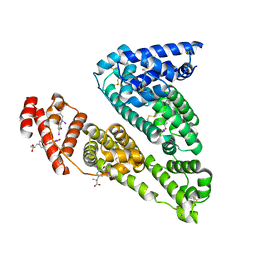 | | HUMAN SERUM ALBUMIN MUTANT R218H COMPLEXED WITH THYROXINE (3,3',5,5'-TETRAIODO-L-THYRONINE) | | Descriptor: | 3,5,3',5'-TETRAIODO-L-THYRONINE, SERUM ALBUMIN | | Authors: | Petitpas, I, Petersen, C.E, Ha, C.E, Bhattacharya, A.A, Zunszain, P.A, Ghuman, J, Bhagavan, N.V, Curry, S. | | Deposit date: | 2003-03-05 | | Release date: | 2003-05-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural Basis of Albumin-Thyroxine Interactions and Familial Dysalbuminemic Hyperthyroxinemia
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1HK1
 
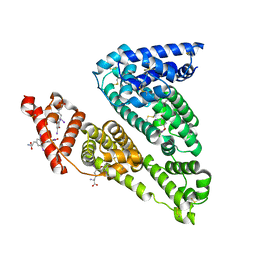 | | HUMAN SERUM ALBUMIN COMPLEXED WITH THYROXINE (3,3',5,5'-TETRAIODO-L-THYRONINE) | | Descriptor: | 3,5,3',5'-TETRAIODO-L-THYRONINE, SERUM ALBUMIN | | Authors: | Petitpas, I, Petersen, C.E, Ha, C.E, Bhattacharya, A.A, Zunszain, P.A, Ghuman, J, Bhagavan, N.V, Curry, S. | | Deposit date: | 2003-03-05 | | Release date: | 2003-05-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structural Basis of Albumin-Thyroxine Interactions and Familial Dysalbuminemic Hyperthyroxinemia
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
4W60
 
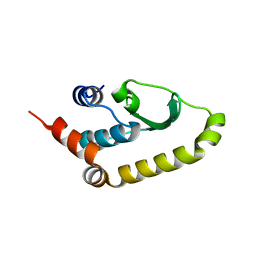 | | The structure of Vaccina virus H7 protein displays A Novel Phosphoinositide binding fold required for membrane biogenesis | | Descriptor: | Late protein H7 | | Authors: | Kolli, S, Meng, X, Wu, X, Shengjuler, D, Cameron, C.E, Xiang, Y, Deng, J. | | Deposit date: | 2014-08-19 | | Release date: | 2014-12-31 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure-function analysis of vaccinia virus h7 protein reveals a novel phosphoinositide binding fold essential for poxvirus replication.
J.Virol., 89, 2015
|
|
4W5U
 
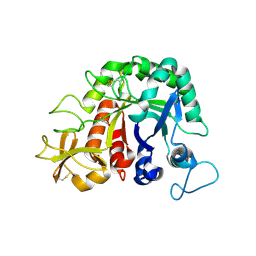 | |
6RB9
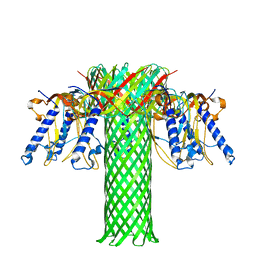 
 | | The pore structure of Clostridium perfringens epsilon toxin | | Descriptor: | Epsilon-toxin type B | | Authors: | Savva, C.G, Clark, A.R, Naylor, C.E, Popoff, M.R, Moss, D.S, Basak, A.K, Titball, R.W, Bokori-Brown, M. | | Deposit date: | 2019-04-09 | | Release date: | 2019-06-19 | | Last modified: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The pore structure of Clostridium perfringens epsilon toxin.
Nat Commun, 10, 2019
|
|
8B3M
 
