1FRP
 
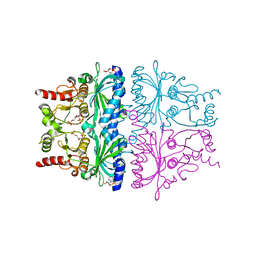 | | CRYSTAL STRUCTURE OF FRUCTOSE-1,6-BISPHOSPHATASE COMPLEXED WITH FRUCTOSE-2,6-BISPHOSPHATE, AMP AND ZN2+ AT 2.0 ANGSTROMS RESOLUTION. ASPECTS OF SYNERGISM BETWEEN INHIBITORS | | Descriptor: | 2,6-di-O-phosphono-beta-D-fructofuranose, ADENOSINE MONOPHOSPHATE, FRUCTOSE 1,6-BISPHOSPHATASE, ... | | Authors: | Xue, Y, Huang, S, Liang, J.-Y, Zhang, Y, Lipscomb, W.N. | | Deposit date: | 1994-08-26 | | Release date: | 1994-11-30 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of fructose-1,6-bisphosphatase complexed with fructose 2,6-bisphosphate, AMP, and Zn2+ at 2.0-A resolution: aspects of synergism between inhibitors.
Proc.Natl.Acad.Sci.USA, 91, 1994
|
|
4H1D
 
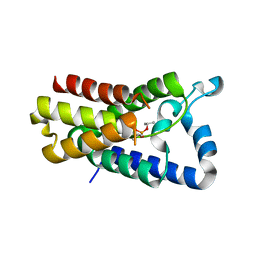 | | Cocrystal structure of GlpG and DFP | | Descriptor: | DIISOPROPYL PHOSPHONATE, Rhomboid protease GlpG | | Authors: | Xue, Y, Ha, Y. | | Deposit date: | 2012-09-10 | | Release date: | 2013-05-01 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.8975 Å) | | Cite: | Large lateral movement of transmembrane helix s5 is not required for substrate access to the active site of rhomboid intramembrane protease.
J.Biol.Chem., 288, 2013
|
|
2VB2
 
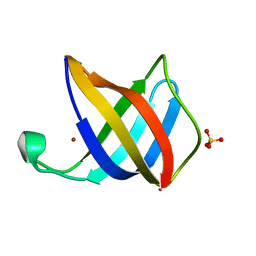 | | Crystal structure of Cu(I)CusF | | Descriptor: | CATION EFFLUX SYSTEM PROTEIN CUSF, COPPER (II) ION, SULFATE ION | | Authors: | Xue, Y, Davis, A.V, Balakrishnan, G, Stasser, J.P, Staehlin, B.M, Focia, P, Spiro, T.G, Penner-Hahn, J.E, O'Halloran, T.V. | | Deposit date: | 2007-09-06 | | Release date: | 2007-12-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Cu(I) Recognition Via Cation-Pi and Methionine Interactions in Cusf.
Nat.Chem.Biol., 4, 2008
|
|
2VB3
 
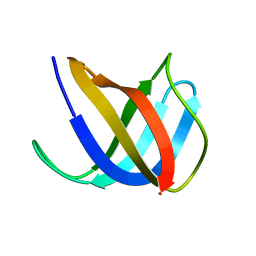 | | Crystal structure of Ag(I)CusF | | Descriptor: | CATION EFFLUX SYSTEM PROTEIN CUSF, SILVER ION | | Authors: | Xue, Y, Davis, A.V, Balakrishnan, G, Stasser, J.P, Staehlin, B.M, Focia, P, Spiro, T.G, Penner-Hahn, J.E, O'Halloran, T.V. | | Deposit date: | 2007-09-06 | | Release date: | 2007-12-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Cu(I) Recognition Via Cation-Pi and Methionine Interactions in Cusf.
Nat.Chem.Biol., 4, 2008
|
|
8YK7
 
 | | Structure of Rib domain from surface adhesin of Limosilactobacillus reuteri | | Descriptor: | 1,2-ETHANEDIOL, COPPER (II) ION, SODIUM ION, ... | | Authors: | Xue, Y, Kang, X. | | Deposit date: | 2024-03-04 | | Release date: | 2024-09-11 | | Last modified: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal structure of the Rib domain of the cell-wall-anchored surface protein from Limosilactobacillus reuteri.
Acta Crystallogr.,Sect.F, 80, 2024
|
|
8YKE
 | |
1CSM
 
 | |
3UBB
 | |
9AVK
 
 | | Structure of long Rib domain from Limosilactobacillus reuteri | | Descriptor: | SODIUM ION, THIOCYANATE ION, YSIRK signal domain/LPXTG anchor domain surface protein | | Authors: | Xue, Y, Kang, X. | | Deposit date: | 2024-03-04 | | Release date: | 2024-05-08 | | Last modified: | 2024-06-12 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | Crystal structure of the long Rib domain of the LPXTG-anchored surface protein from Limosilactobacillus reuteri.
Acta Crystallogr.,Sect.F, 80, 2024
|
|
3Q7G
 
