5JNA
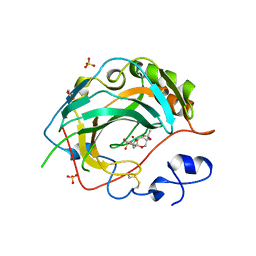 
 | | Crystal structure for the complex of human carbonic anhydrase IV and topiramate | | Descriptor: | ACETATE ION, Carbonic anhydrase 4, GLYCEROL, ... | | Authors: | Chen, Z, Waheed, A, Di Cera, E, Sly, W.S. | | Deposit date: | 2016-04-29 | | Release date: | 2017-05-03 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Intrinsic thermodynamics of high affinity inhibitor binding to recombinant human carbonic anhydrase IV.
Eur. Biophys. J., 47, 2018
|
|
2PP0
 
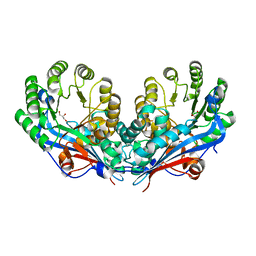 | | Crystal structure of L-talarate/galactarate dehydratase from Salmonella typhimurium LT2 | | Descriptor: | GLYCEROL, L-talarate/Galactarate Dehydratase | | Authors: | Fedorov, A.A, Fedorov, E.V, Yew, W.S, Gerlt, J.A, Almo, S.C. | | Deposit date: | 2007-04-27 | | Release date: | 2007-08-07 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Evolution of enzymatic activities in the enolase superfamily: L-talarate/galactarate dehydratase from Salmonella typhimurium LT2.
Biochemistry, 46, 2007
|
|
2PP1
 
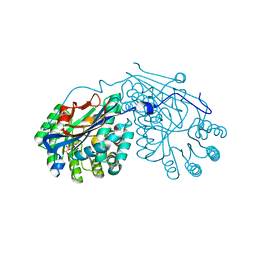 | | Crystal structure of L-talarate/galactarate dehydratase from Salmonella typhimurium LT2 liganded with Mg and L-lyxarohydroxamate | | Descriptor: | (2R,3S,4R)-2,3,4-TRIHYDROXY-5-(HYDROXYAMINO)-5-OXOPENTANOIC ACID, L-talarate/galactarate dehydratase, MAGNESIUM ION | | Authors: | Fedorov, A.A, Fedorov, E.V, Yew, W.S, Gerlt, J.A, Almo, S.C. | | Deposit date: | 2007-04-27 | | Release date: | 2007-08-07 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Evolution of enzymatic activities in the enolase superfamily: L-talarate/galactarate dehydratase from Salmonella typhimurium LT2.
Biochemistry, 46, 2007
|
|
2QIJ
 
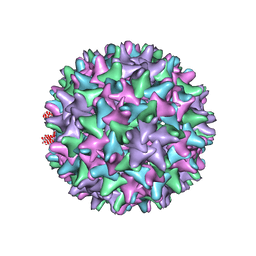 | | Hepatitis B Capsid Protein with an N-terminal extension modelled into 8.9 A data. | | Descriptor: | Core antigen | | Authors: | Tan, W.S, McNae, I.W, Ho, K.L, Walkinshaw, M.D. | | Deposit date: | 2007-07-04 | | Release date: | 2007-12-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (8.9 Å) | | Cite: | Crystallization and X-ray analysis of the T = 4 particle of hepatitis B capsid protein with an N-terminal extension.
Acta Crystallogr.,Sect.F, 63, 2007
|
|
4XPK
 
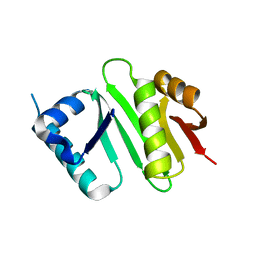 | | The crystal structure of Campylobacter jejuni N-acetyltransferase PseH | | Descriptor: | N-Acetyltransferase, PseH | | Authors: | Song, W.S, Nam, M.S, Namgung, B, Yoon, S.I. | | Deposit date: | 2015-01-17 | | Release date: | 2015-03-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural analysis of PseH, the Campylobacter jejuni N-acetyltransferase involved in bacterial O-linked glycosylation.
Biochem.Biophys.Res.Commun., 458, 2015
|
|
4XPL
 
