8G1M
 
 | |
8G1L
 
 | |
4B62
 
 | | The structure of the cell wall anchor of the T6SS from Pseudomonas aeruginosa | | Descriptor: | 1,2-ETHANEDIOL, PHOSPHATE ION, TSSL1 | | Authors: | Robb, C.S, Carlson, M, Nano, F.E, Boraston, A.B. | | Deposit date: | 2012-08-08 | | Release date: | 2013-10-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Crystal Structure of the Periplasmic Peptidoglycan Binding Anchor of a T6Ss from Pseudomonas Aeruginosa.
To be Published
|
|
8DF2
 
 | |
8DEK
 
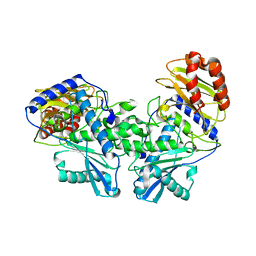 | |
4B28
 
 | | Crystal structure of DMSP lyase RdDddP from Roseobacter denitrificans | | Descriptor: | FE (III) ION, METALLOPEPTIDASE, FAMILY M24, ... | | Authors: | Hehemann, J.H, Law, A, Redecke, L, Boraston, A.B. | | Deposit date: | 2012-07-12 | | Release date: | 2012-07-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | The Structure of Rddddp from Roseobacter Denitrificans Reveals that Dmsp Lyases in the Dddp-Family are Metalloenzymes.
Plos One, 9, 2014
|
|
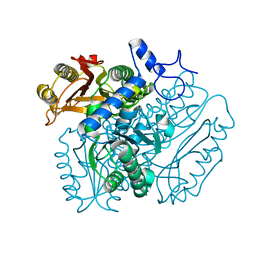
4B29
 
 | | Crystal structures of DMSP lyases RdDddP and RnDddQII | | Descriptor: | 1,2-ETHANEDIOL, BROMIDE ION, DIMETHYLSULFONIOPROPIONATE LYASE, ... | | Authors: | Hehemann, J.H, Law, A, Redecke, L, Boraston, A.B. | | Deposit date: | 2012-07-12 | | Release date: | 2012-07-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Crystal Structures of Dmsp Lyases Rddddp and Rndddqii
To be Published
|
|
4AFT
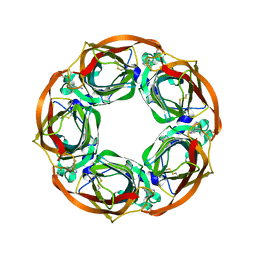 
 | | Aplysia californica AChBP in complex with Varenicline | | Descriptor: | SOLUBLE ACETYLCHOLINE RECEPTOR, VARENICLINE | | Authors: | Rucktooa, P, Haseler, C.A, vanElke, R, Smit, A.B, Gallagher, T, Sixma, T.K. | | Deposit date: | 2012-01-23 | | Release date: | 2012-05-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural Characterization of Binding Mode of Smoking Cessation Drugs to Nicotinic Acetylcholine Receptors Through Study of Ligand Complexes with Acetylcholine-Binding Protein.
J.Biol.Chem., 287, 2012
|
|
4BQT
 
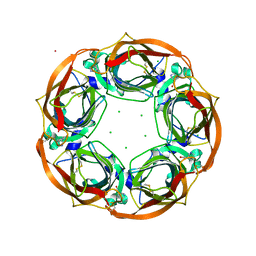 | | Aplysia californica AChBP in complex with Cytisine | | Descriptor: | (1R,5S)-1,2,3,4,5,6-HEXAHYDRO-8H-1,5-METHANOPYRIDO[1,2-A][1,5]DIAZOCIN-8-ONE, CHLORIDE ION, COBALT (II) ION, ... | | Authors: | Rucktooa, P, Haseler, C.A, vanElke, R, Smit, A.B, Gallagher, T, Sixma, T.K. | | Deposit date: | 2013-06-02 | | Release date: | 2013-06-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.88 Å) | | Cite: | Structural Characterization of Binding Mode of Smoking Cessation Drugs to Nicotinic Acetylcholine Receptors Through Study of Ligand Complexes with Acetylcholine-Binding Protein.
J.Biol.Chem., 287, 2012
|
|
8GA8
 
