8QJZ
 
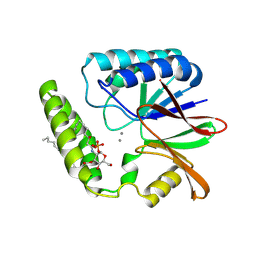 | | Crystal structure of E. coli LpxH in complex with lipid X | | Descriptor: | (R)-((2R,3S,4R,5R,6R)-3-HYDROXY-2-(HYDROXYMETHYL)-5-((R)-3-HYDROXYTETRADECANAMIDO)-6-(PHOSPHONOOXY)TETRAHYDRO-2H-PYRAN-4-YL) 3-HYDROXYTETRADECANOATE, MANGANESE (II) ION, UDP-2,3-diacylglucosamine hydrolase | | Authors: | Sooriyaarachchi, S, Bergfors, T, Jones, T.A, Mowbray, S.L. | | Deposit date: | 2023-09-14 | | Release date: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Antibiotic class with potent in vivo activity targeting lipopolysaccharide synthesis in Gram-negative bacteria.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8QKA
 
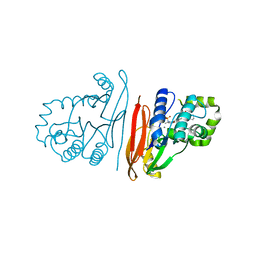 | | Structure of K. pneumoniae LpxH in complex with JEDI-852 | | Descriptor: | 2-[methyl(methylsulfonyl)amino]-~{N}-(4-piperidin-1-ylsulfonylphenyl)benzamide, MANGANESE (II) ION, UDP-2,3-diacylglucosamine hydrolase | | Authors: | Sooriyaarachchi, S, Bergfors, T, Jones, T.A, Mowbray, S.L. | | Deposit date: | 2023-09-14 | | Release date: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Antibiotic class with potent in vivo activity targeting lipopolysaccharide synthesis in Gram-negative bacteria.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8QK9
 
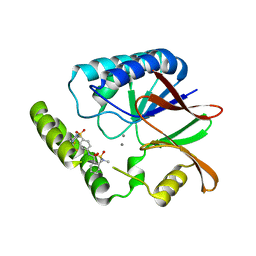 | | Structure of E. coli LpxH in complex with JEDI-1444 | | Descriptor: | 2-[methyl(methylsulfonyl)amino]-~{N}-[4-[4-[3-(trifluoromethyl)phenyl]piperazin-1-yl]sulfonylphenyl]benzamide, MANGANESE (II) ION, UDP-2,3-diacylglucosamine hydrolase | | Authors: | Sooriyaarachchi, S, Bergfors, T, Jones, T.A, Mowbray, S.L. | | Deposit date: | 2023-09-14 | | Release date: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Antibiotic class with potent in vivo activity targeting lipopolysaccharide synthesis in Gram-negative bacteria.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
4JRZ
 
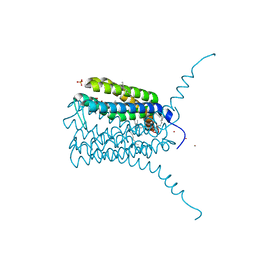 | | Human LTC4 synthase in complex with product analogs - implications for enzyme catalysis | | Descriptor: | Leukotriene C4 synthase, NICKEL (II) ION, PALMITIC ACID, ... | | Authors: | Niegowski, D, Rinaldo-Matthis, A, Haeggstrom, J.Z. | | Deposit date: | 2013-03-22 | | Release date: | 2014-01-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structures of Leukotriene C4 Synthase in Complex with Product Analogs: IMPLICATIONS FOR THE ENZYME MECHANISM.
J.Biol.Chem., 289, 2014
|
|
6YVC
 
