3FL5
 
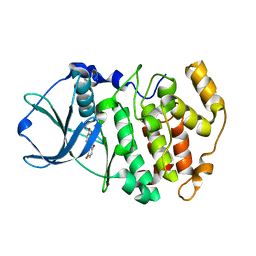 | | Protein kinase CK2 in complex with the inhibitor Quinalizarin | | Descriptor: | 1,2,5,8-tetrahydroxyanthracene-9,10-dione, Casein kinase II subunit alpha, DI(HYDROXYETHYL)ETHER | | Authors: | Mazzorana, M, Franchin, C, Battistutta, R. | | Deposit date: | 2008-12-18 | | Release date: | 2009-08-18 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Quinalizarin as a potent, selective and cell-permeable inhibitor of protein kinase CK2
Biochem.J., 421, 2009
|
|
6FSW
 
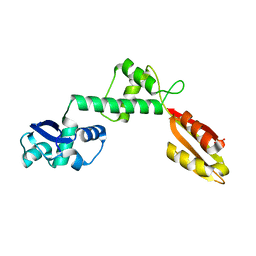 | | Structure of Archaeoglobus fulgidus SBDS protein at 1.9 Angstrom | | Descriptor: | DI(HYDROXYETHYL)ETHER, Ribosome maturation protein SDO1-like protein | | Authors: | Mazzorana, M, Foadi, J, Siliqi, D, Sanchez-Puig, N. | | Deposit date: | 2018-02-20 | | Release date: | 2018-03-21 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Conformational flexibility of proteins involved in ribosome biogenesis: investigations via Small Angle X-ray Scattering (SAXS)
Crystals, 8, 2018
|
|
4A7C
 
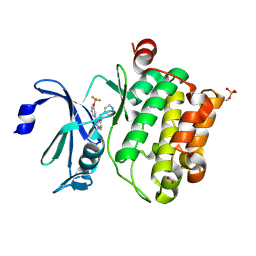 | | Crystal structure of PIM1 kinase with ETP46546 | | Descriptor: | ACETATE ION, IMIDAZOLE, N-(piperidin-4-ylmethyl)-3-[3-(trifluoromethyloxy)phenyl]-[1,2,3]triazolo[4,5-b]pyridin-5-amine, ... | | Authors: | Mazzorana, M, Montoya, G. | | Deposit date: | 2011-11-12 | | Release date: | 2012-02-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Hit to Lead Evaluation of 1,2,3-Triazolo[4,5-B]Pyridines as Pim Kinase Inhibitors.
Bioorg.Med.Chem.Lett., 22, 2012
|
|
8R5J
 
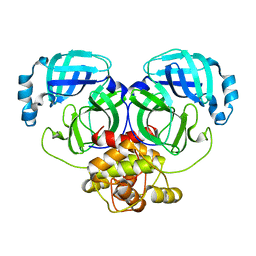 | | Crystal structure of MERS-CoV main protease | | Descriptor: | Non-structural protein 11 | | Authors: | Balcomb, B.H, Fairhead, M, Koekemoer, L, Lithgo, R.M, Aschenbrenner, J.C, Chandran, A.V, Godoy, A.S, Lukacik, P, Marples, P.G, Mazzorana, M, Ni, X, Strain-Damerell, C, Thompson, W, Tomlinson, C.W.E, Wild, C, Winokan, M, Fearon, D, Walsh, M.A, von Delft, F. | | Deposit date: | 2023-11-16 | | Release date: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.898 Å) | | Cite: | Crystal structure of MERS-CoV main protease
To Be Published
|
|
6QS5
 
 | |
4DGN
 
 | | Crystal Structure of maize CK2 in complex with the inhibitor luteolin | | Descriptor: | 2-(3,4-dihydroxyphenyl)-5,7-dihydroxy-4H-chromen-4-one, Casein kinase II subunit alpha | | Authors: | Lolli, G, Mazzorana, M, Battistutta, R. | | Deposit date: | 2012-01-26 | | Release date: | 2012-08-01 | | Last modified: | 2013-01-02 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Inhibition of protein kinase CK2 by flavonoids and tyrphostins. A structural insight.
Biochemistry, 51, 2012
|
|
4DGM
 
 | | Crystal Structure of maize CK2 in complex with the inhibitor apigenin | | Descriptor: | 1,2-ETHANEDIOL, 5,7-dihydroxy-2-(4-hydroxyphenyl)-4H-chromen-4-one, Casein kinase II subunit alpha, ... | | Authors: | Lolli, G, Mazzorana, M, Battistutta, R. | | Deposit date: | 2012-01-26 | | Release date: | 2012-08-01 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Inhibition of protein kinase CK2 by flavonoids and tyrphostins. A structural insight.
Biochemistry, 51, 2012
|
|
8AGI
 
