7TOL
 
 | | X-ray crystal structure of glycerol dibiphytanyl glycerol tetraether - macrocyclic archaeol synthase (GDGT-MAS) from Methanocaldococcus jannaschii with archaeal lipid, 5'deoxyadenosine, and methionine bound | | Descriptor: | 3-PHOSPHORYL-[1,2-DI-PHYTANYL]GLYCEROL, 3-[GLYCEROLYLPHOSPHONYL]-[1,2-DI-PHYTANYL]GLYCEROL, 5'-DEOXYADENOSINE, ... | | Authors: | Lloyd, C.T, Booker, S.J, Boal, A.K. | | Deposit date: | 2022-01-24 | | Release date: | 2022-08-31 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Discovery, structure and mechanism of a tetraether lipid synthase.
Nature, 609, 2022
|
|
7TOM
 
 | | X-ray crystal structure of glycerol dibiphytanyl glycerol tetraether - macrocyclic archaeol synthase (GDGT-MAS) from Methanocaldococcus jannaschii with bacterial lipid substrate analog, 5'deoxyadenosine, and methionine bound | | Descriptor: | 2-(HEXADECANOYLOXY)-1-[(PHOSPHONOOXY)METHYL]ETHYL HEXADECANOATE, 5'-DEOXYADENOSINE, FE (III) ION, ... | | Authors: | Lloyd, C.T, Booker, S.J, Boal, A.K. | | Deposit date: | 2022-01-24 | | Release date: | 2022-08-31 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Discovery, structure and mechanism of a tetraether lipid synthase.
Nature, 609, 2022
|
|
4YDX
 
 | | Crystal structure of cisplatin bound to a human copper chaperone (monomer) - new refinement | | Descriptor: | 3,3',3''-phosphanetriyltripropanoic acid, Copper transport protein ATOX1, PLATINUM (II) ION, ... | | Authors: | Shabalin, I.G, Boal, A.K, Dauter, Z, Jaskolski, M, Minor, W, Rosenzweig, A.C, Wlodawer, A. | | Deposit date: | 2015-02-23 | | Release date: | 2015-03-04 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.602 Å) | | Cite: | Crystallography and chemistry should always go together: a cautionary tale of protein complexes with cisplatin and carboplatin.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
7TWA
 | | Crystal structure of apo BesC from Streptomyces cattleya | | Descriptor: | 1,3-BUTANEDIOL, 4-chloro-allylglycine synthase, ACETATE ION, ... | | Authors: | Neugebauer, M.E, McBride, M.J, Boal, A.K, Chang, M.C.Y. | | Deposit date: | 2022-02-07 | | Release date: | 2022-04-13 | | Last modified: | 2022-04-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Substrate-Triggered mu-Peroxodiiron(III) Intermediate in the 4-Chloro-l-Lysine-Fragmenting Heme-Oxygenase-like Diiron Oxidase (HDO) BesC: Substrate Dissociation from, and C4 Targeting by, the Intermediate.
Biochemistry, 61, 2022
|
|
4D8G
 
 | | Chlamydia trachomatis NrdB with a Mn/Fe cofactor (procedure 2 - low Mn) | | Descriptor: | FE (III) ION, MANGANESE (II) ION, Ribonucleoside-diphosphate reductase subunit beta | | Authors: | Dassama, L.M.K, Boal, A.K, Krebs, C, Rosenzweig, A.C, Bollinger Jr, J.M. | | Deposit date: | 2012-01-10 | | Release date: | 2012-02-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Evidence that the beta subunit of Chlamydia trachomatis ribonucleotide reductase is active with the manganese ion of its manganese(IV)/iron(III) cofactor in site 1.
J.Am.Chem.Soc., 134, 2012
|
|
5V7O
 
 | | Crystal Structure of NosK from Streptomyces actuosus | | Descriptor: | NosK, SULFATE ION | | Authors: | Booker, S.J, Boal, A.K, Grove, T.L, Badding, E.D. | | Deposit date: | 2017-03-20 | | Release date: | 2017-04-12 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Rerouting the Pathway for the Biosynthesis of the Side Ring System of Nosiheptide: The Roles of NosI, NosJ, and NosK.
J. Am. Chem. Soc., 139, 2017
|
|
5UY1
 
 | | X-ray crystal structure of apo Halotag | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Haloalkane dehalogenase | | Authors: | Dunham, N.P, Boal, A.K. | | Deposit date: | 2017-02-23 | | Release date: | 2017-03-08 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | The Cation-pi Interaction Enables a Halo-Tag Fluorogenic Probe for Fast No-Wash Live Cell Imaging and Gel-Free Protein Quantification.
Biochemistry, 56, 2017
|
|
5UXZ
 | |
5VNP
 | |
5HR6
 