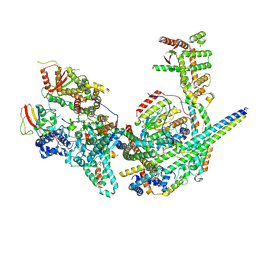 | | Structure of the yeast (HDAC) Rpd3L complex | | Descriptor: | Histone deacetylase RPD3, Transcriptional regulatory protein DEP1, Transcriptional regulatory protein PHO23, ... | | Authors: | Patel, A.B, Radhakrishnan, I, He, Y. | | Deposit date: | 2023-02-22 | | Release date: | 2023-07-12 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Cryo-EM structure of the Saccharomyces cerevisiae Rpd3L histone deacetylase complex.
Nat Commun, 14, 2023
|
|
8D0Z
 
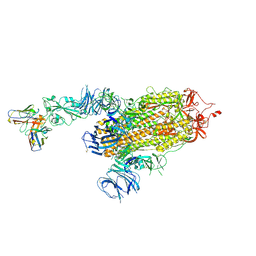 | | S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (focused refinement) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, S728-1157 Fab heavy chain variable region, S728-1157 Fab light chain variable region, ... | | Authors: | Ozorowski, G, Torres, J.L, Ward, A.B. | | Deposit date: | 2022-05-26 | | Release date: | 2023-03-22 | | Last modified: | 2023-05-03 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Site of vulnerability on SARS-CoV-2 spike induces broadly protective antibody against antigenically distinct Omicron subvariants.
J.Clin.Invest., 133, 2023
|
|
8GJE
 
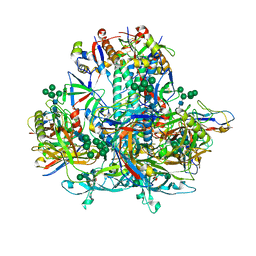 | | HIV-1 Env subtype C CZA97.12 SOSIP.664 in complex with 3BNC117 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3BNC117 Fab heavy chain, ... | | Authors: | Ozorowski, G, Lee, J.H, Ward, A.B. | | Deposit date: | 2023-03-15 | | Release date: | 2023-10-18 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Glycan heterogeneity as a cause of the persistent fraction in HIV-1 neutralization.
Plos Pathog., 19, 2023
|
|
4BMQ
 
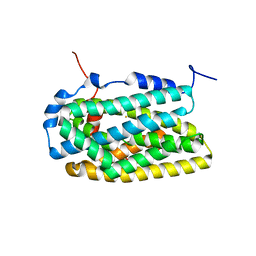 | | Crystal Structure of Ribonucleotide Reductase apo-NrdF from Bacillus cereus (space group C2) | | Descriptor: | FE (II) ION, RIBONUCLEOSIDE-DIPHOSPHATE REDUCTASE SUBUNIT BETA | | Authors: | Tomter, A.B, Hersleth, H.-P, Hammerstad, M, Rohr, A.K, Andersson, K.K. | | Deposit date: | 2013-05-10 | | Release date: | 2014-03-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of Bacillus Cereus Class Ib Ribonucleotide Reductase Di-Iron Nrdf in Complex with Nrdi.
Acs Chem.Biol., 9, 2014
|
|
4BQ3
 
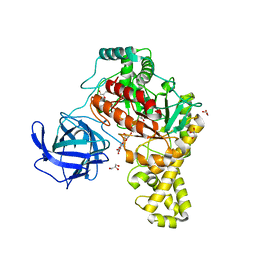 | | Structural analysis of an exo-beta-agarase | | Descriptor: | B-AGARASE, CALCIUM ION, GLYCEROL, ... | | Authors: | Pluvinage, B, Hehemann, J.H, Boraston, A.B. | | Deposit date: | 2013-05-29 | | Release date: | 2013-08-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Substrate Recognition and Hydrolysis by a Family 50 Exo-Beta-Agarase Aga50D from the Marine Bacterium Saccharophagus Degradans
J.Biol.Chem., 288, 2013
|
|
8DC0
 