 | | Structure of human Heat shock protein 90-alpha N-terminal domain (Hsp90-NTD) in complex with JMC31 | | Descriptor: | 1-[2,4-bis(oxidanyl)-5-propan-2-yl-phenyl]-5-[4-[(cyclohexylmethylamino)methyl]phenyl]-~{N}-ethyl-1,2,3-triazole-4-carboxamide, Heat shock protein HSP 90-alpha | | Authors: | Tassone, G, Pozzi, C, Mazzorana, M, Mangani, S, Maramai, S. | | Deposit date: | 2022-07-20 | | Release date: | 2022-09-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Characterization of Human Heat Shock Protein 90 N-Terminal Domain and Its Variants K112R and K112A in Complex with a Potent 1,2,3-Triazole-Based Inhibitor.
Int J Mol Sci, 23, 2022
|
|
8AGJ
 
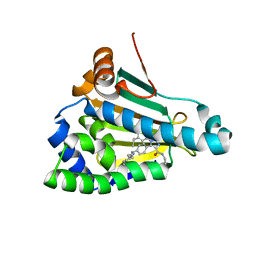 | | Structure of human Heat shock protein 90-alpha N-terminal domain (Hsp90-NTD) variant K112A in complex with JMC31 | | Descriptor: | 1-[2,4-bis(oxidanyl)-5-propan-2-yl-phenyl]-5-[4-[(cyclohexylmethylamino)methyl]phenyl]-~{N}-ethyl-1,2,3-triazole-4-carboxamide, Heat shock protein HSP 90-alpha | | Authors: | Tassone, G, Pozzi, C, Mazzorana, M, Mangani, S, Maramai, S. | | Deposit date: | 2022-07-20 | | Release date: | 2022-09-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Structural Characterization of Human Heat Shock Protein 90 N-Terminal Domain and Its Variants K112R and K112A in Complex with a Potent 1,2,3-Triazole-Based Inhibitor.
Int J Mol Sci, 23, 2022
|
|
8AGL
 
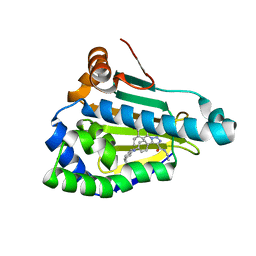 | | Structure of human Heat shock protein 90-alpha N-terminal domain (Hsp90-NTD) variant K112R in complex with JMC31 | | Descriptor: | 1-[2,4-bis(oxidanyl)-5-propan-2-yl-phenyl]-5-[4-[(cyclohexylmethylamino)methyl]phenyl]-~{N}-ethyl-1,2,3-triazole-4-carboxamide, Heat shock protein HSP 90-alpha | | Authors: | Tassone, G, Pozzi, C, Mazzorana, M, Mangani, S, Maramai, S. | | Deposit date: | 2022-07-20 | | Release date: | 2022-09-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural Characterization of Human Heat Shock Protein 90 N-Terminal Domain and Its Variants K112R and K112A in Complex with a Potent 1,2,3-Triazole-Based Inhibitor.
Int J Mol Sci, 23, 2022
|
|
3PWD
 
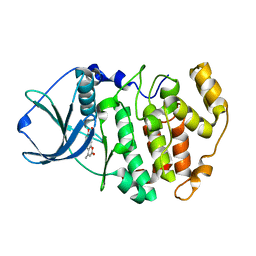 | | Crystal structure of maize CK2 in complex with NBC (Z1) | | Descriptor: | 8-hydroxy-4-methyl-9-nitro-2H-benzo[g]chromen-2-one, Casein kinase II subunit alpha | | Authors: | Battistutta, R, Mazzorana, M. | | Deposit date: | 2010-12-08 | | Release date: | 2011-11-02 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural features underlying the selectivity of the kinase inhibitors NBC and dNBC: role of a nitro group that discriminates between CK2 and DYRK1A
Cell.Mol.Life Sci., 69, 2012
|
|
1ZOH
 