 | |
6H3N
 
 | |
7RT7
 
 | | Crystal structure of the RhsP2 C-terminal toxin domain in complex with its immunity protein, RhsI2 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, RhsI2, RhsP2 | | Authors: | Bullen, N.P, Prehna, G, Whitney, J.C. | | Deposit date: | 2021-08-12 | | Release date: | 2022-08-17 | | Last modified: | 2023-03-29 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | An ADP-ribosyltransferase toxin kills bacterial cells by modifying structured non-coding RNAs.
Mol.Cell, 82, 2022
|
|
5I7J
 
 | |
5I7K
 
 | | Crystal Structure of Human SPLUNC1 Dolphin Mutant D1 (G58A, S61A, G62E, G63D, G66D, I67T) | | Descriptor: | BPI fold-containing family A member 1 | | Authors: | Walton, W.G, Redinbo, M.R. | | Deposit date: | 2016-02-17 | | Release date: | 2016-05-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.552 Å) | | Cite: | Structural Features Essential to the Antimicrobial Functions of Human SPLUNC1.
Biochemistry, 55, 2016
|
|
5I7L
 
 | |
4J7T
 
 | | Human LTC4 synthase in complex with product analogs - implications for enzyme catalysis | | Descriptor: | D-gamma-glutamyl-S-(4-phenylbutyl)-L-cysteinylglycine, Leukotriene C4 synthase, NICKEL (II) ION, ... | | Authors: | Niegowski, D, Rinaldo-Matthis, A, Haeggstrom, J.Z. | | Deposit date: | 2013-02-14 | | Release date: | 2014-01-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.203 Å) | | Cite: | Crystal Structures of Leukotriene C4 Synthase in Complex with Product Analogs: IMPLICATIONS FOR THE ENZYME MECHANISM.
J.Biol.Chem., 289, 2014
|
|
4J7Y
 
 | | Human LTC4 synthase in complex with product analogs - implications for enzyme catalysis | | Descriptor: | D-gamma-glutamyl-(Z)-N-(carboxymethylidene)-S-[(2R)-2-hydroxy-4-phenylbutyl]-L-cysteinamide, Leukotriene C4 synthase, NICKEL (II) ION, ... | | Authors: | Niegowski, D, Rinaldo-Matthis, A, Haeggstrom, J.Z. | | Deposit date: | 2013-02-14 | | Release date: | 2014-01-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.901 Å) | | Cite: | Crystal Structures of Leukotriene C4 Synthase in Complex with Product Analogs: IMPLICATIONS FOR THE ENZYME MECHANISM.
J.Biol.Chem., 289, 2014
|
|
6SSR
 
 | |
4JC7
 
 | | Human LTC4 synthase in complex with product analogs - implications for enzyme catalysis | | Descriptor: | Leukotriene C4 synthase, NICKEL (II) ION, PALMITIC ACID, ... | | Authors: | Niegowski, D, Rinaldo-Matthis, A, Haeggstrom, J.Z. | | Deposit date: | 2013-02-21 | | Release date: | 2014-01-01 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structures of Leukotriene C4 Synthase in Complex with Product Analogs: IMPLICATIONS FOR THE ENZYME MECHANISM.
J.Biol.Chem., 289, 2014
|
|
4JCZ
 
 | | Human LTC4 synthase in complex with product analogs - implications for enzyme catalysis | | Descriptor: | Leukotriene C4 synthase, NICKEL (II) ION, PALMITIC ACID, ... | | Authors: | Niegowski, D, Rinaldo-Matthis, A, Haeggstrom, J.Z. | | Deposit date: | 2013-02-22 | | Release date: | 2014-01-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Crystal Structures of Leukotriene C4 Synthase in Complex with Product Analogs: IMPLICATIONS FOR THE ENZYME MECHANISM.
J.Biol.Chem., 289, 2014
|
|
6SSS
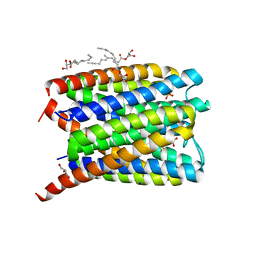 
 | | Crystal structure of Human Microsomal Glutathione S-Transferase 2 | | Descriptor: | 1-(8Z-hexadecenoyl)-sn-glycerol, GLYCEROL, Microsomal glutathione S-transferase 2, ... | | Authors: | Thulasingam, M, Nji, E, Haeggstrom, J.Z. | | Deposit date: | 2019-09-09 | | Release date: | 2021-02-03 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.498 Å) | | Cite: | Crystal structures of human MGST2 reveal synchronized conformational changes regulating catalysis.
Nat Commun, 12, 2021
|
|
6SSW
 
