4XXO
 
 | |
9PYF
 
 | | uPA Inhibitory Fab AB2 Complex | | Descriptor: | AB2 Fab Heavy Chain, AB2 Fab Light Chain, Urokinase-type plasminogen activator | | Authors: | Anderson, K.J, Bohn, M.F. | | Deposit date: | 2025-08-07 | | Release date: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structure of an affinity-matured inhibitory recombinant fab against urokinase plasminogen activator reveals basis of potency and specificity.
Biochim Biophys Acta Proteins Proteom, 1869, 2021
|
|
9FYT
 
 | | mAbs in complex with cobratoxin at pH 4.5 | | Descriptor: | Alpha-cobratoxin, CHLORIDE ION, GLYCEROL, ... | | Authors: | Wade, J, Bohn, M.F, Laustsen, A.H, Morth, J.P. | | Deposit date: | 2024-07-03 | | Release date: | 2025-07-16 | | Last modified: | 2025-11-19 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Rational design of antibodies with pH-dependent antigen-binding properties using structural insights from broadly neutralizing antibodies against alpha-neurotoxins.
Mabs, 17, 2025
|
|
9FYS
 
 | | D11 mAbs bound to alpha-Bungarotoxin | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Alpha-bungarotoxin, D11 mAbs Light chain, ... | | Authors: | Wade, J, Bohn, M.F, Laustsen, A.H, Morth, J.P. | | Deposit date: | 2024-07-03 | | Release date: | 2025-07-16 | | Last modified: | 2025-11-19 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Rational design of antibodies with pH-dependent antigen-binding properties using structural insights from broadly neutralizing antibodies against alpha-neurotoxins.
Mabs, 17, 2025
|
|
9HUO
 
 | | A01 mAbs bound to cobratoxin at pH 5.5 | | Descriptor: | Alpha-cobratoxin, CHLORIDE ION, GLYCEROL, ... | | Authors: | Wade, J.W, Bohn, M.F, Laustsen, A.H, Morth, J.P. | | Deposit date: | 2024-12-23 | | Release date: | 2025-11-19 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Rational design of antibodies with pH-dependent antigen-binding properties using structural insights from broadly neutralizing antibodies against alpha-neurotoxins.
Mabs, 17, 2025
|
|
9HUB
 
 | | D11 mAbs bound to alpha-Bungarotoxin at pH 7.5 | | Descriptor: | 1,2-ETHANEDIOL, Alpha-bungarotoxin, heavy chain, ... | | Authors: | Wade, J.W, Bohn, M.F, Laustsen, A.H, Morth, J.P. | | Deposit date: | 2024-12-21 | | Release date: | 2025-11-19 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | Rational design of antibodies with pH-dependent antigen-binding properties using structural insights from broadly neutralizing antibodies against alpha-neurotoxins.
Mabs, 17, 2025
|
|
9HXO
 
 | | A01 mAbs bound to cobratoxin at pH 6.0 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Alpha-cobratoxin, CHLORIDE ION, ... | | Authors: | Wade, J.W, Bohn, M.F, Laustsen, A.H, Morth, J.P. | | Deposit date: | 2025-01-07 | | Release date: | 2025-11-19 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Rational design of antibodies with pH-dependent antigen-binding properties using structural insights from broadly neutralizing antibodies against alpha-neurotoxins.
Mabs, 17, 2025
|
|
7KYD
 
 | | Drosophila melanogaster long-chain fatty-acyl-CoA synthetase CG6178 | | Descriptor: | 1,2-ETHANEDIOL, 5'-O-[(S)-hydroxy(octanoyloxy)phosphoryl]adenosine, Long-chain fatty-acyl-CoA synthetase CG6178 | | Authors: | Adams, S.T, Zephyr, J, Bohn, M.F, Schiffer, C.A, Miller, S.C. | | Deposit date: | 2020-12-07 | | Release date: | 2022-01-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | FruitFire: a luciferase based on a fruit fly metabolic enzyme.
Biorxiv, 2023
|
|
4IOU
 