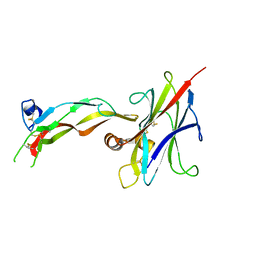 | |
4C21
 
 | | L-Fucose Isomerase In Complex With Fucitol | | Descriptor: | 1,2-ETHANEDIOL, FUCITOL, L-FUCOSE ISOMERASE, ... | | Authors: | Higgins, M.A, Suits, M.D.L, Marsters, C, Boraston, A.B. | | Deposit date: | 2013-08-16 | | Release date: | 2013-12-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structural and Functional Analysis of Fucose-Processing Enzymes from Streptococcus Pneumoniae.
J.Mol.Biol., 426, 2014
|
|
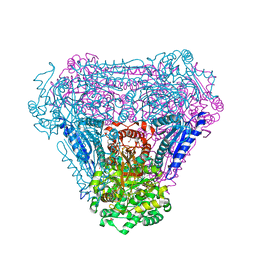
4BMT
 
 | | Crystal Structure of Ribonucleotide Reductase di-iron NrdF from Bacillus cereus | | Descriptor: | FE (II) ION, RIBONUCLEOSIDE-DIPHOSPHATE REDUCTASE SUBUNIT BETA | | Authors: | Hersleth, H.-P, Tomter, A.B, Hammerstad, M, Rohr, A.K, Andersson, K.K. | | Deposit date: | 2013-05-10 | | Release date: | 2014-03-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structure of Bacillus Cereus Class Ib Ribonucleotide Reductase Di-Iron Nrdf in Complex with Nrdi.
Acs Chem.Biol., 9, 2014
|
|
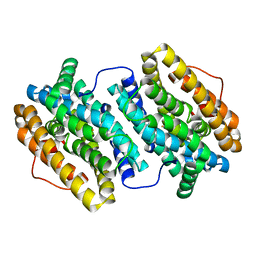
4C23
 
 | | L-fuculose kinase | | Descriptor: | 1,2-ETHANEDIOL, L-FUCULOSE KINASE FUCK | | Authors: | Higgins, M.A, Suits, M.D.L, Marsters, C, Boraston, A.B. | | Deposit date: | 2013-08-16 | | Release date: | 2013-12-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Functional Analysis of Fucose-Processing Enzymes from Streptococcus Pneumoniae.
J.Mol.Biol., 426, 2014
|
|
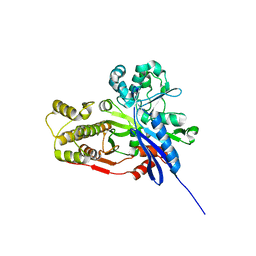
4BMU
 
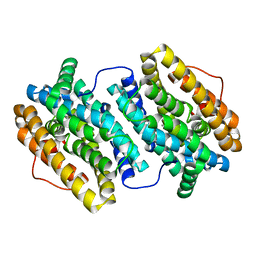 | | Crystal Structure of Ribonucleotide Reductase di-manganese(II) NrdF from Bacillus cereus | | Descriptor: | MANGANESE (II) ION, RIBONUCLEOSIDE-DIPHOSPHATE REDUCTASE SUBUNIT BETA | | Authors: | Hersleth, H.-P, Tomter, A.B, Hammerstad, M, Rohr, A.K, Andersson, K.K. | | Deposit date: | 2013-05-10 | | Release date: | 2014-03-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of Bacillus Cereus Class Ib Ribonucleotide Reductase Di-Iron Nrdf in Complex with Nrdi.
Acs Chem.Biol., 9, 2014
|
|
4C24
 
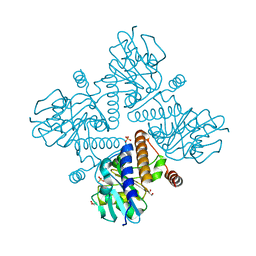 | | L-fuculose 1-phosphate aldolase | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, L-FUCULOSE PHOSPHATE ALDOLASE, ... | | Authors: | Higgins, M.A, Suits, M.D.L, Marsters, C, Boraston, A.B. | | Deposit date: | 2013-08-16 | | Release date: | 2013-12-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural and Functional Analysis of Fucose-Processing Enzymes from Streptococcus Pneumoniae.
J.Mol.Biol., 426, 2014
|
|
4AWD
 