 | |
6SSU
 
 | |
6B12
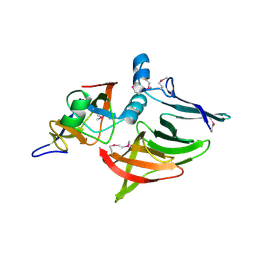 
 | | Structure of Tne2 in complex with Tni2 | | Descriptor: | Tne2, Tni2 | | Authors: | Tang, J.Y, Whitney, J.C. | | Deposit date: | 2017-09-15 | | Release date: | 2017-12-20 | | Last modified: | 2020-01-08 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Diverse NADase effector families mediate interbacterial antagonism via the type VI secretion system.
J. Biol. Chem., 293, 2018
|
|
6AXJ
 
 | |
3QMM
 
 | |
5Y81
 
 | | NuA4 TEEAA sub-complex | | Descriptor: | Actin, Actin-related protein 4, Chromatin modification-related protein EAF1, ... | | Authors: | Wang, X, Cai, G. | | Deposit date: | 2017-08-18 | | Release date: | 2018-04-18 | | Last modified: | 2019-11-06 | | Method: | ELECTRON MICROSCOPY (4.7 Å) | | Cite: | Architecture of the Saccharomyces cerevisiae NuA4/TIP60 complex
Nat Commun, 9, 2018
|
|
5YWW
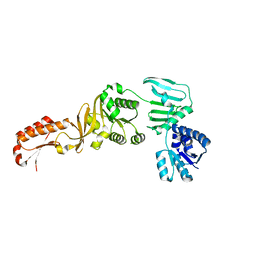 
 | | Archael RuvB-like Holiday junction helicase | | Descriptor: | GLYCEROL, Nucleotide binding protein PINc | | Authors: | Zhai, B, Yuan, Z, Han, X, DuPrez, K, Shen, Y, Fan, L. | | Deposit date: | 2017-11-30 | | Release date: | 2018-06-13 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | The archaeal ATPase PINA interacts with the helicase Hjm via its carboxyl terminal KH domain remodeling and processing replication fork and Holliday junction.
Nucleic Acids Res., 46, 2018
|
|
1T2N
 
 | |
6V2F
 
 | | Crystal structure of the HIV capsid hexamer bound to the small molecule long-acting inhibitor, GS-6207 | | Descriptor: | HIV-1 capsid, N-[(1S)-1-(3-{4-chloro-3-[(methylsulfonyl)amino]-1-(2,2,2-trifluoroethyl)-1H-indazol-7-yl}-6-[3-methyl-3-(methylsulfonyl)but-1-yn-1-yl]pyridin-2-yl)-2-(3,5-difluorophenyl)ethyl]-2-[(3bS,4aR)-5,5-difluoro-3-(trifluoromethyl)-3b,4,4a,5-tetrahydro-1H-cyclopropa[3,4]cyclopenta[1,2-c]pyrazol-1-yl]acetamide | | Authors: | Appleby, T.C, Link, J.O, Yant, S.R, Villasenor, A.G, Somoza, J.R, Hu, E.Y, Schroeder, S.D, Cihlar, T. | | Deposit date: | 2019-11-22 | | Release date: | 2020-07-01 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Clinical targeting of HIV capsid protein with a long-acting small molecule.
Nature, 584, 2020
|
|
