6RIO
 
 | | Imidazole Polyamide-DNA complex NMR structure (5'-CGATGTACATCG-3') | | Descriptor: | 3-[3-[[4-[[4-[[4-[[4-[[(2~{R})-2-azaniumyl-4-[[1-methyl-4-[[1-methyl-4-[[1-methyl-4-[(1-methylimidazol-2-yl)carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]butanoyl]amino]-1-methyl-imidazol-2-yl]carbonylamino]-1-methyl-pyrrol-2-yl]carbonylamino]-1-methyl-pyrrol-2-yl]carbonylamino]-1-methyl-pyrrol-2-yl]carbonylamino]propanoylamino]propyl-dimethyl-azanium, DNA (5'-(*(DC5)P*GP*AP*TP*GP*TP*AP*CP*AP*TP*CP*(DG3))-3') | | Authors: | Padroni, G, Withers, J.M, Taladriz-Sender, A, Reichenbach, L.F, Parkinson, J.A, Burley, G.A. | | Deposit date: | 2019-04-24 | | Release date: | 2019-06-12 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Sequence-Selective Minor Groove Recognition of a DNA Duplex Containing Synthetic Genetic Components.
J.Am.Chem.Soc., 141, 2019
|
|
6I4N
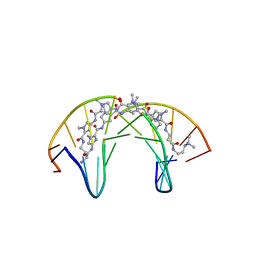 
 | | Dodecamer DNA containing the synthetic base pair P-Z in complex with a pyrrole-imidazole polyamide | | Descriptor: | 3-[3-[[4-[[4-[[4-[[4-[[(2~{R})-2-azaniumyl-4-[[1-methyl-4-[[1-methyl-4-[[1-methyl-4-[(1-methylimidazol-2-yl)carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]butanoyl]amino]-1-methyl-imidazol-2-yl]carbonylamino]-1-methyl-pyrrol-2-yl]carbonylamino]-1-methyl-pyrrol-2-yl]carbonylamino]-1-methyl-pyrrol-2-yl]carbonylamino]propanoylamino]propyl-dimethyl-azanium, DNA (5'-D(*CP*GP*AP*TP*(DP)P*TP*AP*(DZ)P*AP*TP*CP*G)-3') | | Authors: | Padroni, G, Parkison, J, Burley, G.A. | | Deposit date: | 2018-11-10 | | Release date: | 2019-06-12 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Sequence-Selective Minor Groove Recognition of a DNA Duplex Containing Synthetic Genetic Components.
J.Am.Chem.Soc., 141, 2019
|
|
6I4O
 
 | |
5OCZ
 
 | | Free DNA_hairpin polyamides studies | | Descriptor: | DNA (5'-D(*CP*GP*AP*TP*GP*TP*AP*CP*AP*TP*CP*G)-3') | | Authors: | Padroni, G, Parkinson, J, Burley, G.A. | | Deposit date: | 2017-07-04 | | Release date: | 2017-12-06 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis of DNA duplex distortion induced by thiazole-containing hairpin polyamides.
Nucleic Acids Res., 46, 2018
|
|
5ODM
 
 | | NtiPr polyamide in complex with 5'CGATGTACTACG3 | | Descriptor: | 3-(3-azaniumylpropanoylamino)propyl-dimethyl-azanium, 4-AMINO-(1-METHYLIMIDAZOLE)-2-CARBOXYLIC ACID, 4-AMINO-(1-METHYLPYRROLE)-2-CARBOXYLIC ACID, ... | | Authors: | Padroni, G, Parkinson, J, Burley, G.A. | | Deposit date: | 2017-07-06 | | Release date: | 2017-12-06 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis of DNA duplex distortion induced by thiazole-containing hairpin polyamides.
Nucleic Acids Res., 46, 2018
|
|
5ODF
 