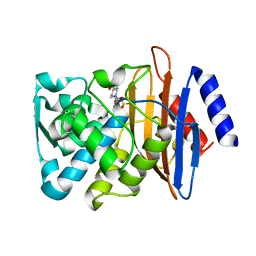 | | Millisecond cryo-trapping by the spitrobot crystal plunger, CTXM-14 Avibactam complex, SSX, 1 sec | | Descriptor: | (2S,5R)-1-formyl-5-[(sulfooxy)amino]piperidine-2-carboxamide, Beta-lactamase | | Authors: | Mehrabi, P, Sung, S, von Stetten, D, Prester, A, Hatton, C.E, Kleine-Doepke, S, Berkes, A, Gore, G, Leimkohl, J.P, Schikora, H, Kollewe, M, Rohde, H, Wilmanns, M, Tellkamp, F, Schulz, E.C. | | Deposit date: | 2022-09-16 | | Release date: | 2023-05-24 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Millisecond cryo-trapping by the spitrobot crystal plunger simplifies time-resolved crystallography.
Nat Commun, 14, 2023
|
|
8B56
 
 | | Crystal structure of SARS-CoV-2 main protease (MPro) in complex with the inhibitor GD-9 | | Descriptor: | (2~{S})-4-(2-chloranylethanoyl)-1-(3,4-dichlorophenyl)-~{N}-(thiophen-2-ylmethyl)piperazine-2-carboxamide, 3C-like proteinase nsp5, BROMIDE ION, ... | | Authors: | Straeter, N, Muller, C.E, Claff, T, Sylvester, K, Weisse, R, Gao, S, Song, L, Liu, X, Zhan, P. | | Deposit date: | 2022-09-21 | | Release date: | 2023-08-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.823 Å) | | Cite: | Discovery and Crystallographic Studies of Nonpeptidic Piperazine Derivatives as Covalent SARS-CoV-2 Main Protease Inhibitors.
J.Med.Chem., 65, 2022
|
|
4F2E
 
 | |
4F2F
 
 | | Crystal structure of the metal binding domain (MBD) of the Streptococcus pneumoniae D39 Cu(I) exporting P-type ATPase CopA with Cu(I) | | Descriptor: | CHLORIDE ION, COPPER (I) ION, Cation-transporting ATPase, ... | | Authors: | Fu, Y, Dann III, C.E, Giedroc, D.P. | | Deposit date: | 2012-05-07 | | Release date: | 2013-01-30 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | A new structural paradigm in copper resistance in Streptococcus pneumoniae.
Nat.Chem.Biol., 9, 2013
|
|
6DV3
 
 | | Structure of the Salmonella SPI-1 type III secretion injectisome secretin InvG in the open gate state | | Descriptor: | Protein InvG | | Authors: | Hu, J, Worrall, L.J, Vuckovic, M, Atkinson, C.E, Strynadka, N.C.J. | | Deposit date: | 2018-06-22 | | Release date: | 2018-10-03 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Cryo-EM analysis of the T3S injectisome reveals the structure of the needle and open secretin.
Nat Commun, 9, 2018
|
|
6DV6
 | | Structure of the Salmonella SPI-1 type III secretion injectisome secretin InvG (residues 176-end) in the open gate state | | Descriptor: | Protein InvG | | Authors: | Hu, J, Worrall, L.J, Vuckovic, M, Atkinson, C.E, Strynadka, N.C.J. | | Deposit date: | 2018-06-22 | | Release date: | 2018-10-03 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Cryo-EM analysis of the T3S injectisome reveals the structure of the needle and open secretin.
Nat Commun, 9, 2018
|
|
6DUZ
 | | Structure of the periplasmic domains of PrgH and PrgK from the assembled Salmonella type III secretion injectisome needle complex | | Descriptor: | Lipoprotein PrgK, Protein PrgH | | Authors: | Hu, J, Worrall, L.J, Vuckovic, M, Atkinson, C.E, Strynadka, N.C.J. | | Deposit date: | 2018-06-22 | | Release date: | 2018-10-03 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Cryo-EM analysis of the T3S injectisome reveals the structure of the needle and open secretin.
Nat Commun, 9, 2018
|
|
3LK9
 
