8ATX
 
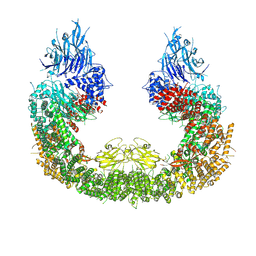 | |
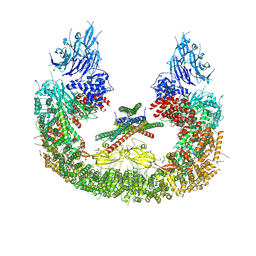
8AUW
 
 | |
8ATU
 
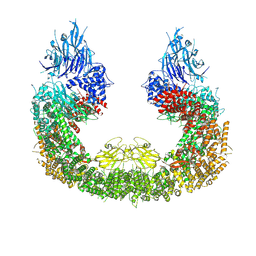 | | Cryo-EM structure of human BIRC6 | | Descriptor: | Baculoviral IAP repeat-containing protein 6, ZINC ION | | Authors: | Ehrmann, J.F, Grabarczyk, D.B, Clausen, T. | | Deposit date: | 2022-08-24 | | Release date: | 2023-02-15 | | Last modified: | 2023-03-29 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Structural basis for regulation of apoptosis and autophagy by the BIRC6/SMAC complex.
Science, 379, 2023
|
|
8AUK
 
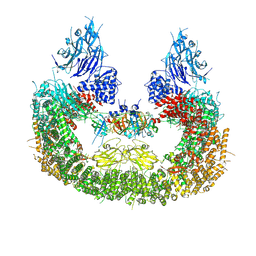 | | Cryo-EM structure of human BIRC6 in complex with HTRA2. | | Descriptor: | Baculoviral IAP repeat-containing protein 6, Serine protease HTRA2, mitochondrial, ... | | Authors: | Ehrmann, J.F, Grabarczyk, D.B, Clausen, T. | | Deposit date: | 2022-08-25 | | Release date: | 2023-02-15 | | Last modified: | 2023-05-24 | | Method: | ELECTRON MICROSCOPY (6.2 Å) | | Cite: | Structural basis for regulation of apoptosis and autophagy by the BIRC6/SMAC complex.
Science, 379, 2023
|
|
8B9O
 
 | |
8B9U
 
 | |
6QDJ
 
 | | Molecular features of the UNC-45 chaperone critical for binding and folding muscle myosin | | Descriptor: | 1,4-BUTANEDIOL, 2,5,8,11,14,17-HEXAOXANONADECAN-19-OL, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Meinhart, A, Clausen, T, Arnese, R. | | Deposit date: | 2019-01-02 | | Release date: | 2019-10-30 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.884 Å) | | Cite: | Molecular features of the UNC-45 chaperone critical for binding and folding muscle myosin.
Nat Commun, 10, 2019
|
|
6QDL
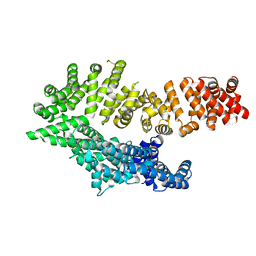 
 | |
6QDM
 
 | |
1C7N
 
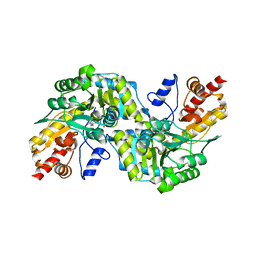 | | CRYSTAL STRUCTURE OF CYSTALYSIN FROM TREPONEMA DENTICOLA CONTAINS A PYRIDOXAL 5'-PHOSPHATE COFACTOR | | Descriptor: | CYSTALYSIN, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Krupka, H.I, Huber, R, Holt, S.C, Clausen, T. | | Deposit date: | 2000-03-16 | | Release date: | 2000-07-26 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of cystalysin from Treponema denticola: a pyridoxal 5'-phosphate-dependent protein acting as a haemolytic enzyme.
EMBO J., 19, 2000
|
|
1C7O
 
 | | CRYSTAL STRUCTURE OF CYSTALYSIN FROM TREPONEMA DENTICOLA CONTAINS A PYRIDOXAL 5'-PHOSPHATE-L-AMINOETHOXYVINYLGLYCINE COMPLEX | | Descriptor: | (2E,3E)-4-(2-aminoethoxy)-2-[({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methyl)imino]but-3-enoic acid, CYSTALYSIN | | Authors: | Krupka, H.I, Huber, R, Holt, S.C, Clausen, T. | | Deposit date: | 2000-03-16 | | Release date: | 2000-07-26 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of cystalysin from Treponema denticola: a pyridoxal 5'-phosphate-dependent protein acting as a haemolytic enzyme.
EMBO J., 19, 2000
|
|
1SOZ
 