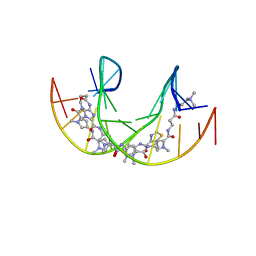 | |
5OE1
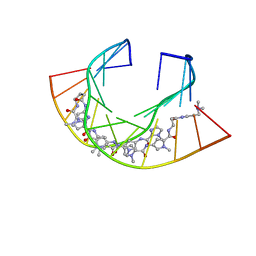 
 | | Im polyamide in complex with 5'CGATGTACATCG3'- hairpin polyamides studies | | Descriptor: | 1-methylimidazole-2-carboxylic acid, 3-(3-azaniumylpropanoylamino)propyl-dimethyl-azanium, 4-AMINO-(1-METHYLIMIDAZOLE)-2-CARBOXYLIC ACID, ... | | Authors: | Padroni, G, Parkinson, J, Burley, G.A. | | Deposit date: | 2017-07-07 | | Release date: | 2017-12-06 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis of DNA duplex distortion induced by thiazole-containing hairpin polyamides.
Nucleic Acids Res., 46, 2018
|
|
1JHI
 
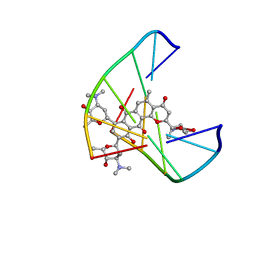 | | Solution Structure of a Hedamycin-DNA complex | | Descriptor: | 5'-D(*AP*CP*CP*(HEH)GP*GP*T)-3', HEDAMYCIN | | Authors: | Owen, E.A, Burley, G.A, Carver, J.A, Wickham, G, Keniry, M.A. | | Deposit date: | 2001-06-27 | | Release date: | 2003-07-08 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural investigation of the hedamycin:d(ACCGGT)2 complex by NMR and restrained molecular dynamics.
Biochem.Biophys.Res.Commun., 290, 2002
|
|
9GN4
 
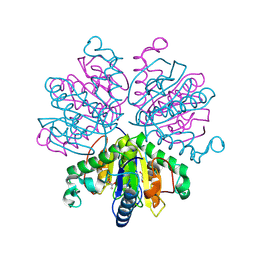 | | Nucleoside-2'-deoxyribosyltransferase from Lactobacillus leichmannii. Y7F/D72N mutant with cytidine | | Descriptor: | 4-AMINO-1-BETA-D-RIBOFURANOSYL-2(1H)-PYRIMIDINONE, Nucleoside deoxyribosyltransferase | | Authors: | Ascham, A, Salihovic, A, Burley, G, Grogan, G. | | Deposit date: | 2024-08-30 | | Release date: | 2024-12-25 | | Last modified: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Biocatalytic synthesis of ribonucleoside analogues using nucleoside transglycosylase-2.
Chem Sci, 16, 2025
|
|
9GN2
 
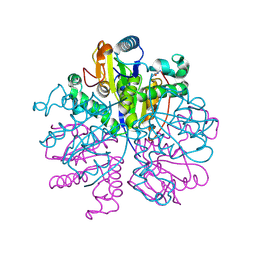 | | Nucleoside-2'-deoxyribosyltransferase from Lactobacillus leichmannii. Complex with ribose and cytosine | | Descriptor: | 6-AMINOPYRIMIDIN-2(1H)-ONE, Nucleoside deoxyribosyltransferase, alpha-D-ribofuranose | | Authors: | Ascham, A, Salihovic, A, Burley, G, Grogan, G. | | Deposit date: | 2024-08-30 | | Release date: | 2024-12-25 | | Last modified: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Biocatalytic synthesis of ribonucleoside analogues using nucleoside transglycosylase-2.
Chem Sci, 16, 2025
|
|
9EZK
 
 | |
9F08
 
 | | Nucleoside-2'-deoxyribosyltransferase from Lactobacillus leichmannii. Covalent complex with 2-deoxyribose. | | Descriptor: | 2-deoxy-beta-D-erythro-pentofuranose, Nucleoside deoxyribosyltransferase | | Authors: | Ascham, A, Salihovic, A, Burley, G, Grogan, G. | | Deposit date: | 2024-04-15 | | Release date: | 2024-12-04 | | Method: | X-RAY DIFFRACTION (2.371 Å) | | Cite: | Gram-scale enzymatic synthesis of 2'-deoxyribonucleoside analogues using nucleoside transglycosylase-2.
Chem Sci, 15, 2024
|
|
9F09
 
