1VF3
 
 | | cGSTA1-1 in complex with glutathione conjugate of CDNB | | Descriptor: | ACETIC ACID, GLUTATHIONE S-(2,4 DINITROBENZENE), Glutathione S-transferase 3 | | Authors: | Lin, S.C, Lo, Y.C, Tam, M.F, Liaw, Y.C. | | Deposit date: | 2004-04-07 | | Release date: | 2005-08-23 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal structures of chicken glutathione S-transferase A1-1
To be Published
|
|
1VF4
 
 | | cGSTA1-1 apo form | | Descriptor: | ACETIC ACID, Glutathione S-transferase 3 | | Authors: | Lin, S.C, Lo, Y.C, Tam, M.F, Liaw, Y.C. | | Deposit date: | 2004-04-08 | | Release date: | 2005-08-23 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Crystal structures of chicken glutathione S-transferase A1-1
To be Published
|
|
1VF1
 
 | |
1VF2
 
 | |
1U2A
 
 | |
2U2A
 
 | |
5WVC
 
 | | Structure of the CARD-CARD disk | | Descriptor: | Apoptotic protease-activating factor 1, Caspase, IODIDE ION | | Authors: | Lin, S.C, Lo, Y.C, Su, T.W. | | Deposit date: | 2016-12-24 | | Release date: | 2017-04-19 | | Method: | X-RAY DIFFRACTION (2.993 Å) | | Cite: | Structural Insights into DD-Fold Assembly and Caspase-9 Activation by the Apaf-1 Apoptosome.
Structure, 25, 2017
|
|
2POP
 
 | | The Crystal Structure of TAB1 and BIR1 complex | | Descriptor: | Baculoviral IAP repeat-containing protein 4, Mitogen-activated protein kinase kinase kinase 7-interacting protein 1, ZINC ION | | Authors: | Lin, S.C. | | Deposit date: | 2007-04-27 | | Release date: | 2007-07-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | XIAP Induces NF-kappaB Activation via the BIR1/TAB1 Interaction and BIR1 Dimerization.
Mol.Cell, 26, 2007
|
|
2POM
 
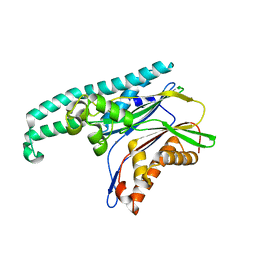 | | TAB1 with manganese ion | | Descriptor: | MANGANESE (II) ION, Mitogen-activated protein kinase kinase kinase 7-interacting protein 1 | | Authors: | Lin, S.C. | | Deposit date: | 2007-04-26 | | Release date: | 2007-07-03 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | XIAP Induces NF-kappaB Activation via the BIR1/TAB1 Interaction and BIR1 Dimerization.
Mol.Cell, 26, 2007
|
|
5H07
 
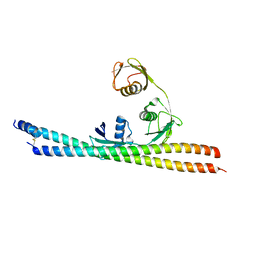 | | TNIP2-Ub complex, C2 form | | Descriptor: | Polyubiquitin-C, TNFAIP3-interacting protein 2 | | Authors: | Lo, Y.C, Lin, S.C. | | Deposit date: | 2016-10-04 | | Release date: | 2017-03-08 | | Method: | X-RAY DIFFRACTION (2.586 Å) | | Cite: | Structural Insights into Linear Tri-ubiquitin Recognition by A20-Binding Inhibitor of NF-kappa B, ABIN-2
Structure, 25, 2017
|
|
6UMX
 
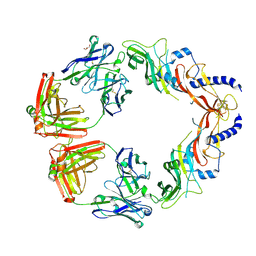 | | Structural basis for specific inhibition of extracellular activation of pro/latent myostatin by SRK-015 | | Descriptor: | GL29H4-16 Fab Heavy Chain,GL29H4-16 Fab Heavy Chain, GL29H4-16 Fab Light Chain,GL29H4-16 Fab Light Chain, GLYCEROL, ... | | Authors: | Dagbay, K.B, Treece, E, Streich Jr, F.C, Jackson, J.W, Faucette, R.R, Nikiforov, A, Lin, S.C, Bostion, C.J, Nicholls, S.B, Capili, A.D, Carven, G.J. | | Deposit date: | 2019-10-10 | | Release date: | 2020-02-26 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | Structural basis of specific inhibition of extracellular activation of pro- or latent myostatin by the monoclonal antibody SRK-015.
J.Biol.Chem., 295, 2020
|
|
3MRS
 
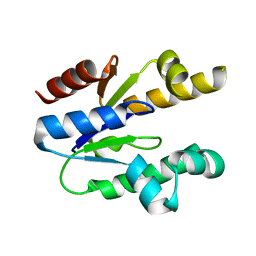 | |
3N2E
 | | Crystal structure of Helicobactor pylori shikimate kinase in complex with NSC162535 | | Descriptor: | 7-amino-4-hydroxy-3-[(E)-(5-hydroxy-7-sulfonaphthalen-2-yl)diazenyl]naphthalene-2-sulfonic acid, L(+)-TARTARIC ACID, Shikimate kinase | | Authors: | Cheng, W.C, Chen, T.J, Lin, S.C, Wang, W.C. | | Deposit date: | 2010-05-18 | | Release date: | 2011-05-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Structures of Helicobacter pylori shikimate kinase reveal a selective inhibitor-induced-fit mechanism
Plos One, 7, 2012
|
|
3MUF
 