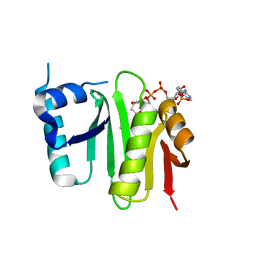 | | The crystal structure of Campylobacter jejuni N-acetyltransferase PseH in complex with acetyl coenzyme A | | Descriptor: | ACETYL COENZYME *A, N-Acetyltransferase, PseH | | Authors: | Song, W.S, Nam, M.S, Namgung, B, Yoon, S.I. | | Deposit date: | 2015-01-17 | | Release date: | 2015-03-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural analysis of PseH, the Campylobacter jejuni N-acetyltransferase involved in bacterial O-linked glycosylation.
Biochem.Biophys.Res.Commun., 458, 2015
|
|
4WH4
 
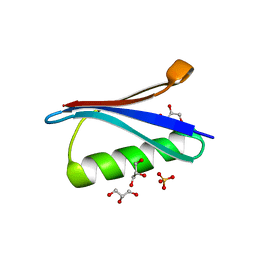 | | Protein GB1 Quadruple Mutant I6H/N8H/K28H/Q32H | | Descriptor: | GLYCEROL, Immunoglobulin G-binding protein G, SULFATE ION | | Authors: | Cunningham, T.C, Horne, W.S, Saxena, S. | | Deposit date: | 2014-09-19 | | Release date: | 2015-08-05 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The double-histidine Cu2+-binding motif: a highly rigid, site-specific spin probe for electron spin resonance distance measurements.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
3K1E
 
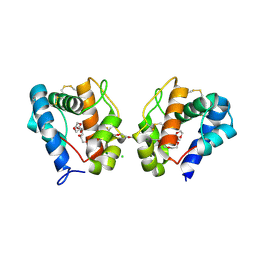 | | Crystal structure of odorant binding protein 1 (AaegOBP1) from Aedes aegypti | | Descriptor: | 2,5,8,11,14,17,20,23,26,29,32,35,38,41,44,47,50,53,56,59,62,65,68,71,74,77,80-HEPTACOSAOXADOOCTACONTAN-82-OL, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Leite, N.R, Krogh, R, Leal, W.S, Iulek, J, Oliva, G. | | Deposit date: | 2009-09-27 | | Release date: | 2009-12-08 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structure of an odorant-binding protein from the mosquito Aedes aegypti suggests a binding pocket covered by a pH-sensitive "Lid".
Plos One, 4, 2009
|
|
3IYS
 
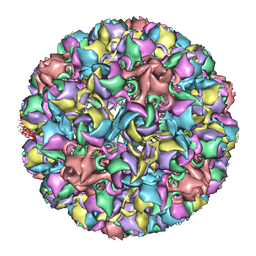 | | Homology model of avian polyomavirus asymmetric unit | | Descriptor: | Major capsid protein VP1 | | Authors: | Shen, P.S, Enderlein, D, Nelson, C.D.S, Carter, W.S, Kawano, M, Xing, L, Swenson, R.D, Olson, N.H, Baker, T.S, Cheng, R.H, Atwood, W.J, Johne, R, Belnap, D.M. | | Deposit date: | 2010-04-19 | | Release date: | 2011-01-26 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (11.3 Å) | | Cite: | The structure of avian polyomavirus reveals variably sized capsids, non-conserved inter-capsomere interactions, and a possible location of the minor capsid protein VP4.
Virology, 411, 2011
|
|
3HEX
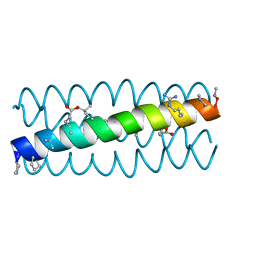 
 | | Cyclic residues in alpha/beta-peptide helix bundles: GCN4-pLI side chain sequence on an (alpha-alpha-beta) backbone with cyclic beta-residues at positions 1, 4, 19 and 28 | | Descriptor: | alpha/beta-peptide based on the GCN4-pLI side chain sequence with an (alpha-alpha-beta) backbone and cyclic beta-residues at positions 1, 4, 19 and 28 | | Authors: | Horne, W.S, Price, J.L, Gellman, S.H. | | Deposit date: | 2009-05-10 | | Release date: | 2010-04-21 | | Last modified: | 2024-07-10 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural consequences of beta-amino acid preorganization in a self-assembling alpha/beta-peptide: fundamental studies of foldameric helix bundles.
J.Am.Chem.Soc., 132, 2010
|
|
4YJY
 