 | | X-ray crystal structure of C118A RlmN with cross-linked tRNA purified from Escherichia coli | | Descriptor: | 5'-DEOXYADENOSINE, IRON/SULFUR CLUSTER, MAGNESIUM ION, ... | | Authors: | Schwalm, E.L, Grove, T.L, Booker, S.J, Boal, A.K. | | Deposit date: | 2016-01-22 | | Release date: | 2016-04-13 | | Last modified: | 2016-04-27 | | Method: | X-RAY DIFFRACTION (2.88 Å) | | Cite: | Crystallographic capture of a radical S-adenosylmethionine enzyme in the act of modifying tRNA.
Science, 352, 2016
|
|
5HR7
 | | X-ray crystal structure of C118A RlmN from Escherichia coli with cross-linked in vitro transcribed tRNA | | Descriptor: | 5'-DEOXYADENOSINE, Dual-specificity RNA methyltransferase RlmN, IRON/SULFUR CLUSTER, ... | | Authors: | Schwalm, E.L, Grove, T.L, Booker, S.J, Boal, A.K. | | Deposit date: | 2016-01-22 | | Release date: | 2016-04-13 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystallographic capture of a radical S-adenosylmethionine enzyme in the act of modifying tRNA.
Science, 352, 2016
|
|
5IQV
 
 | | WelO5 bound to Fe, Cl, 2-oxoglutarate, 12-epifischerindole U, and nitric oxide | | Descriptor: | (6aS,9R,10R,10aS)-9-ethyl-10-isocyano-6,6,9-trimethyl-5,6,6a,7,8,9,10,10a-octahydroindeno[2,1-b]indole, 2-OXOGLUTARIC ACID, CHLORIDE ION, ... | | Authors: | Mitchell, A.J, Boal, A.K. | | Deposit date: | 2016-03-11 | | Release date: | 2016-06-29 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for halogenation by iron- and 2-oxo-glutarate-dependent enzyme WelO5.
Nat.Chem.Biol., 12, 2016
|
|
5IQT
 
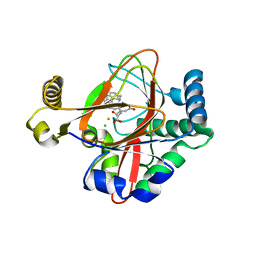 | | WelO5 bound to Fe(II), Cl, 2-oxoglutarate, and 12-epifischerindole U | | Descriptor: | (6aS,9R,10R,10aS)-9-ethyl-10-isocyano-6,6,9-trimethyl-5,6,6a,7,8,9,10,10a-octahydroindeno[2,1-b]indole, 2-OXOGLUTARIC ACID, CHLORIDE ION, ... | | Authors: | Mitchell, A.J, Boal, A.K. | | Deposit date: | 2016-03-11 | | Release date: | 2016-06-29 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for halogenation by iron- and 2-oxo-glutarate-dependent enzyme WelO5.
Nat.Chem.Biol., 12, 2016
|
|
5IQU
 
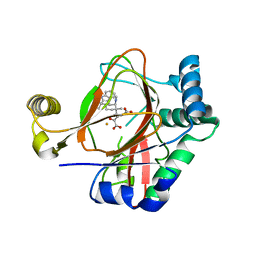 | | WelO5 G166D variant bound to Fe(II), 2-oxoglutarate, and 12-epifischerindole U | | Descriptor: | (6aS,9R,10R,10aS)-9-ethyl-10-isocyano-6,6,9-trimethyl-5,6,6a,7,8,9,10,10a-octahydroindeno[2,1-b]indole, 2-OXOGLUTARIC ACID, FE (II) ION, ... | | Authors: | Mitchell, A.J, Maggiolo, A.O, Boal, A.K. | | Deposit date: | 2016-03-11 | | Release date: | 2016-06-29 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Structural basis for halogenation by iron- and 2-oxo-glutarate-dependent enzyme WelO5.
Nat.Chem.Biol., 12, 2016
|
|
5IQS
 
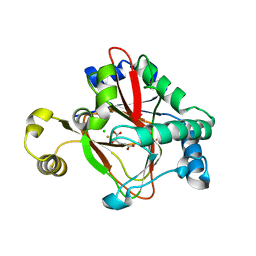 | | WelO5 bound to Fe(II), Cl, and 2-oxoglutarate | | Descriptor: | 2-OXOGLUTARIC ACID, CHLORIDE ION, FE (II) ION, ... | | Authors: | Mitchell, A.J, Ananth, N, Boal, A.K. | | Deposit date: | 2016-03-11 | | Release date: | 2016-06-29 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis for halogenation by iron- and 2-oxo-glutarate-dependent enzyme WelO5.
Nat.Chem.Biol., 12, 2016
|
|
6ONP
 