 | | Crystal Structure of E2 domain of Human Amyloid Precursor-Like Protein 1 in complex with SOS (sucrose octasulfate) | | Descriptor: | 1,3,4,6-tetra-O-sulfo-beta-D-fructofuranose-(2-1)-2,3,4,6-tetra-O-sulfonato-alpha-D-glucopyranose, Amyloid-like protein 1 | | Authors: | Xue, Y, Ha, Y. | | Deposit date: | 2011-01-04 | | Release date: | 2011-06-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of the E2 Domain of Amyloid Precursor Protein-like Protein 1 in Complex with Sucrose Octasulfate.
J.Biol.Chem., 286, 2011
|
|
3Q7L
 
 | |
3TXT
 
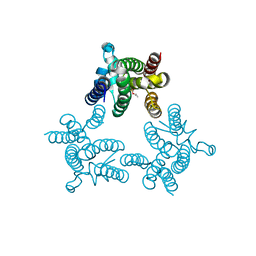 | | Crystal structure of GlpG in complex with inhibitor DFP | | Descriptor: | DIISOPROPYL PHOSPHONATE, Rhomboid protease glpG | | Authors: | Xue, Y, Ha, Y. | | Deposit date: | 2011-09-23 | | Release date: | 2011-12-07 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The crystal structure of rhomboid protease GlpG in complex with a mechanism-based inhibitor
To be Published
|
|
3QMK
 
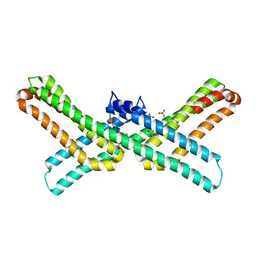 | | Crystal structure of the E2 domain of APLP1 in complex with heparin hexasaccharide | | Descriptor: | 2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose, Amyloid-like protein 1, SULFATE ION | | Authors: | Xue, Y, Ha, Y. | | Deposit date: | 2011-02-04 | | Release date: | 2011-10-12 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | Crystal structure of amyloid precursor-like protein 1 and heparin complex suggests a dual role of heparin in E2 dimerization.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
2AFV
 
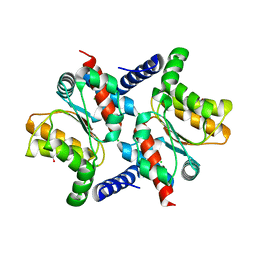 | | The Crystal Structure of Putative Precorrin Isomerase CbiC in Cobalamin Biosynthesis | | Descriptor: | 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, CHLORIDE ION, HEXAETHYLENE GLYCOL, ... | | Authors: | Xue, Y, Wei, Z. | | Deposit date: | 2005-07-26 | | Release date: | 2006-04-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The crystal structure of putative precorrin isomerase CbiC in cobalamin biosynthesis
J.Struct.Biol., 153, 2006
|
|
2AFR
 
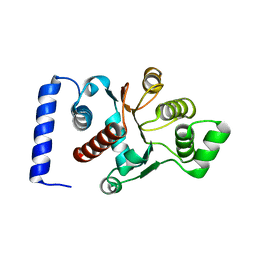 | |
1AG6
 | | PLASTOCYANIN FROM SPINACH | | Descriptor: | COPPER (II) ION, PLASTOCYANIN | | Authors: | Xue, Y, Okvist, M, Young, S. | | Deposit date: | 1997-04-02 | | Release date: | 1998-10-21 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of spinach plastocyanin at 1.7 A resolution.
Protein Sci., 7, 1998
|
|
1BIC
 
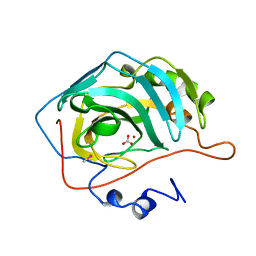 | | CRYSTALLOGRAPHIC ANALYSIS OF THR-200-> HIS HUMAN CARBONIC ANHYDRASE II AND ITS COMPLEX WITH THE SUBSTRATE, HCO3- | | Descriptor: | BICARBONATE ION, CARBONIC ANHYDRASE II, METHYL MERCURY ION, ... | | Authors: | Xue, Y, Vidgren, J, Svensson, L.A, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-01 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystallographic analysis of Thr-200-->His human carbonic anhydrase II and its complex with the substrate, HCO3-.
Proteins, 15, 1993
|
|
1CAJ
 