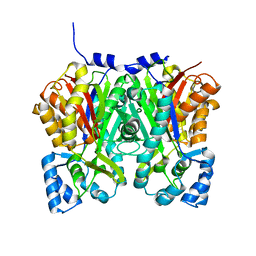 | |
2INV
 
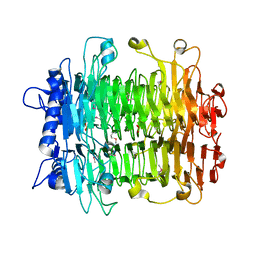 | |
2ID5
 
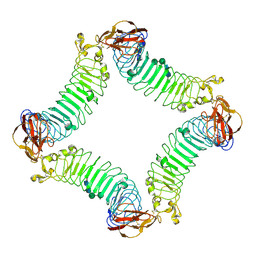 | | Crystal Structure of the Lingo-1 Ectodomain | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Leucine rich repeat neuronal 6A, ... | | Authors: | Mosyak, L, Wood, A, Dwyer, B, Johnson, M, Stahl, M.L, Somers, W.S. | | Deposit date: | 2006-09-14 | | Release date: | 2006-09-26 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.698 Å) | | Cite: | The structure of the Lingo-1 ectodomain, a module implicated in central nervous system repair inhibition.
J.Biol.Chem., 281, 2006
|
|
2HXU
 
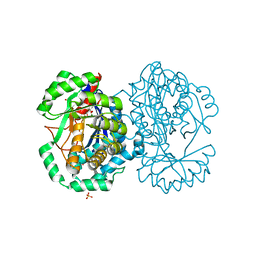 | | Crystal structure of K220A mutant of L-Fuconate Dehydratase from Xanthomonas campestris liganded with Mg++ and L-fuconate | | Descriptor: | 6-deoxy-L-galactonic acid, L-fuconate dehydratase, MAGNESIUM ION, ... | | Authors: | Fedorov, A.A, Fedorov, E.V, Yew, W.S, Rakus, J.F, Gerlt, J.A, Almo, S.C. | | Deposit date: | 2006-08-03 | | Release date: | 2006-12-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Evolution of Enzymatic Activities in the Enolase Superfamily: l-Fuconate Dehydratase from Xanthomonas campestris.
Biochemistry, 45, 2006
|
|
2INU
 
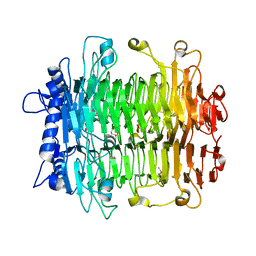 | |
2HNE
 
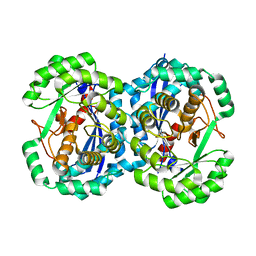 | | Crystal structure of l-fuconate dehydratase from xanthomonas campestris pv. campestris str. ATCC 33913 | | Descriptor: | L-fuconate dehydratase, MAGNESIUM ION | | Authors: | Fedorov, A.A, Fedorov, E.V, Yew, W.S, Gerlt, J.A, Almo, S.C, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2006-07-12 | | Release date: | 2006-07-25 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of l-fuconate dehydratase from xanthomonas campestris pv. campestris str. ATCC 33913
To be Published
|
|
2HXT
 
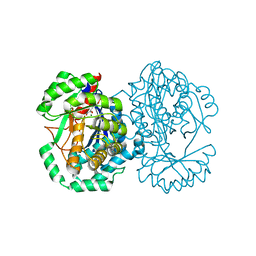 | | Crystal structure of L-Fuconate Dehydratase from Xanthomonas campestris liganded with Mg++ and D-erythronohydroxamate | | Descriptor: | (2R,3R)-N,2,3,4-TETRAHYDROXYBUTANAMIDE, L-fuconate dehydratase, MAGNESIUM ION | | Authors: | Fedorov, A.A, Fedorov, E.V, Yew, W.S, Rakus, J.F, Gerlt, J.A, Almo, S.C. | | Deposit date: | 2006-08-03 | | Release date: | 2006-12-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Evolution of Enzymatic Activities in the Enolase Superfamily: l-Fuconate Dehydratase from Xanthomonas campestris.
Biochemistry, 45, 2006
|
|
2JPO
 
