1DCH
 
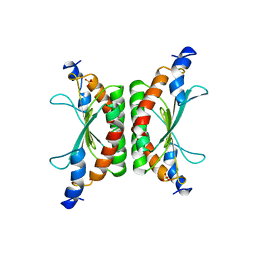 | | CRYSTAL STRUCTURE OF DCOH, A BIFUNCTIONAL, PROTEIN-BINDING TRANSCRIPTION COACTIVATOR | | Descriptor: | DCOH (DIMERIZATION COFACTOR OF HNF-1), SULFATE ION | | Authors: | Endrizzi, J.A, Cronk, J.D, Wang, W, Crabtree, G.R, Alber, T. | | Deposit date: | 1995-01-24 | | Release date: | 1996-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of DCoH, a bifunctional, protein-binding transcriptional coactivator.
Science, 268, 1995
|
|
1DEB
 
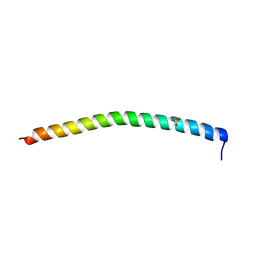 | |
1EKX
 
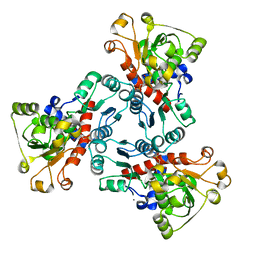 | | THE ISOLATED, UNREGULATED CATALYTIC TRIMER OF ASPARTATE TRANSCARBAMOYLASE COMPLEXED WITH BISUBSTRATE ANALOG PALA (N-(PHOSPHONACETYL)-L-ASPARTATE) | | Descriptor: | ASPARTATE TRANSCARBAMOYLASE, CALCIUM ION, N-(PHOSPHONACETYL)-L-ASPARTIC ACID | | Authors: | Endrizzi, J.A, Beernink, P.T, Alber, T, Schachman, H.K. | | Deposit date: | 2000-03-09 | | Release date: | 2000-05-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Binding of bisubstrate analog promotes large structural changes in the unregulated catalytic trimer of aspartate transcarbamoylase: implications for allosteric regulation induced cell migration.
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
6CCD
 
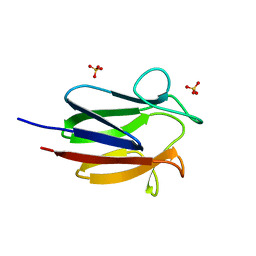 | |
1RU0
 
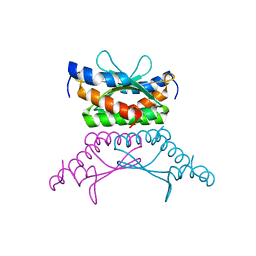 | | Crystal structure of DCoH2, a paralog of DCoH, the Dimerization Cofactor of HNF-1 | | Descriptor: | DcoH-like protein DCoHm | | Authors: | Rose, R.B, Pullen, K.E, Bayle, J.H, Crabtree, G.R, Alber, T. | | Deposit date: | 2003-12-10 | | Release date: | 2004-10-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Biochemical and structural basis for partially redundant enzymatic and transcriptional functions of DCoH and DCoH2
Biochemistry, 43, 2004
|
|
1TGG
 
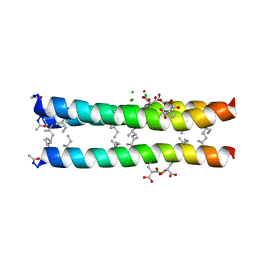 | | RH3 DESIGNED RIGHT-HANDED COILED COIL TRIMER | | Descriptor: | CHLORIDE ION, NICKEL (II) ION, right-handed coiled coil trimer | | Authors: | Plecs, J.J, Harbury, P.B, Kim, P.S, Alber, T. | | Deposit date: | 2004-05-28 | | Release date: | 2004-10-12 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural test of the parameterized-backbone method for protein design
J.Mol.Biol., 342, 2004
|
|
2NT4
 
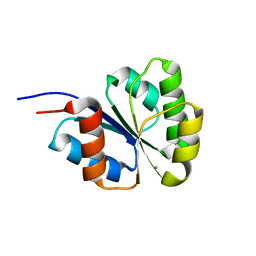 | | Receiver domain from Myxococcus xanthus social motility protein FrzS (H92F mutant) | | Descriptor: | CHLORIDE ION, Response regulator homolog | | Authors: | Echols, N, Fraser, J, Weisfield, S, Merlie, J, Zusman, D, Alber, T. | | Deposit date: | 2006-11-06 | | Release date: | 2007-03-13 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.02 Å) | | Cite: | An atypical receiver domain controls the dynamic polar localization of the Myxococcus xanthus social motility protein FrzS.
Mol.Microbiol., 65, 2007
|
|
2NT3
 
