1PWC
 
 | | penicilloyl acyl enzyme complex of the Streptomyces R61 DD-peptidase with penicillin G | | Descriptor: | D-alanyl-D-alanine carboxypeptidase, OPEN FORM - PENICILLIN G | | Authors: | Silvaggi, N.R, Josephine, H.R, Pratt, R.F, Kelly, J.A. | | Deposit date: | 2003-07-01 | | Release date: | 2004-07-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Crystal structures of complexes between the R61 DD-peptidase and peptidoglycan-mimetic beta-lactams: a non-covalent complex with a "perfect penicillin"
J.Mol.Biol., 345, 2005
|
|
1PW8
 
 | | Covalent Acyl Enzyme Complex Of The R61 DD-Peptidase with A Highly Specific Cephalosporin | | Descriptor: | (6R,7R)-3-[(ACETYLOXY)METHYL]-7-{[(6S)-6-(GLYCYLAMINO)-7-OXIDO-7-OXOHEPTANOYL]AMINO}-8-OXO-5-THIA-1-AZABICYCLO[4.2.0]OCTANE-2-CARBOXYLATE, D-alanyl-D-alanine carboxypeptidase, GLYCEROL | | Authors: | Silvaggi, N.R, Josephine, H.R, Pratt, R.F, Kelly, J.A. | | Deposit date: | 2003-07-01 | | Release date: | 2004-07-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal structures of complexes between the R61 DD-peptidase and peptidoglycan-mimetic beta-lactams: a non-covalent complex with a "perfect penicillin"
J.Mol.Biol., 345, 2005
|
|
1PW1
 
 | | Non-Covalent Complex Of Streptomyces R61 DD-Peptidase With A Highly Specific Penicillin | | Descriptor: | (2S,5R,6R)-6-{[(6R)-6-(GLYCYLAMINO)-7-OXIDO-7-OXOHEPTANOYL]AMINO}-3,3-DIMETHYL-7-OXO-4-THIA-1-AZABICYCLO[3.2.0]HEPTANE-2-CARBOXYLATE, D-alanyl-D-alanine carboxypeptidase, FORMYL GROUP, ... | | Authors: | Silvaggi, N.R, Josephine, H.R, Pratt, R.F, Kelly, J.A. | | Deposit date: | 2003-06-30 | | Release date: | 2004-07-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Crystal structures of complexes between the R61 DD-peptidase and peptidoglycan-mimetic beta-lactams: a non-covalent complex with a "perfect penicillin"
J.Mol.Biol., 345, 2005
|
|
1PWG
 
 | | Covalent Penicilloyl Acyl Enzyme Complex Of The Streptomyces R61 DD-Peptidase With A Highly Specific Penicillin | | Descriptor: | (2R,4S)-2-[(1R)-1-{[(6S)-6-carboxy-6-(glycylamino)hexanoyl]amino}-2-oxoethyl]-5,5-dimethyl-1,3-thiazolidine-4-carboxylic acid, D-alanyl-D-alanine carboxypeptidase | | Authors: | Silvaggi, N.R, Josephine, H.R, Pratt, R.F, Kelly, J.A. | | Deposit date: | 2003-07-01 | | Release date: | 2004-07-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.074 Å) | | Cite: | Crystal structures of complexes between the R61 DD-peptidase and peptidoglycan-mimetic beta-lactams: a non-covalent complex with a "perfect penicillin"
J.Mol.Biol., 345, 2005
|
|
2JKV
 
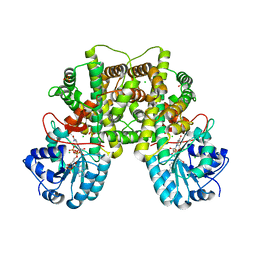 | | Structure of human Phosphogluconate Dehydrogenase in complex with NADPH at 2.53A | | Descriptor: | 6-PHOSPHOGLUCONATE DEHYDROGENASE, DECARBOXYLATING, CHLORIDE ION, ... | | Authors: | Pilka, E.S, Kavanagh, K.L, von Delft, F, Muniz, J.R.C, Murray, J, Picaud, S, Guo, K, Edwards, A, Arrowsmith, C.H, Weigelt, J, Bountra, C, Oppermann, U. | | Deposit date: | 2008-09-01 | | Release date: | 2009-09-01 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.532 Å) | | Cite: | Structure of Human Phosphogluconate Dehydrogenase in Complex with Nadph at 2.53A
To be Published
|
|
4LIU
 