 | |
7TCZ
 
 | | Human cytomegalovirus protease mutant (C84A, C87A, C138A, C202A) in complex with inhibitor | | Descriptor: | Assemblin, [1-(2-oxopropyl)-4-phenyl-1H-1,2,3-triazol-5-yl]methyl benzylcarbamate | | Authors: | Hulce, K.R, Bohn, M, Ongpipattanakul, C, Jaishankar, P, Renslo, A.R, Craik, C.S. | | Deposit date: | 2021-12-29 | | Release date: | 2022-01-12 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.67 Å) | | Cite: | Inhibiting a dynamic viral protease by targeting a non-catalytic cysteine.
Cell Chem Biol, 29, 2022
|
|
6EFK
 
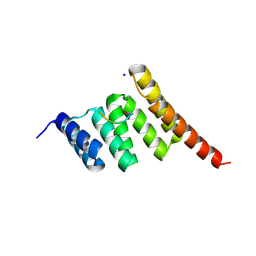 | | Crystal structure of the human CHIP TPR domain in complex with a 5mer acetylated HSP70 peptide | | Descriptor: | ACE-ILE-GLU-GLU-VAL-ASP, E3 ubiquitin-protein ligase CHIP, SODIUM ION | | Authors: | Basu, K, Ravalin, M, Bohn, M.-F, Craik, C.S, Gestwicki, J.E. | | Deposit date: | 2018-08-16 | | Release date: | 2019-07-31 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Specificity for latent C termini links the E3 ubiquitin ligase CHIP to caspases.
Nat.Chem.Biol., 15, 2019
|
|
6NSV
 
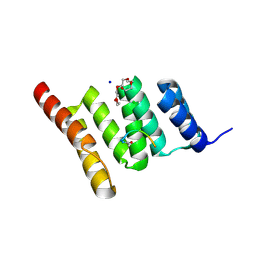 | | Crystal structure of the human CHIP TPR domain in complex with a 5mer acetylated optimized peptide | | Descriptor: | ACE-LEU-TRP-TRP-PRO-ASP, CHLORIDE ION, E3 ubiquitin-protein ligase CHIP, ... | | Authors: | Basu, K, Ravalin, M, Bohn, M.-F, Craik, C.S, Gestwicki, J.E. | | Deposit date: | 2019-01-25 | | Release date: | 2019-07-31 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.305 Å) | | Cite: | Specificity for latent C termini links the E3 ubiquitin ligase CHIP to caspases.
Nat.Chem.Biol., 15, 2019
|
|
5KEG
 
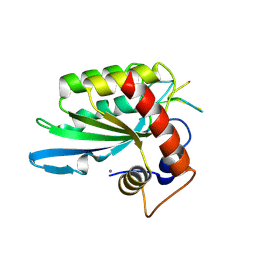 | | Crystal structure of APOBEC3A in complex with a single-stranded DNA | | Descriptor: | CALCIUM ION, CHLORIDE ION, DNA (5'-D(*TP*TP*CP*TP*T)-3'), ... | | Authors: | Kouno, T, Hilbert, B.J, Silvas, T, Royer, W.E, Matsuo, H, Schiffer, C.A. | | Deposit date: | 2016-06-09 | | Release date: | 2017-05-10 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of APOBEC3A bound to single-stranded DNA reveals structural basis for cytidine deamination and specificity.
Nat Commun, 8, 2017
|
|
5V5D
 