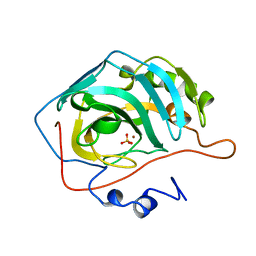 | | STRUCTURAL ANALYSIS OF THE ZINC HYDROXIDE-THR 199-GLU 106 HYDROGEN BONDING NETWORK IN HUMAN CARBONIC ANHYDRASE II | | Descriptor: | CARBONIC ANHYDRASE II, SULFATE ION, ZINC ION | | Authors: | Xue, Y, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-17 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural analysis of the zinc hydroxide-Thr-199-Glu-106 hydrogen-bond network in human carbonic anhydrase II.
Proteins, 17, 1993
|
|
1CAK
 | | STRUCTURAL ANALYSIS OF THE ZINC HYDROXIDE-THR 199-GLU 106 HYDROGEN BONDING NETWORK IN HUMAN CARBONIC ANHYDRASE II | | Descriptor: | CARBONIC ANHYDRASE II, SULFATE ION, ZINC ION | | Authors: | Xue, Y, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-17 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural analysis of the zinc hydroxide-Thr-199-Glu-106 hydrogen-bond network in human carbonic anhydrase II.
Proteins, 17, 1993
|
|
1CAM
 
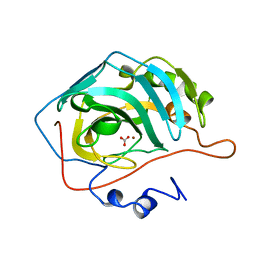 | | STRUCTURAL ANALYSIS OF THE ZINC HYDROXIDE-THR 199-GLU 106 HYDROGEN BONDING NETWORK IN HUMAN CARBONIC ANHYDRASE II | | Descriptor: | BICARBONATE ION, CARBONIC ANHYDRASE II, ZINC ION | | Authors: | Xue, Y, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-17 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural analysis of the zinc hydroxide-Thr-199-Glu-106 hydrogen-bond network in human carbonic anhydrase II.
Proteins, 17, 1993
|
|
4BTJ
 
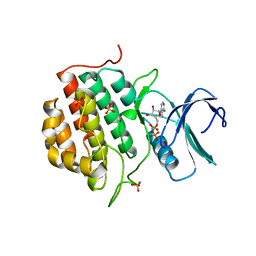 | | TTBK1 in complex with ATP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, PHOSPHATE ION, ... | | Authors: | Xue, Y, Wan, P, Hillertz, P, Schweikart, F, Zhao, Y, Wissler, L, Dekker, N. | | Deposit date: | 2013-06-18 | | Release date: | 2013-09-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | X-Ray Structural Analysis of Tau-Tubulin Kinase 1 and its Interactions with Small Molecular Inhibitors.
Chemmedchem, 8, 2013
|
|
1CAL
 
 | | STRUCTURAL ANALYSIS OF THE ZINC HYDROXIDE-THR 199-GLU 106 HYDROGEN BONDING NETWORK IN HUMAN CARBONIC ANHYDRASE II | | Descriptor: | CARBONIC ANHYDRASE II, ZINC ION | | Authors: | Xue, Y, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-17 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural analysis of the zinc hydroxide-Thr-199-Glu-106 hydrogen-bond network in human carbonic anhydrase II.
Proteins, 17, 1993
|
|
1CAI
 
 | | STRUCTURAL ANALYSIS OF THE ZINC HYDROXIDE-THR 199-GLU 106 HYDROGEN BONDING NETWORK IN HUMAN CARBONIC ANHYDRASE II | | Descriptor: | CARBONIC ANHYDRASE II, SULFATE ION, ZINC ION | | Authors: | Xue, Y, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-17 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural analysis of the zinc hydroxide-Thr-199-Glu-106 hydrogen-bond network in human carbonic anhydrase II.
Proteins, 17, 1993
|
|
4BTK
 
 | | TTBK1 in complex with inhibitor | | Descriptor: | 4-[3-HYDROXYANILINO]-6,7-DIMETHOXYQUINAZOLINE, DIMETHYL SULFOXIDE, TAU-TUBULIN KINASE 1 | | Authors: | Xue, Y, Wan, P, Hillertz, P, Schweikart, F, Zhao, Y, Wissler, L, Dekker, N. | | Deposit date: | 2013-06-18 | | Release date: | 2013-09-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | X-Ray Structural Analysis of Tau-Tubulin Kinase 1 and its Interactions with Small Molecular Inhibitors.
Chemmedchem, 8, 2013
|
|
4BTM
 
 | | TTBK1 in complex with inhibitor | | Descriptor: | DIMETHYL SULFOXIDE, GLYCEROL, SULFATE ION, ... | | Authors: | Xue, Y, Wan, P, Hillertz, P, Schweikart, F, Zhao, Y, Wissler, L, Dekker, N. | | Deposit date: | 2013-06-18 | | Release date: | 2013-09-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | X-Ray Structural Analysis of Tau-Tubulin Kinase 1 and its Interactions with Small Molecular Inhibitors.
Chemmedchem, 8, 2013
|
|