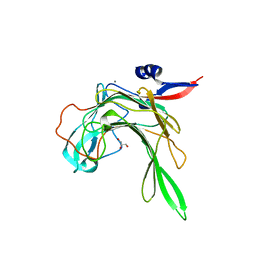 | | Crystal structure of the beta-porphyranase BpGH16B (BACPLE_01689) from the human gut bacterium Bacteroides plebeius | | Descriptor: | BETA-PORPHYRANASE, CALCIUM ION, GLYCEROL | | Authors: | Hehemann, J.H, Kelly, A.G, Pudlo, N.A, Martens, E.C, Boraston, A.B. | | Deposit date: | 2012-06-01 | | Release date: | 2012-07-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Bacteria of the human gut microbiome catabolize red seaweed glycans with carbohydrate-active enzyme updates from extrinsic microbes.
Proc. Natl. Acad. Sci. U.S.A., 109, 2012
|
|
4CC5
 
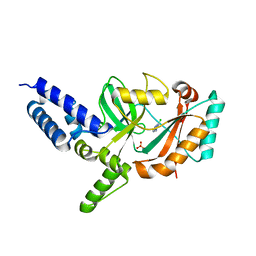 | | Fragment-Based Discovery of 6 Azaindazoles As Inhibitors of Bacterial DNA Ligase | | Descriptor: | 2-chloranyl-6-(1H-1,2,4-triazol-3-yl)pyrazine, DNA LIGASE, SULFATE ION | | Authors: | Howard, S, Amin, N, Benowitz, A.B, Chiarparin, E, Cui, H, Deng, X, Heightman, T.D, Holmes, D.J, Hopkins, A, Huang, J, Jin, Q, Kreatsoulas, C, Martin, A.C.L, Massey, F, McCloskey, L, Mortenson, P.N, Pathuri, P, Tisi, D, Williams, P.A. | | Deposit date: | 2013-10-18 | | Release date: | 2014-06-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Fragment-Based Discovery of 6-Azaindazoles as Inhibitors of Bacterial DNA Ligase.
Acs Med.Chem.Lett., 4, 2013
|
|
4C18
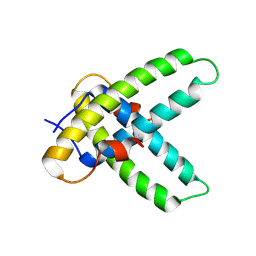 
 | |
8EAQ
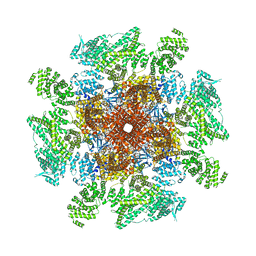 
 | | Structure of the full-length IP3R1 channel determined at high Ca2+ | | Descriptor: | (9R,11S)-9-({[(1S)-1-HYDROXYHEXADECYL]OXY}METHYL)-2,2-DIMETHYL-5,7,10-TRIOXA-2LAMBDA~5~-AZA-6LAMBDA~5~-PHOSPHAOCTACOSANE-6,6,11-TRIOL, CALCIUM ION, Inositol 1,4,5-trisphosphate receptor type 1, ... | | Authors: | Fan, G, Baker, M.R, Terry, L.E, Arige, V, Chen, M, Seryshev, A.B, Baker, M.L, Ludtke, S.J, Yule, D.I, Serysheva, I.I. | | Deposit date: | 2022-08-29 | | Release date: | 2022-11-23 | | Method: | ELECTRON MICROSCOPY (3.26 Å) | | Cite: | Conformational motions and ligand-binding underlying gating and regulation in IP 3 R channel.
Nat Commun, 13, 2022
|
|
8EAR
 
 | | Structure of the full-length IP3R1 channel determined in the presence of Calcium/IP3/ATP | | Descriptor: | (9R,11S)-9-({[(1S)-1-HYDROXYHEXADECYL]OXY}METHYL)-2,2-DIMETHYL-5,7,10-TRIOXA-2LAMBDA~5~-AZA-6LAMBDA~5~-PHOSPHAOCTACOSANE-6,6,11-TRIOL, ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, ... | | Authors: | Fan, G, Baker, M.R, Terry, L.E, Arige, V, Chen, M, Seryshev, A.B, Baker, M.L, Ludtke, S.J, Yule, D.I, Serysheva, I.I. | | Deposit date: | 2022-08-29 | | Release date: | 2022-11-23 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Conformational motions and ligand-binding underlying gating and regulation in IP 3 R channel.
Nat Commun, 13, 2022
|
|
