6A9K
 
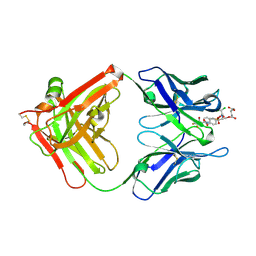 | | Crystal structure of the complex of the hydrolytic antibody Fab 9C10 with a transition-state analog | | Descriptor: | 5-[(2R,3R)-2-[2,2-bis(chloranyl)ethanoylamino]-3-(4-nitrophenyl)-3-[oxidanyl-[[4-[2,2,2-tris(fluoranyl)ethanoylamino]phenyl]methyl]phosphoryl]oxy-propoxy]-5-oxidanylidene-pentanoic acid, IMMUNOGLOBULIN 9C10 H CHAIN, IMMUNOGLOBULIN 9C10 L CHAIN | | Authors: | Tsuchiya, Y, Fujii, I, Tada, T, Yamaguchi, A, Tsumuraya, T, Kumon, A. | | Deposit date: | 2018-07-13 | | Release date: | 2019-07-24 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the complex of the hydrolytic antibody Fab 9C10 with a transition-state analog
To Be Published
|
|
6IGU
 
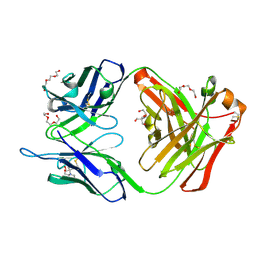 | | Crystal structure of the hydrolytic antibody Fab 9C10 | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, IMMUNOGLOBULIN 9C10 H CHAIN, IMMUNOGLOBULIN 9C10 L CHAIN, ... | | Authors: | Yamaguchi, A, Tada, T, Tsuchiya, Y, Tsumuraya, T, Fujii, I. | | Deposit date: | 2018-09-26 | | Release date: | 2019-10-09 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Crystal structure of the complex of the hydrolytic antibody Fab 9C10 and a transition-state analog
To Be Published
|
|
6I2U
 
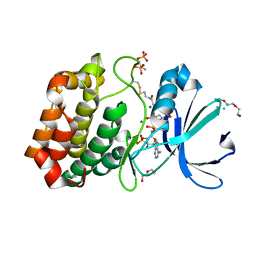 | |
6A1C
 
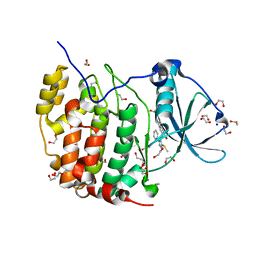 | | Crystal structure of the CK2a1-go289 complex | | Descriptor: | 1,2-ETHANEDIOL, 5-bromanyl-2-methoxy-4-[(E)-(3-methylsulfanyl-5-phenyl-1,2,4-triazol-4-yl)iminomethyl]phenol, Casein kinase II subunit alpha, ... | | Authors: | Kinoshita, T, Tsuyuguchi, M. | | Deposit date: | 2018-06-07 | | Release date: | 2019-03-06 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Cell-based screen identifies a new potent and highly selective CK2 inhibitor for modulation of circadian rhythms and cancer cell growth.
Sci Adv, 5, 2019
|
|
5WQJ
 
 | |
5WQK
 
 | |
8J6N
 | | Crystal structure of Cystathionine gamma-lyase in complex with compound 1 | | Descriptor: | 1,2-ETHANEDIOL, Cystathionine gamma-lyase, GLYCEROL, ... | | Authors: | Hibi, R, Toma-Fukai, S, Shimizu, T, Hanaoka, K. | | Deposit date: | 2023-04-26 | | Release date: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Discovery of a cystathionine gamma-lyase (CSE) selective inhibitor targeting active-site pyridoxal 5'-phosphate (PLP) via Schiff base formation.
Sci Rep, 13, 2023
|
|
5B2E
 