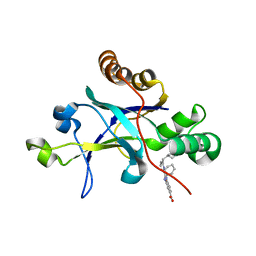 | | Room temperature (280K) crystal structure of Kaposi's sarcoma-associated herpesvirus protease in complex with allosteric inhibitor (compound 250) | | Descriptor: | 4-{[6-(cyclohexylmethyl)pyridine-2-carbonyl]amino}-3-(phenylamino)benzoic acid, ORF 17 | | Authors: | Thompson, M.C, Acker, T.M, Fraser, J.S, Craik, C.S. | | Deposit date: | 2017-03-14 | | Release date: | 2017-04-12 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.104 Å) | | Cite: | Allosteric Inhibitors, Crystallography, and Comparative Analysis Reveal Network of Coordinated Movement across Human Herpesvirus Proteases.
J. Am. Chem. Soc., 139, 2017
|
|
5V5E
 
 | | Room temperature (280K) crystal structure of Kaposi's sarcoma-associated herpesvirus protease in complex with allosteric inhibitor (compound 733) | | Descriptor: | 4-{[6-(cyclohexylmethyl)pyridine-2-carbonyl]amino}-3-{[3-(trifluoromethoxy)phenyl]amino}benzoic acid, ORF 17 | | Authors: | Thompson, M.C, Acker, T.M, Fraser, J.S, Craik, C.S. | | Deposit date: | 2017-03-14 | | Release date: | 2017-04-12 | | Last modified: | 2024-12-25 | | Method: | X-RAY DIFFRACTION (2.299 Å) | | Cite: | Allosteric Inhibitors, Crystallography, and Comparative Analysis Reveal Network of Coordinated Movement across Human Herpesvirus Proteases.
J. Am. Chem. Soc., 139, 2017
|
|
8FYU
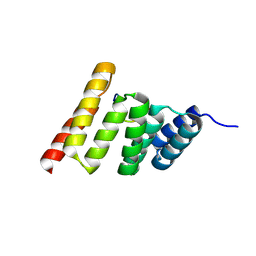 
 | | Crystal structure of the human CHIP-TPR domain in complex with a 10mer acetylated tau peptide | | Descriptor: | ACE-SER-SER-THR-GLY-SER-ILE-ASP-MET-VAL-ASP, E3 ubiquitin-protein ligase CHIP | | Authors: | Wucherer, K, Bohn, M.F, Basu, K, Nadel, C.M, Gestwicki, J.E, Craik, C.S. | | Deposit date: | 2023-01-26 | | Release date: | 2023-08-30 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.84839141 Å) | | Cite: | Phosphorylation of tau at a single residue inhibits binding to the E3 ubiquitin ligase, CHIP.
Nat Commun, 15, 2024
|
|
7KKH
 
 | | P1A4 Fab in complex with ARS1620 | | Descriptor: | (S)-1-{4-[6-chloro-8-fluoro-7-(2-fluoro-6-hydroxyphenyl)quinazolin-4-yl] piperazin-1-yl}propan-1-one, P1A4 Fab Heavy Chain, P1A4 Fab Light Chain, ... | | Authors: | Basu, K, Rohweder, P.J, Zhang, Z, Bohn, M.-F, Shokat, K, Craik, C.S. | | Deposit date: | 2020-10-27 | | Release date: | 2022-04-27 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A covalent inhibitor of K-Ras(G12C) induces MHC class I presentation of haptenated peptide neoepitopes targetable by immunotherapy.
Cancer Cell, 40, 2022
|
|
8GCK
 
 | | Crystal structure of the human CHIP-TPR domain in complex with a 6mer acetylated tau peptide | | Descriptor: | ACE-SER-ILE-ASP-MET-VAL-ASP, E3 ubiquitin-protein ligase CHIP | | Authors: | Wucherer, K, Bohn, M.F, Basu, K, Nadel, C.M, Gestwicki, J.E, Craik, C.S. | | Deposit date: | 2023-03-02 | | Release date: | 2024-03-06 | | Last modified: | 2025-10-01 | | Method: | X-RAY DIFFRACTION (1.36823535 Å) | | Cite: | Intersecting PTMS regulate clearance of pathogenic tau by the CHIP ubiquitin ligase
Alzheimer's Dement., 18, 2022
|
|