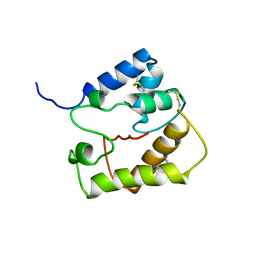 | |
1HLA
 
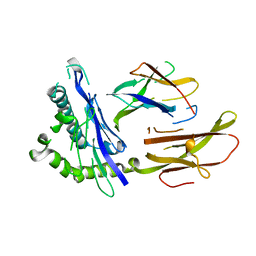 | | STRUCTURE OF THE HUMAN CLASS I HISTOCOMPATIBILITY ANTIGEN, HLA-A2 | | Descriptor: | BETA 2-MICROGLOBULIN, CLASS I HISTOCOMPATIBILITY ANTIGEN (HLA-A2) (ALPHA CHAIN) | | Authors: | Bjorkman, P.J, Saper, M.A, Samraoui, B, Bennett, W.S, Strominger, J.L, Wiley, D.C. | | Deposit date: | 1987-10-15 | | Release date: | 1988-01-16 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structure of the human class I histocompatibility antigen, HLA-A2.
Nature, 329, 1987
|
|
5U5B
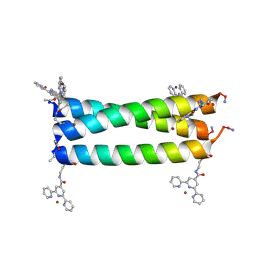 
 | | Coiled Coil Peptide Metal Coordination Framework: Trimer Fold | | Descriptor: | COPPER (II) ION, Designed trimeric coiled coil peptide with two terpyridine side chains, GLYCEROL | | Authors: | Tavenor, N.A, Murnin, M.J, Horne, W.S. | | Deposit date: | 2016-12-06 | | Release date: | 2017-02-22 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Supramolecular Metal-Coordination Polymers, Nets, and Frameworks from Synthetic Coiled-Coil Peptides.
J. Am. Chem. Soc., 139, 2017
|
|
5U59
 
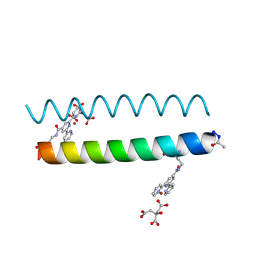 | | Coiled Coil Peptide Metal Coordination Framework: Dimer Fold Grown with Citrate | | Descriptor: | CITRIC ACID, COPPER (II) ION, Designed dimeric coiled coil peptide with two terpyridine side chains | | Authors: | Tavenor, N.A, Murnin, M.J, Horne, W.S. | | Deposit date: | 2016-12-06 | | Release date: | 2017-02-22 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Supramolecular Metal-Coordination Polymers, Nets, and Frameworks from Synthetic Coiled-Coil Peptides.
J. Am. Chem. Soc., 139, 2017
|
|
5U5C
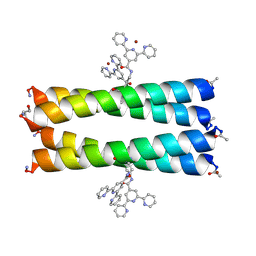 
 | | Coiled Coil Peptide Metal Coordination Framework: Tetramer Fold | | Descriptor: | COPPER (II) ION, Designed tetrameric coiled coil peptide with one terpyridine side chain, HEXANE-1,6-DIOL | | Authors: | Tavenor, N.A, Murnin, M.J, Horne, W.S. | | Deposit date: | 2016-12-06 | | Release date: | 2017-02-22 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Supramolecular Metal-Coordination Polymers, Nets, and Frameworks from Synthetic Coiled-Coil Peptides.
J. Am. Chem. Soc., 139, 2017
|
|
5U5A
 
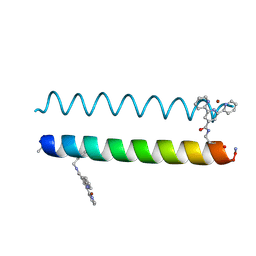 | | Coiled Coil Peptide Metal Coordination Framework: Dimer Fold | | Descriptor: | COPPER (II) ION, Designed dimeric coiled coil peptide with two terpyridine side chains | | Authors: | Tavenor, N.A, Murnin, M.J, Horne, W.S. | | Deposit date: | 2016-12-06 | | Release date: | 2017-02-22 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.23 Å) | | Cite: | Supramolecular Metal-Coordination Polymers, Nets, and Frameworks from Synthetic Coiled-Coil Peptides.
J. Am. Chem. Soc., 139, 2017
|
|
6PV1
 
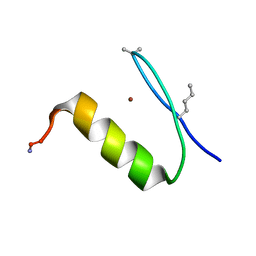 | |
5YCL
 
 | |