 | | N,N'-diacetylchitobiose deacetylase from Pyrococcus horikoshii complexed with its inhibitor MPG (acetate-containing condition) | | Descriptor: | 2-deoxy-2-{[(S)-hydroxy(methyl)phosphoryl]amino}-beta-D-glucopyranose, HEXANE-1,6-DIOL, Putative uncharacterized protein PH0499, ... | | Authors: | Nakamura, T, Niiyama, M, Ida, K, Uegaki, K. | | Deposit date: | 2016-01-15 | | Release date: | 2016-08-10 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Substrate recognition of N,N'-diacetylchitobiose deacetylase from Pyrococcus horikoshii
J.Struct.Biol., 195, 2016
|
|
5B2F
 | | N,N'-diacetylchitobiose deacetylase from Pyrococcus horikoshii complexed with its inhibitor MPG (phosphate-containing condition) | | Descriptor: | 2-deoxy-2-{[(S)-hydroxy(methyl)phosphoryl]amino}-beta-D-glucopyranose, Putative uncharacterized protein PH0499, ZINC ION | | Authors: | Nakamura, T, Niiyama, M, Ida, K, Uegaki, K. | | Deposit date: | 2016-01-15 | | Release date: | 2016-08-10 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Substrate recognition of N,N'-diacetylchitobiose deacetylase from Pyrococcus horikoshii
J.Struct.Biol., 195, 2016
|
|
3WQU
 
 | | Staphylococcus aureus FtsA complexed with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Cell division protein FtsA, MAGNESIUM ION | | Authors: | Fujita, J, Maeda, Y, Miyazaki, Y, Inoue, T, Matsumura, H. | | Deposit date: | 2014-02-01 | | Release date: | 2014-10-08 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of FtsA from Staphylococcus aureus
FEBS Lett., 588, 2014
|
|
3WQT
 
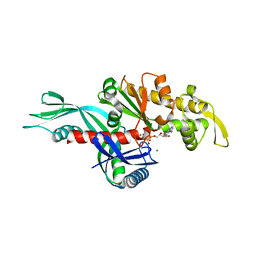 | | Staphylococcus aureus FtsA complexed with AMPPNP | | Descriptor: | CHLORIDE ION, Cell division protein FtsA, MAGNESIUM ION, ... | | Authors: | Fujita, J, Maeda, Y, Miyazaki, Y, Inoue, T, Matsumura, H. | | Deposit date: | 2014-02-01 | | Release date: | 2014-10-08 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of FtsA from Staphylococcus aureus
FEBS Lett., 588, 2014
|
|
5CBK
 
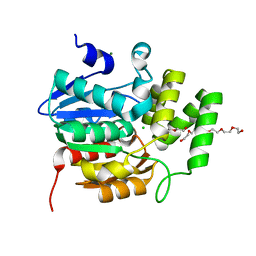 | | Crystal structure of the strigolactone receptor ShHTL5 from Striga hermonthica | | Descriptor: | 3,6,9,12,15,18,21,24,27,30,33,36,39-TRIDECAOXAHENTETRACONTANE-1,41-DIOL, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Stogios, P.J, Onopriyenko, O, Yim, V, Savchenko, A. | | Deposit date: | 2015-07-01 | | Release date: | 2015-10-21 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.462 Å) | | Cite: | Structure-function analysis identifies highly sensitive strigolactone receptors in Striga.
Science, 350, 2015
|
|
7C87
 
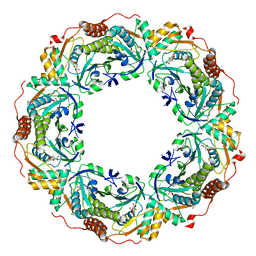 | |
7C89
 | |
7C8A
 | |
7CQJ
 | |
1Q31
 
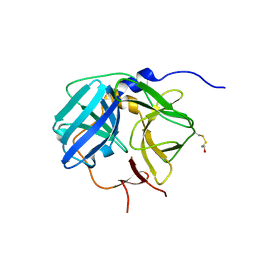 | | Crystal Structure of the Tobacco Etch Virus Protease C151A mutant | | Descriptor: | BETA-MERCAPTOETHANOL, Nuclear inclusion protein A | | Authors: | Nunn, C.M, Djordjevic, S, George, R.R, Urquhart, G.T, Chao, L.H, Tsuchiya, Y. | | Deposit date: | 2003-07-28 | | Release date: | 2004-11-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of tobacco etch virus protease shows the protein C terminus bound within the active site.
J.Mol.Biol., 350, 2005
|
|