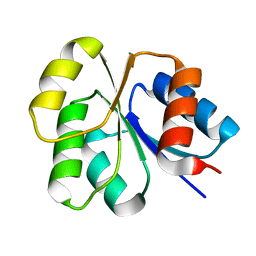 | | Receiver domain from Myxococcus xanthus social motility protein FrzS (Y102A Mutant) | | Descriptor: | Response regulator homolog | | Authors: | Fraser, J.S, Echols, N, Merlie, J.P, Zusman, D.R, Alber, T. | | Deposit date: | 2006-11-06 | | Release date: | 2007-03-13 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | An atypical receiver domain controls the dynamic polar localization of the Myxococcus xanthus social motility protein FrzS.
Mol.Microbiol., 65, 2007
|
|
3RMR
 
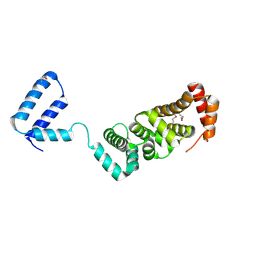 | |
3TGP
 
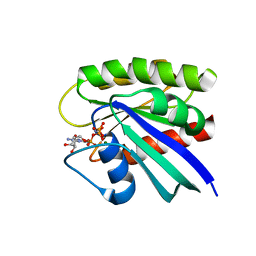 | | Room temperature H-ras | | Descriptor: | GTPase HRas, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER | | Authors: | Fraser, J.S, Alber, T. | | Deposit date: | 2011-08-17 | | Release date: | 2011-10-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.3075 Å) | | Cite: | Accessing protein conformational ensembles using room-temperature X-ray crystallography.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3CSU
 
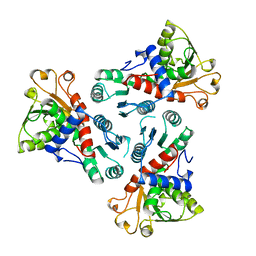 | | CATALYTIC TRIMER OF ESCHERICHIA COLI ASPARTATE TRANSCARBAMOYLASE | | Descriptor: | CALCIUM ION, PROTEIN (ASPARTATE CARBAMOYLTRANSFERASE) | | Authors: | Beernink, P.T, Endrizzi, J.A, Alber, T, Schachman, H.K. | | Deposit date: | 1999-04-22 | | Release date: | 1999-05-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Assessment of the allosteric mechanism of aspartate transcarbamoylase based on the crystalline structure of the unregulated catalytic subunit.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
3F61
 
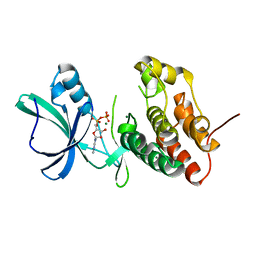 | |
3GOF
 
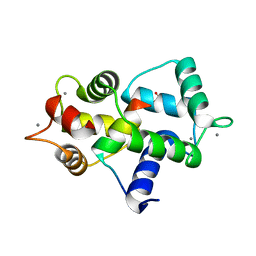 | | Calmodulin bound to peptide from macrophage nitric oxide synthase | | Descriptor: | CALCIUM ION, Calmodulin, Nitric oxide synthase, ... | | Authors: | Ng, H.L, Greenstein, A, Marletta, M, Wand, A.J, Alber, T. | | Deposit date: | 2009-03-19 | | Release date: | 2010-03-31 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structural diversity in calmodulin recognition of nitric oxide synthases
To be Published
|
|
1I1J
 
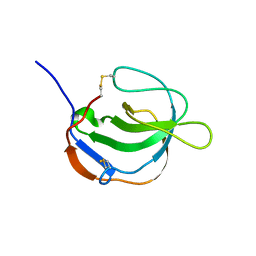 | | STRUCTURE OF MELANOMA INHIBITORY ACTIVITY PROTEIN: A MEMBER OF A NEW FAMILY OF SECRETED PROTEINS | | Descriptor: | MELANOMA DERIVED GROWTH REGULATORY PROTEIN | | Authors: | Lougheed, J.C, Holton, J.M, Alber, T, Bazan, J.F, Handel, T.M. | | Deposit date: | 2001-02-02 | | Release date: | 2001-05-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | Structure of melanoma inhibitory activity protein, a member of a recently identified family of secreted proteins.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
3IP2
 
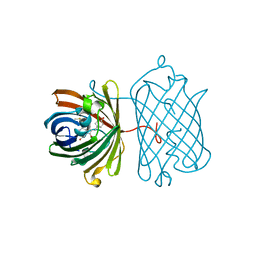 | | Crystal structure of red fluorescent protein Neptune at pH 7.0 | | Descriptor: | Neptune red fluorescent protein | | Authors: | Lin, M.Z, McKeown, M.R, Ng, H.L, Aguilera, T.A, Shaner, N.C, Ma, W, Adams, S.R, Campbell, R.E, Alber, T, Tsien, R.Y. | | Deposit date: | 2009-08-15 | | Release date: | 2009-12-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Autofluorescent proteins with excitation in the optical window for intravital imaging in mammals.
Chem.Biol., 16, 2009
|
|
3K0P
 