 | | Nucleoside-2'-deoxyribosyltransferase from Lactobacillus leichmannii. Complex with 2-deoxyribose, 7-Bromo-1H-imidazo[4,5-b]pyridine and 2'-deoxycytidine | | Descriptor: | 2'-DEOXYCYTIDINE, 2-deoxy-beta-D-erythro-pentofuranose, 7-bromanyl-3~{H}-imidazo[4,5-b]pyridine, ... | | Authors: | Ascham, A, Salihovic, A, Burley, G, Grogan, G. | | Deposit date: | 2024-04-15 | | Release date: | 2024-12-04 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Gram-scale enzymatic synthesis of 2'-deoxyribonucleoside analogues using nucleoside transglycosylase-2.
Chem Sci, 15, 2024
|
|
6RZ2
 
 | | SalL with Chloroadenosine | | Descriptor: | 5'-CHLORO-5'-DEOXYADENOSINE, Adenosyl-chloride synthase | | Authors: | McKean, I, Frese, A, Cuetos, A, Burley, G, Grogan, G. | | Deposit date: | 2019-06-12 | | Release date: | 2020-04-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | S-Adenosyl Methionine Cofactor Modifications Enhance the Biocatalytic Repertoire of Small Molecule C-Alkylation.
Angew.Chem.Int.Ed.Engl., 58, 2019
|
|
6RYZ
 
 | | SalL with S-adenosyl methionine | | Descriptor: | 1,2-ETHANEDIOL, Adenosyl-chloride synthase, CHLORIDE ION, ... | | Authors: | McKean, I, Frese, A, Cuetos, A, Burley, G, Grogan, G. | | Deposit date: | 2019-06-12 | | Release date: | 2020-04-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | S-Adenosyl Methionine Cofactor Modifications Enhance the Biocatalytic Repertoire of Small Molecule C-Alkylation.
Angew.Chem.Int.Ed.Engl., 58, 2019
|
|
6GZ7
 
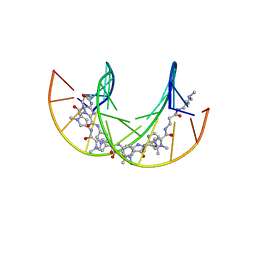 | | Polyamide - DNA complex NMR structure | | Descriptor: | DNA (5'-D(*CP*GP*AP*TP*GP*TP*AP*CP*AP*TP*CP*G)-3'), dimethyl-[3-[3-[[1-methyl-4-[[1-methyl-4-[[1-methyl-4-[[1-methyl-4-[4-[[1-methyl-4-[[1-methyl-4-[[1-methyl-4-[(1-propan-2-ylimidazol-2-yl)carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]butanoylamino]imidazol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]propanoylamino]propyl]azanium | | Authors: | Aman, K. | | Deposit date: | 2018-07-03 | | Release date: | 2018-11-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural and Kinetic Profiling of Allosteric Modulation of Duplex DNA Induced by DNA-Binding Polyamide Analogues.
Chemistry, 25, 2019
|
|
5KTV
 
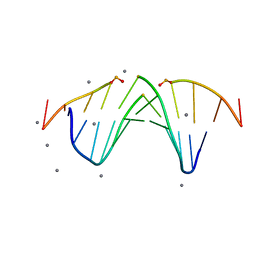 | |
5LJ4
 
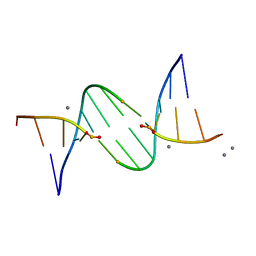 | |
5MGZ
 
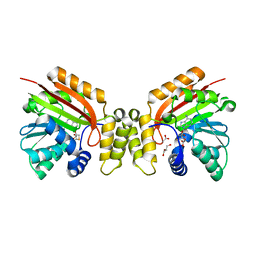 | | Streptomyces Spheroides NovO (8-demethylnovbiocic acid methyltransferase) with SAH | | Descriptor: | 8-demethylnovobiocic acid C(8)-methyltransferase, ACETATE ION, GLYCEROL, ... | | Authors: | Chung, C.-W, Mosley, J, Sadler, J.C. | | Deposit date: | 2016-11-22 | | Release date: | 2017-01-18 | | Last modified: | 2025-12-10 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural and Functional Basis of C-Methylation of Coumarin Scaffolds by NovO.
ACS Chem. Biol., 12, 2017
|
|