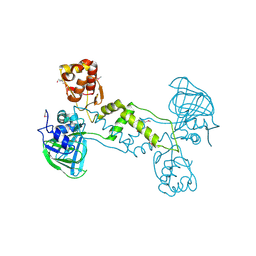 | | Structure of YcfD, a Ribosomal oxygenase from Escherichia coli. | | Descriptor: | 50S ribosomal protein L16 arginine hydroxylase, PHOSPHATE ION, TRIS(HYDROXYETHYL)AMINOMETHANE | | Authors: | Brissett, N.C, Doherty, A.J, Fox, G.C. | | Deposit date: | 2013-07-03 | | Release date: | 2014-05-14 | | Last modified: | 2014-07-09 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Ribosomal oxygenases are structurally conserved from prokaryotes to humans.
Nature, 509, 2014
|
|
4LIV
 
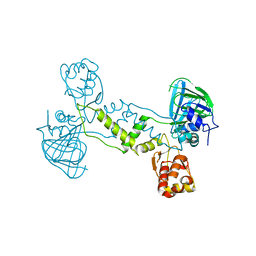 | |
4LIT
 
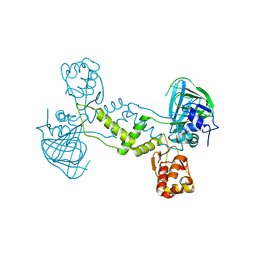 | |
4KQO
 
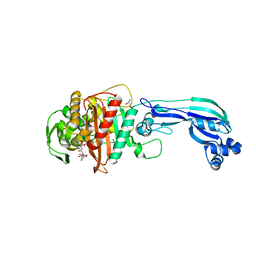 | | Crystal structure of penicillin-binding protein 3 from pseudomonas aeruginosa in complex with piperacillin | | Descriptor: | CHLORIDE ION, GLYCEROL, IMIDAZOLE, ... | | Authors: | Nettleship, J.E, Stuart, D.I, Owens, R.J, Ren, J. | | Deposit date: | 2013-05-15 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Binding of (5S)-Penicilloic Acid to Penicillin Binding Protein 3.
Acs Chem.Biol., 8, 2013
|
|
4KQQ
 
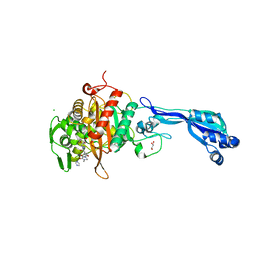 | | CRYSTAL STRUCTURE OF PENICILLIN-BINDING PROTEIN 3 FROM PSEUDOMONAS AERUGINOSA IN COMPLEX WITH (5S)-Penicilloic Acid | | Descriptor: | (2S,4S)-2-[(R)-carboxy{[(2R)-2-{[(4-ethyl-2,3-dioxopiperazin-1-yl)carbonyl]amino}-2-phenylacetyl]amino}methyl]-5,5-dimethyl-1,3-thiazolidine-4-carboxylic acid, CHLORIDE ION, GLYCEROL, ... | | Authors: | Nettleship, J.E, Stuart, D.I, Owens, R.J, Ren, J. | | Deposit date: | 2013-05-15 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Binding of (5S)-Penicilloic Acid to Penicillin Binding Protein 3.
Acs Chem.Biol., 8, 2013
|
|
4KQR
 