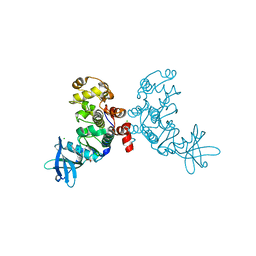 | | Crystal structure of protein kinase CK2 in complex with TBB-derivatives inhibitors | | Descriptor: | 1,2-ETHANEDIOL, 5,6,7,8-TETRABROMO-1-METHYL-2,3-DIHYDRO-1H-IMIDAZO[1,2-A]BENZIMIDAZOLE, CHLORIDE ION, ... | | Authors: | Battistutta, R, Mazzorana, M, Sarno, S, Kazimierczuk, Z, Zanotti, G, Pinna, L.A. | | Deposit date: | 2005-05-13 | | Release date: | 2005-11-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Inspecting the structure-activity relationship of protein kinase CK2 inhibitors derived from tetrabromo-benzimidazole.
Chem.Biol., 12, 2005
|
|
1ZOE
 
 | | Crystal structure of protein kinase CK2 in complex with TBB-derivatives inhibitors | | Descriptor: | 4,5,6,7-TETRABROMO-N,N-DIMETHYL-1H-BENZIMIDAZOL-2-AMINE, CHLORIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Battistutta, R, Mazzorana, M, Sarno, S, Kazimierczuk, Z, Zanotti, G, Pinna, L.A. | | Deposit date: | 2005-05-13 | | Release date: | 2005-11-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Inspecting the structure-activity relationship of protein kinase CK2 inhibitors derived from tetrabromo-benzimidazole.
Chem.Biol., 12, 2005
|
|
1ZOG
 
 | | Crystal structure of protein kinase CK2 in complex with TBB-derivatives | | Descriptor: | 4,5,6,7-TETRABROMO-2-(METHYLSULFANYL)-1H-BENZIMIDAZOLE, PROTEIN KINASE CK2, alpha SUBUNIT | | Authors: | Battistutta, R, Mazzorana, M, Sarno, S, Kazimierczuk, Z, Zanotti, G, Pinna, L.A. | | Deposit date: | 2005-05-13 | | Release date: | 2005-11-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Inspecting the structure-activity relationship of protein kinase CK2 inhibitors derived from tetrabromo-benzimidazole.
Chem.Biol., 12, 2005
|
|
3KXM
 
 | | Crystal structure of Z. mays CK2 kinase alpha subunit in complex with the inhibitor K74 | | Descriptor: | Casein kinase II subunit alpha, N-methyl-2-[(4,5,6,7-tetrabromo-1-methyl-1H-benzimidazol-2-yl)sulfanyl]acetamide | | Authors: | Papinutto, E, Franchin, C, Battistutta, R. | | Deposit date: | 2009-12-03 | | Release date: | 2010-11-17 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | ATP site-directed inhibitors of protein kinase CK2: an update.
Curr Top Med Chem, 11, 2011
|
|
3KXG
 
 | | Crystal structure of Z. mays CK2 kinase alpha subunit in complex with the inhibitor 3,4,5,6,7-pentabromo-1H-indazole (K64) | | Descriptor: | 3,4,5,6,7-pentabromo-1H-indazole, Casein kinase II subunit alpha | | Authors: | Papinutto, E, Franchin, C, Battistutta, R. | | Deposit date: | 2009-12-03 | | Release date: | 2010-11-17 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | ATP site-directed inhibitors of protein kinase CK2: an update.
Curr Top Med Chem, 11, 2011
|
|
3R0T
 
 | | Crystal structure of human protein kinase CK2 alpha subunit in complex with the inhibitor CX-5279 | | Descriptor: | 1,2-ETHANEDIOL, 3-(cyclopropylamino)-5-{[3-(trifluoromethyl)phenyl]amino}pyrimido[4,5-c]quinoline-8-carboxylic acid, Casein kinase II subunit alpha, ... | | Authors: | Battistutta, R, Papinutto, E, Lolli, G, Pierre, F, Haddach, M, Ryckman, D.M. | | Deposit date: | 2011-03-09 | | Release date: | 2011-12-07 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Unprecedented selectivity and structural determinants of a new class of protein kinase CK2 inhibitors in clinical trials for the treatment of cancer.
Biochemistry, 50, 2011
|
|
6YB7
 
 | | SARS-CoV-2 main protease with unliganded active site (2019-nCoV, coronavirus disease 2019, COVID-19). | | Descriptor: | 3C-like proteinase, DI(HYDROXYETHYL)ETHER, DIMETHYL SULFOXIDE | | Authors: | Owen, C.D, Lukacik, P, Strain-Damerell, C.M, Douangamath, A, Powell, A.J, Fearon, D, Brandao-Neto, J, Crawshaw, A.D, Aragao, D, Williams, M, Flaig, R, Hall, D.R, McAuley, K.E, Mazzorana, M, Stuart, D.I, von Delft, F, Walsh, M.A. | | Deposit date: | 2020-03-16 | | Release date: | 2020-03-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | COVID-19 main protease with unliganded active site
To Be Published
|
|
6Y84
 