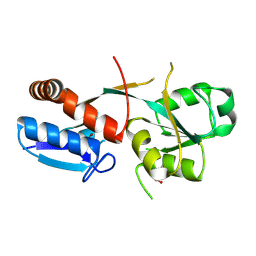 | |
6ONQ
 
 | |
6OXJ
 
 | | X-ray crystal structure of Y140F FtmOx1 bound to Fe(II) | | Descriptor: | FE (II) ION, Verruculogen synthase | | Authors: | Dunham, N.P, Boal, A.K. | | Deposit date: | 2019-05-13 | | Release date: | 2019-07-03 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Hydrogen Donation but not Abstraction by a Tyrosine (Y68) during Endoperoxide Installation by Verruculogen Synthase (FtmOx1).
J.Am.Chem.Soc., 141, 2019
|
|
6OXH
 
 | |
6P5Q
 
 | | X-ray structure of Fe(II)-soaked UndA bound to lauric acid | | Descriptor: | FE (III) ION, GLYCEROL, LAURIC ACID, ... | | Authors: | Rajakovich, L.J, Mitchell, A.J, Boal, A.K. | | Deposit date: | 2019-05-30 | | Release date: | 2019-09-18 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Substrate-Triggered Formation of a Peroxo-Fe2(III/III) Intermediate during Fatty Acid Decarboxylation by UndA.
J.Am.Chem.Soc., 141, 2019
|
|
6M9S
 
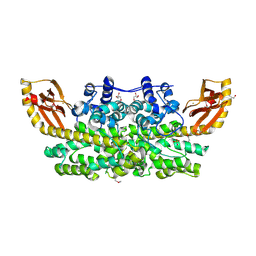 | | Crystal structure of SeMet SznF from Streptomyces achromogenes var. streptozoticus NRRL 2697 | | Descriptor: | FE (III) ION, GLYCEROL, ISOPROPYL ALCOHOL, ... | | Authors: | Rohac, R, Mitchell, A.J, Boal, A.K. | | Deposit date: | 2018-08-24 | | Release date: | 2019-02-06 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | An N-nitrosating metalloenzyme constructs the pharmacophore of streptozotocin.
Nature, 566, 2019
|
|
6M9R
 
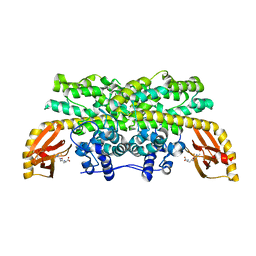 | |
6MP8
 
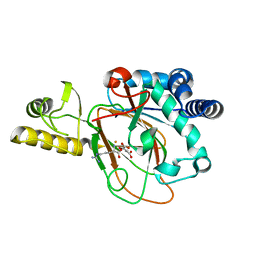 | | X-ray crystal structure of VioC bound to Fe(II), D-arginine, and 2-oxoglutarate | | Descriptor: | 2-OXOGLUTARIC ACID, Alpha-ketoglutarate-dependent L-arginine hydroxylase, D-ARGININE, ... | | Authors: | Dunham, N.P, Boal, A.K. | | Deposit date: | 2018-10-05 | | Release date: | 2018-11-21 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | alpha-Amine Desaturation of d-Arginine by the Iron(II)- and 2-(Oxo)glutarate-Dependent l-Arginine 3-Hydroxylase, VioC.
Biochemistry, 57, 2018
|
|
6MP9
 
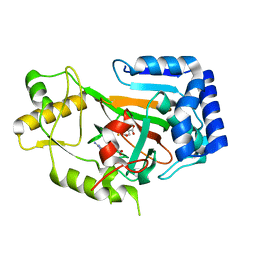 | | X-ray crystal structure of VioC bound to Fe(II), 2-oxo-5-guanidinopentanoic acid, and succinate | | Descriptor: | 5-carbamimidamido-2-oxopentanoic acid, Alpha-ketoglutarate-dependent L-arginine hydroxylase, FE (II) ION, ... | | Authors: | Dunham, N.P, Boal, A.K. | | Deposit date: | 2018-10-05 | | Release date: | 2018-11-21 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | alpha-Amine Desaturation of d-Arginine by the Iron(II)- and 2-(Oxo)glutarate-Dependent l-Arginine 3-Hydroxylase, VioC.
Biochemistry, 57, 2018
|
|
6NPA
 
 | | X-ray crystal structure of TmpB, (R)-1-hydroxy-2-trimethylaminoethylphosphonate oxygenase, with (R)-1-hydroxy-2-trimethylaminoethylphosphonate | | Descriptor: | (2R)-2-hydroxy-N,N,N-trimethyl-2-phosphonoethan-1-aminium, FE (II) ION, FE (III) ION, ... | | Authors: | Rajakovich, L.J, Mitchell, A.J, Boal, A.K. | | Deposit date: | 2019-01-17 | | Release date: | 2019-03-06 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | A New Microbial Pathway for Organophosphonate Degradation Catalyzed by Two Previously Misannotated Non-Heme-Iron Oxygenases.
Biochemistry, 58, 2019
|
|