 | | DNA polymerase beta with a gapped DNA substrate and dTMP(CF2)P(CF2)P | | Descriptor: | 5'-O-[(R)-{[(R)-[difluoro(phosphono)methyl](hydroxy)phosphoryl](difluoro)methyl}(hydroxy)phosphoryl]thymidine, CHLORIDE ION, DNA (5'-D(*CP*CP*GP*AP*CP*AP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3'), ... | | Authors: | Zibinsky, M, Surya Prakash, G.K, Upton, T.G, Kashemirov, B.A, McKenna, C.E, Oertell, K, Goodman, M.F, Batra, V.K, Pedersen, L.C, Beard, W.A, Wilson, S.H. | | Deposit date: | 2010-01-27 | | Release date: | 2011-01-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Synthesis and biological evaluation of fluorinated deoxynucleotide analogs based on bis-(difluoromethylene)triphosphoric acid.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2FQT
 
 | | Crystal structure of B.subtilis LuxS in complex with (2S)-2-Amino-4-[(2R,3S)-2,3-dihydroxy-3-N-hydroxycarbamoyl-propylmercapto]butyric acid | | Descriptor: | (2S)-2-AMINO-4-[(2R,3S)-2,3-DIHYDROXY-3-N-HYDROXYCARBAMOYL-PROPYLMERCAPTO]BUTYRIC ACID, COBALT (II) ION, S-ribosylhomocysteine lyase, ... | | Authors: | Shen, G, Rajan, R, Zhu, J, Bell, C.E, Pei, D. | | Deposit date: | 2006-01-18 | | Release date: | 2006-05-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Design and Synthesis of Substrate Analogue Inhibitors of S-Ribosylhomocysteinase (LuxS)
J.Med.Chem., 49, 2006
|
|
2FYR
 
 | |
2FYQ
 
 | |
2GEM
 
 | |
4V97
 
 | | Crystal structure of the bacterial ribosome ram mutation G299A. | | Descriptor: | 16S rRNA, 23S rRNA, 30S ribosomal protein S10, ... | | Authors: | Fagan, C.E, Dunkle, J.A, Maehigashi, T, Dunham, C.M. | | Deposit date: | 2012-04-06 | | Release date: | 2014-07-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.516 Å) | | Cite: | Reorganization of an intersubunit bridge induced by disparate 16S ribosomal ambiguity mutations mimics an EF-Tu-bound state.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4F4L
 
 | | Open Channel Conformation of a Voltage Gated Sodium Channel | | Descriptor: | Ion transport protein | | Authors: | McCusker, E.C, Bagneris, C, Naylor, C.E, Cole, A.R, D'Avanzo, N, Nichols, C.G, Wallace, B.A. | | Deposit date: | 2012-05-10 | | Release date: | 2012-10-03 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.49 Å) | | Cite: | Structure of a bacterial voltage-gated sodium channel pore reveals mechanisms of opening and closing.
Nat Commun, 3, 2012
|
|
6IEJ
 
 | | The C2 domain of cytosolic phospholipase A2 alpha bound to phosphatidylcholine | | Descriptor: | 1,2-dihexanoyl-sn-glycero-3-phosphocholine, CALCIUM ION, Cytosolic phospholipase A2, ... | | Authors: | Hirano, Y, Gao, Y.G, Stephenson, D.J, Vu, N.T, Malinina, L, Chalfant, C.E, Patel, D.J, Brown, R.E. | | Deposit date: | 2018-09-14 | | Release date: | 2019-05-22 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.206 Å) | | Cite: | Structural basis of phosphatidylcholine recognition by the C2-domain of cytosolic phospholipase A2alpha.
Elife, 8, 2019
|
|
4V8J
 
 | | Crystal structure of the bacterial ribosome ram mutation G347U. | | Descriptor: | 16S rRNA, 23S rRNA, 30S ribosomal protein S10, ... | | Authors: | Fagan, C.E, Dunkle, J.A, Maehigashi, T, Dunham, C.M. | | Deposit date: | 2011-12-20 | | Release date: | 2014-07-09 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (3.9 Å) | | Cite: | Reorganization of an intersubunit bridge induced by disparate 16S ribosomal ambiguity mutations mimics an EF-Tu-bound state.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