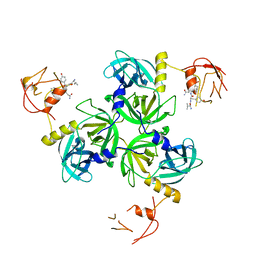 | | Crystal Structure of DegS protease in complex with an activating peptide | | Descriptor: | Protease degS, activating peptide | | Authors: | Wilken, C, Kitzing, K, Kurzbauer, R, Ehrmann, M, Clausen, T. | | Deposit date: | 2004-03-16 | | Release date: | 2004-06-08 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the DegS stress sensor: How a PDZ domain recognizes misfolded protein and activates a protease
Cell(Cambridge,Mass.), 117, 2004
|
|
1SOT
 
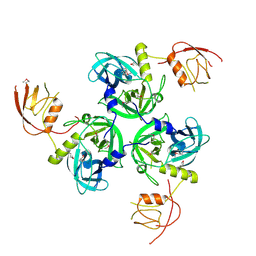 | | Crystal Structure of the DegS stress sensor | | Descriptor: | Protease degS | | Authors: | Wilken, C, Kitzing, K, Kurzbauer, R, Ehrmann, M, Clausen, T. | | Deposit date: | 2004-03-15 | | Release date: | 2004-06-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the DegS stress sensor: How a PDZ domain recognizes misfolded protein and activates a protease
Cell(Cambridge,Mass.), 117, 2004
|
|
7OIK
 
 | |
7OIM
 
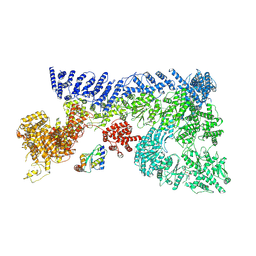
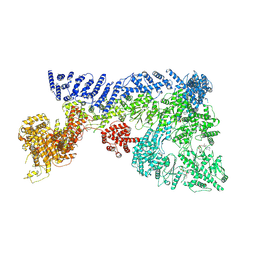 | | Mouse RNF213, with mixed nucleotides bound | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, E3 ubiquitin-protein ligase RNF213, ... | | Authors: | Grabarczyk, D, Ahel, J, Clausen, T. | | Deposit date: | 2021-05-11 | | Release date: | 2022-06-01 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | E3 ubiquitin ligase RNF213 employs a non-canonical zinc finger active site and is allosterically regulated by ATP
To Be Published
|
|
6QDK
 
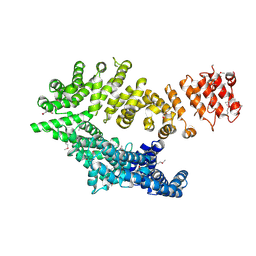 | |
3NWU
 
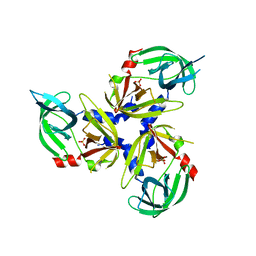 | | Substrate induced remodeling of the active site regulates HtrA1 activity | | Descriptor: | SULFATE ION, Serine protease HTRA1 | | Authors: | Truebestein, L, Tennstaedt, A, Hauske, P, Krojer, T, Kaiser, M, Clausen, T, Ehrmann, M. | | Deposit date: | 2010-07-11 | | Release date: | 2011-02-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Substrate-induced remodeling of the active site regulates human HTRA1 activity.
Nat.Struct.Mol.Biol., 18, 2011
|
|
3NUM
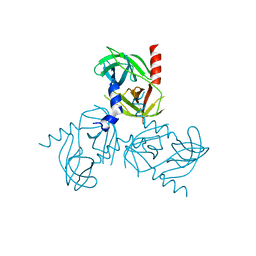 
 | | Substrate induced remodeling of the active site regulates HtrA1 activity | | Descriptor: | Serine protease HTRA1 | | Authors: | Truebestein, L, Tennstaedt, A, Hauske, P, Krojer, T, Kaiser, M, Clausen, T, Ehrmann, M. | | Deposit date: | 2010-07-07 | | Release date: | 2011-02-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Substrate-induced remodeling of the active site regulates human HTRA1 activity.
Nat.Struct.Mol.Biol., 18, 2011
|
|
3NZI
 
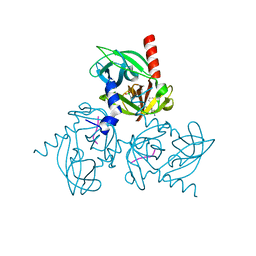 | | Substrate induced remodeling of the active site regulates HtrA1 activity | | Descriptor: | Citrate synthase, Serine protease HTRA1 | | Authors: | Truebestein, L, Tennstaedt, A, Hauske, P, Krojer, T, Kaiser, M, Clausen, T, Ehrmann, M. | | Deposit date: | 2010-07-16 | | Release date: | 2011-02-23 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Substrate-induced remodeling of the active site regulates human HTRA1 activity.
Nat.Struct.Mol.Biol., 18, 2011
|
|
6ZK7
 