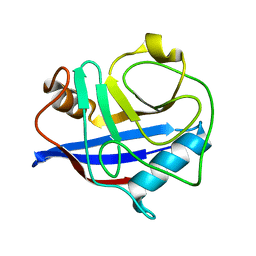 | |
1YWF
 
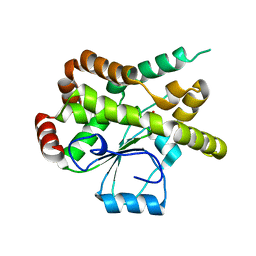 | |
3KA4
 
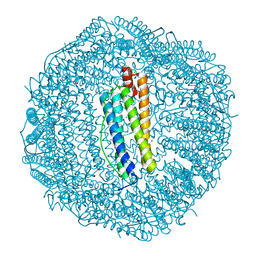 | | Frog M-ferritin with cobalt | | Descriptor: | CHLORIDE ION, COBALT (II) ION, Ferritin, ... | | Authors: | Tosha, T, Ng, H.L, Theil, E, Alber, T, Bhattasali, O. | | Deposit date: | 2009-10-18 | | Release date: | 2010-10-06 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Moving Metal Ions through Ferritin-Protein Nanocages from Three-Fold Pores to Catalytic Sites.
J.Am.Chem.Soc., 132, 2010
|
|
3KA6
 
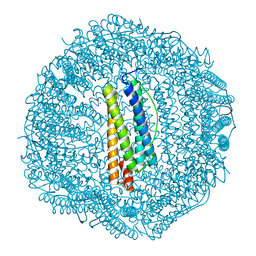 | | Frog M-ferritin, EED mutant, with cobalt | | Descriptor: | CHLORIDE ION, COBALT (II) ION, Ferritin, ... | | Authors: | Tosha, T, Ng, H.L, Theil, E, Alber, T, Bhattasali, O. | | Deposit date: | 2009-10-18 | | Release date: | 2010-10-06 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Moving Metal Ions through Ferritin-Protein Nanocages from Three-Fold Pores to Catalytic Sites.
J.Am.Chem.Soc., 132, 2010
|
|
3KA3
 
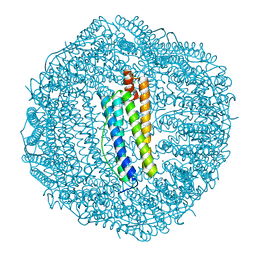 | | Frog M-ferritin with magnesium | | Descriptor: | CHLORIDE ION, Ferritin, middle subunit, ... | | Authors: | Tosha, T, Ng, H.L, Theil, E, Alber, T, Bhattasali, O. | | Deposit date: | 2009-10-18 | | Release date: | 2010-10-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Moving Metal Ions through Ferritin-Protein Nanocages from Three-Fold Pores to Catalytic Sites.
J.Am.Chem.Soc., 132, 2010
|
|
3KA8
 
 | | Frog M-ferritin, EQH mutant, with cobalt | | Descriptor: | CHLORIDE ION, COBALT (II) ION, Ferritin, ... | | Authors: | Tosha, T, Ng, H.L, Theil, E, Alber, T, Bhattasali, O. | | Deposit date: | 2009-10-19 | | Release date: | 2010-10-06 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Moving Metal Ions through Ferritin-Protein Nanocages from Three-Fold Pores to Catalytic Sites.
J.Am.Chem.Soc., 132, 2010
|
|
3KA9
 
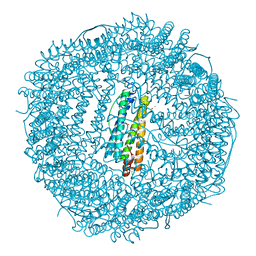 | | Frog M-ferritin, EEH mutant, with cobalt | | Descriptor: | CHLORIDE ION, COBALT (II) ION, Ferritin, ... | | Authors: | Tosha, T, Ng, H.L, Theil, E, Alber, T, Bhattasali, O. | | Deposit date: | 2009-10-19 | | Release date: | 2010-10-06 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Frog M-ferritin, EEH mutant, with cobalt
To be Published
|
|
3M4U
 
 | |
2O7A
 
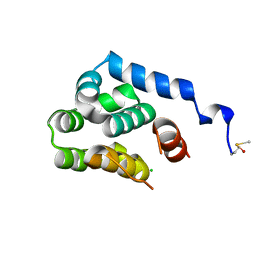 | | T4 lysozyme C-terminal fragment | | Descriptor: | ACETATE ION, CHLORIDE ION, Lysozyme | | Authors: | Echols, N, Kwon, E, Marqusee, S.M, Alber, T. | | Deposit date: | 2006-12-10 | | Release date: | 2007-04-10 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (0.84 Å) | | Cite: | Exploring subdomain cooperativity in T4 lysozyme I: Structural and energetic studies of a circular permutant and protein fragment.
Protein Sci., 16, 2007
|
|
2O60
 
 | |