 | | SARS-CoV-2 main protease with unliganded active site (2019-nCoV, coronavirus disease 2019, COVID-19) | | Descriptor: | 3C-like proteinase nsp5, DIMETHYL SULFOXIDE | | Authors: | Owen, C.D, Lukacik, P, Strain-Damerell, C.M, Douangamath, A, Powell, A.J, Fearon, D, Brandao-Neto, J, Crawshaw, A.D, Aragao, D, Williams, M, Flaig, R, Hall, D.R, McAuley, K.E, Mazzorana, M, Stuart, D.I, von Delft, F, Walsh, M.A. | | Deposit date: | 2020-03-03 | | Release date: | 2020-03-11 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | COVID-19 main protease with unliganded active site
To Be Published
|
|
4ZBC
 
 | | A dehydrated form of glucose isomerase collected at 100K. | | Descriptor: | MANGANESE (II) ION, Xylose isomerase, alpha-D-glucopyranose, ... | | Authors: | Sandy, J. | | Deposit date: | 2015-04-14 | | Release date: | 2016-03-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A generic protocol for protein crystal dehydration using the HC1b humidity controller.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
4ZB5
 
 | | A form of glucose isomerase collected at 100K. | | Descriptor: | MANGANESE (II) ION, Xylose isomerase, alpha-D-glucopyranose | | Authors: | Sandy, J. | | Deposit date: | 2015-04-14 | | Release date: | 2016-03-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A generic protocol for protein crystal dehydration using the HC1b humidity controller.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
4ZB0
 
 | | A dehydrated form of glucose isomerase collected at room temperature. | | Descriptor: | MANGANESE (II) ION, Xylose isomerase, alpha-D-glucopyranose, ... | | Authors: | Sandy, J. | | Deposit date: | 2015-04-14 | | Release date: | 2016-03-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A generic protocol for protein crystal dehydration using the HC1b humidity controller.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
4ZB2
 
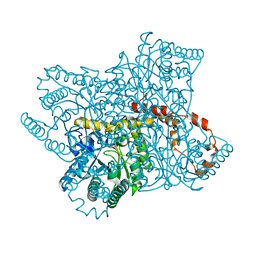 | | A native form of glucose isomerase collected at room temperature. | | Descriptor: | MANGANESE (II) ION, Xylose isomerase, alpha-D-glucopyranose, ... | | Authors: | Sandy, J. | | Deposit date: | 2015-04-14 | | Release date: | 2016-03-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A generic protocol for protein crystal dehydration using the HC1b humidity controller.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
3PE1
 
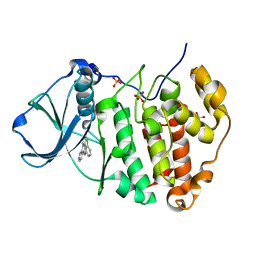 | | Crystal structure of human protein kinase CK2 alpha subunit in complex with the inhibitor CX-4945 | | Descriptor: | 5-[(3-chlorophenyl)amino]benzo[c][2,6]naphthyridine-8-carboxylic acid, Casein kinase II subunit alpha, SULFATE ION | | Authors: | Battistutta, R, Papinutto, E, Lolli, G, Pierre, F, Haddach, M, Ryckman, D.M. | | Deposit date: | 2010-10-25 | | Release date: | 2011-09-07 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Unprecedented selectivity and structural determinants of a new class of protein kinase CK2 inhibitors in clinical trials for the treatment of cancer.
Biochemistry, 50, 2011
|
|
3PE2
 
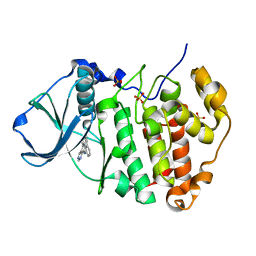 | | Crystal structure of human protein kinase CK2 in complex with the inhibitor CX-5011 | | Descriptor: | 5-[(3-ethynylphenyl)amino]pyrimido[4,5-c]quinoline-8-carboxylic acid, Casein kinase II subunit alpha, SULFATE ION | | Authors: | Battistutta, R, Papinutto, E, Lolli, G, Pierre, F, Haddach, M, Ryckman, D.M. | | Deposit date: | 2010-10-25 | | Release date: | 2011-09-07 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Unprecedented selectivity and structural determinants of a new class of protein kinase CK2 inhibitors in clinical trials for the treatment of cancer.
Biochemistry, 50, 2011
|
|