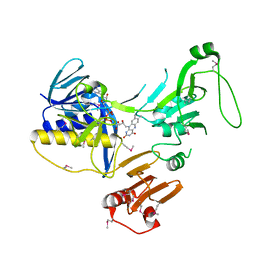 | | Crystal Structure of human PYROXD1/FAD complex | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Pyridine nucleotide-disulfide oxidoreductase domain-containing protein 1 | | Authors: | Meinhart, A, Asanovic, I, Martinez, J, Clausen, T. | | Deposit date: | 2020-06-30 | | Release date: | 2021-05-12 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | The oxidoreductase PYROXD1 uses NAD(P) + as an antioxidant to sustain tRNA ligase activity in pre-tRNA splicing and unfolded protein response.
Mol.Cell, 81, 2021
|
|
4A8D
 
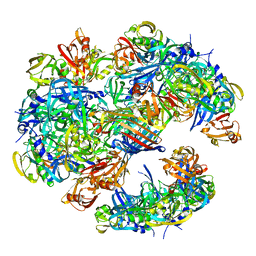 | | DegP dodecamer with bound OMP | | Descriptor: | OUTER MEMBRANE PROTEIN C, PERIPLASMIC SERINE ENDOPROTEASE DEGP | | Authors: | Malet, H, Krojer, T, Sawa, J, Schafer, E, Saibil, H.R, Ehrmann, M, Clausen, T. | | Deposit date: | 2011-11-20 | | Release date: | 2012-01-11 | | Last modified: | 2017-08-30 | | Method: | ELECTRON MICROSCOPY (28 Å) | | Cite: | Newly Folded Substrates Inside the Molecular Cage of the Htra Chaperone Degq
Nat.Struct.Mol.Biol., 19, 2012
|
|
3MH4
 
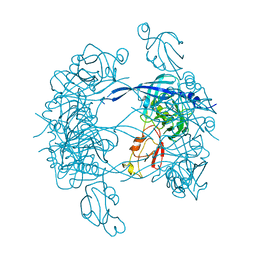 | |
3MH5
 
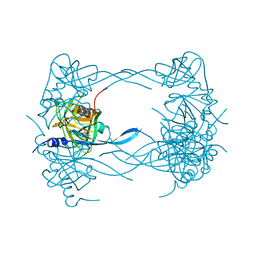 | | HtrA proteases are activated by a conserved mechanism that can be triggered by distinct molecular cues | | Descriptor: | DIISOPROPYL PHOSPHONATE, Protease do | | Authors: | Krojer, T, Sawa, J, Huber, R, Clausen, T. | | Deposit date: | 2010-04-07 | | Release date: | 2010-06-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | HtrA proteases have a conserved activation mechanism that can be triggered by distinct molecular cues
Nat.Struct.Mol.Biol., 17, 2010
|
|
3MH6
 
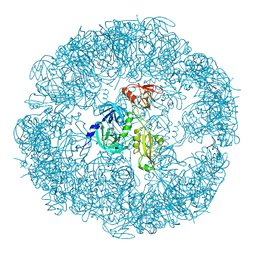 | | HtrA proteases are activated by a conserved mechanism that can be triggered by distinct molecular cues | | Descriptor: | DIISOPROPYL PHOSPHONATE, Protease do | | Authors: | Krojer, T, Sawa, J, Huber, R, Clausen, T. | | Deposit date: | 2010-04-07 | | Release date: | 2010-06-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | HtrA proteases have a conserved activation mechanism that can be triggered by distinct molecular cues
Nat.Struct.Mol.Biol., 17, 2010
|
|
3MH7
 
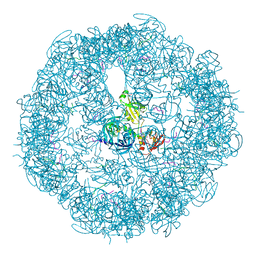 | | HtrA proteases are activated by a conserved mechanism that can be triggered by distinct molecular cues | | Descriptor: | 5-mer peptide, Protease do | | Authors: | Krojer, T, Sawa, J, Huber, R, Clausen, T. | | Deposit date: | 2010-04-07 | | Release date: | 2010-06-30 | | Last modified: | 2014-03-05 | | Method: | X-RAY DIFFRACTION (2.961 Å) | | Cite: | HtrA proteases have a conserved activation mechanism that can be triggered by distinct molecular cues
Nat.Struct.Mol.Biol., 17, 2010
|
|