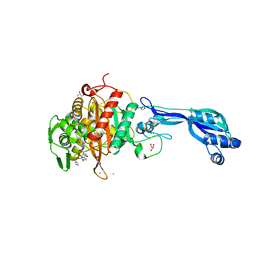 | | CRYSTAL STRUCTURE OF PENICILLIN-BINDING PROTEIN 3 FROM PSEUDOMONAS AERUGINOSA IN COMPLEX WITH (5S)-Penicilloic Acid | | Descriptor: | (2S,4S)-2-[(R)-carboxy{[(2R)-2-{[(4-ethyl-2,3-dioxopiperazin-1-yl)carbonyl]amino}-2-phenylacetyl]amino}methyl]-5,5-dimethyl-1,3-thiazolidine-4-carboxylic acid, CHLORIDE ION, GLYCEROL, ... | | Authors: | Nettleship, J.E, Stuart, D.I, Owens, R.J, Ren, J. | | Deposit date: | 2013-05-15 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Binding of (5S)-Penicilloic Acid to Penicillin Binding Protein 3.
Acs Chem.Biol., 8, 2013
|
|
4C63
 
 | | ULTRA HIGH RESOLUTION DICKERSON-DREW DODECAMER B-DNA WITH 5- METHYLCYSTOSINE MODIFICATION | | Descriptor: | 5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*5CMP*GP*CP*GP)-3', MAGNESIUM ION | | Authors: | Mcdonough, M, El-Sagheer, A.H, Brown, T, Schofield, C.J. | | Deposit date: | 2013-09-17 | | Release date: | 2013-10-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Structural insights into how 5-hydroxymethylation influences transcription factor binding.
Chem. Commun. (Camb.), 50, 2014
|
|
4C64
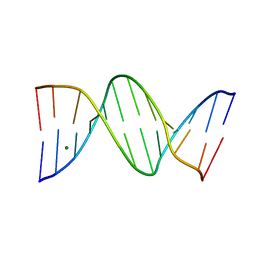 
 | | ULTRA HIGH RESOLUTION DICKERSON-DREW DODECAMER B-DNA | | Descriptor: | 5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*GP)-3', MAGNESIUM ION | | Authors: | Mcdonough, M, El-Sagheer, A.H, Brown, T, Schofield, C.J. | | Deposit date: | 2013-09-17 | | Release date: | 2013-10-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Structural insights into how 5-hydroxymethylation influences transcription factor binding.
Chem. Commun. (Camb.), 50, 2014
|
|
4C5X
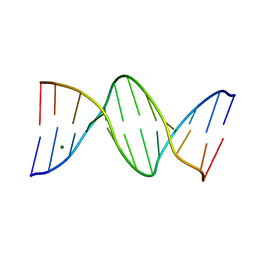 
 | | Ultra High Resolution Dickerson-Drew dodecamer B-DNA with 5-Hydroxymethyl-cytosine Modification | | Descriptor: | 5'-D(CP*GP*CP*GP*AP*AP*TP*TP*5HCP*GP*CP*GP)-3', MAGNESIUM ION | | Authors: | McDonough, M, El-Sagheer, A.H, Brown, T, Schofield, C.J. | | Deposit date: | 2013-09-16 | | Release date: | 2013-10-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Structural insights into how 5-hydroxymethylation influences transcription factor binding.
Chem. Commun. (Camb.), 50, 2014
|
|
5LAS
 
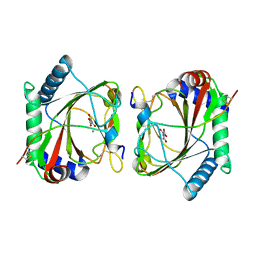 | |
5LB6
 
 | |
5LBE
 
 | |
5LBC
 
 | |
5LBB
 
 | |
5L9B
 
 | |
5LA9
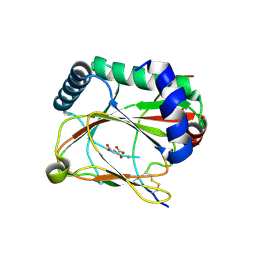 
 | |
5L9R
 
 | |
5LBF
 
 | |
5LAT
 
 | |
2YBS
 
 | |