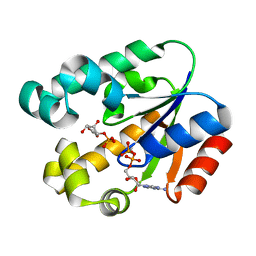 | | Shikimate kinase from Helicobacter pylori in complex with shikimate-3-phosphate and ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, SHIKIMATE-3-PHOSPHATE, Shikimate kinase | | Authors: | Cheng, W.C, Chen, T.J, Lin, S.C, Wang, W.C. | | Deposit date: | 2010-05-03 | | Release date: | 2011-05-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of Helicobacter pylori shikimate kinase reveal a selective inhibitor-induced-fit mechanism
Plos One, 7, 2012
|
|
7EAL
 
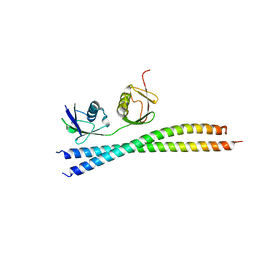 | |
7EAO
 
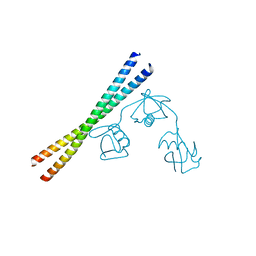 | |
7EB9
 
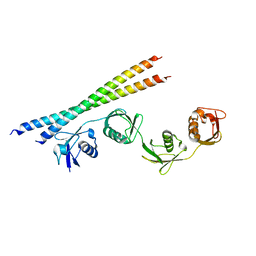 | |
4UFP
 
 | | Laboratory evolved variant R-C1B1D33 of potato epoxide hydrolase StEH1 | | Descriptor: | EPOXIDE HYDROLASE | | Authors: | Carlsson, A.J, Bauer, P, Nilsson, M, Dobritzsch, D, Kamerlin, S.C.L, Widersten, M. | | Deposit date: | 2015-03-17 | | Release date: | 2016-04-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Laboratory Evolved Enzymes Provide Snapshots of the Development of Enantioconvergence in Enzyme-Catalyzed Epoxide Hydrolysis.
Chembiochem, 17, 2016
|
|
4UFO
 
 | | Laboratory evolved variant R-C1B1D33E6 of potato epoxide hydrolase StEH1 | | Descriptor: | EPOXIDE HYDROLASE | | Authors: | Carlsson, A.J, Bauer, P, Nilsson, M, Dobritzsch, D, Kamerlin, S.C.L, Widersten, M. | | Deposit date: | 2015-03-17 | | Release date: | 2016-04-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Laboratory Evolved Enzymes Provide Snapshots of the Development of Enantioconvergence in Enzyme-Catalyzed Epoxide Hydrolysis.
Chembiochem, 17, 2016
|
|
3FX0
 | |
3PHJ
 
 | |
3PHH
 
 | | Shikimate 5-Dehydrogenase (aroE) from Helicobacter pylori in complex with Shikimate | | Descriptor: | (3R,4S,5R)-3,4,5-TRIHYDROXYCYCLOHEX-1-ENE-1-CARBOXYLIC ACID, Shikimate dehydrogenase | | Authors: | Cheng, W.C, Lin, S.C, Wang, W.C. | | Deposit date: | 2010-11-04 | | Release date: | 2011-11-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Shikimate 5-Dehydrogenase (aroE) from Helicobacter pylori in complex with Shikimate
To be Published
|
|
3PHG
 
 | |
3PHI
 
 | | Shikimate 5-Dehydrogenase (aroE) from Helicobacter pylori in complex with Shikimate and NADPH | | Descriptor: | (3R,4S,5R)-3,4,5-TRIHYDROXYCYCLOHEX-1-ENE-1-CARBOXYLIC ACID, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Shikimate dehydrogenase | | Authors: | Cheng, W.C, Lin, S.C, Wang, W.C. | | Deposit date: | 2010-11-04 | | Release date: | 2011-11-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Shikimate 5-Dehydrogenase (aroE) from Helicobacter pylori in complex with Shikimate and NADPH
To be Published
|
|
6Z1M
 
 | | Structure of an Ancestral glycosidase (family 1) bound to heme | | Descriptor: | 1,2-ETHANEDIOL, Ancestral reconstructed glycosidase, GLYCEROL, ... | | Authors: | Gavira, J.A, Risso, V.A, Sanchez-Ruiz, J.M, Gamiz-Arco, G, Gutierrez-Rus, L, Ibarra-Molero, B, Oshino, Y, Petrovic, D, Romero-Rivera, A, Seelig, B, Kamerlin, S.C.L, Gaucher, E.A. | | Deposit date: | 2020-05-14 | | Release date: | 2020-07-22 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Heme-binding enables allosteric modulation in an ancient TIM-barrel glycosidase.
Nat Commun, 12, 2021
|
|
