5ZB3
 
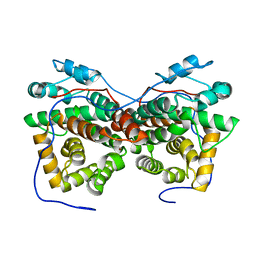 | | Dimeric crystal structure of ORF57 from KSHV | | Descriptor: | ORF57, ZINC ION | | Authors: | Gao, Z.Q, Yuan, F, Dong, Y.H, Lan, K. | | Deposit date: | 2018-02-09 | | Release date: | 2018-08-08 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (3.506 Å) | | Cite: | The crystal structure of KSHV ORF57 reveals dimeric active sites important for protein stability and function.
PLoS Pathog., 14, 2018
|
|
3BW3
 
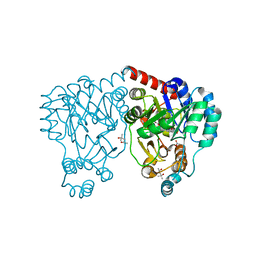 | | Crystal structures and site-directed mutagenesis study of nitroalkane oxidase from Streptomyces ansochromogenes | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-nitropropane dioxygenase, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Li, Y.H, Gao, Z.Q, Hou, H.F. | | Deposit date: | 2008-01-08 | | Release date: | 2009-01-13 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure and site-directed mutagenesis of a nitroalkane oxidase from Streptomyces ansochromogenes
Biochem.Biophys.Res.Commun., 405, 2011
|
|
3BW2
 
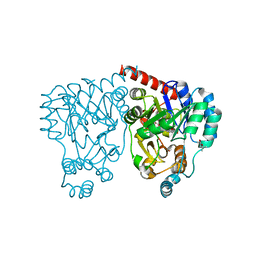 | | Crystal structures and site-directed mutagenesis study of nitroalkane oxidase from Streptomyces ansochromogenes | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-nitropropane dioxygenase, FLAVIN MONONUCLEOTIDE | | Authors: | Li, Y.H, Gao, Z.Q, Hou, H.F. | | Deposit date: | 2008-01-08 | | Release date: | 2009-01-13 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure and site-directed mutagenesis of a nitroalkane oxidase from Streptomyces ansochromogenes
Biochem.Biophys.Res.Commun., 405, 2011
|
|
3BW4
 
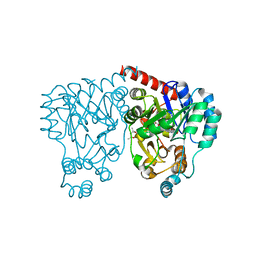 | |
3LCM
 
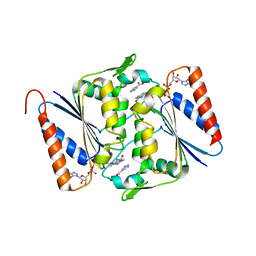 | | Crystal structure of Smu.1420 from Streptococcus mutans UA159 | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Putative oxidoreductase | | Authors: | Wang, Z.X, Su, X.-D. | | Deposit date: | 2010-01-11 | | Release date: | 2011-01-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.799 Å) | | Cite: | Structural and biochemical characterization of MdaB from cariogenic Streptococcus mutans reveals an NADPH-specific quinone oxidoreductase
Acta Crystallogr.,Sect.D, 70, 2014
|
|
5C2O
 
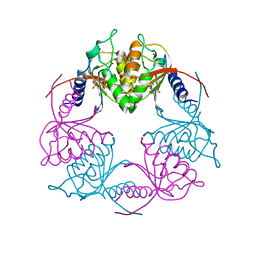 | | Crystal structure of Streptococcus mutans Deoxycytidylate Deaminase complexed with dTTP | | Descriptor: | MAGNESIUM ION, Putative deoxycytidylate deaminase, THYMIDINE-5'-TRIPHOSPHATE, ... | | Authors: | Li, Y.H, Gao, Z.Q, Hou, H.F, Dong, Y.H. | | Deposit date: | 2015-06-16 | | Release date: | 2016-07-13 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Mechanism of the allosteric regulation of Streptococcus mutans 2'-deoxycytidylate deaminase.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
2BB0
 
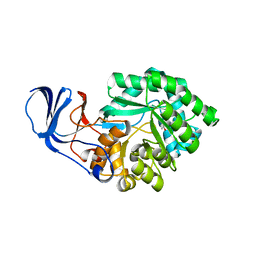 | |
2G3F
 
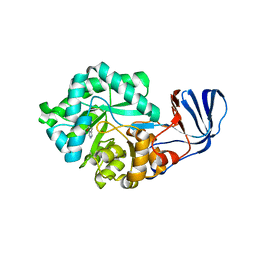 | | Crystal Structure of imidazolonepropionase complexed with imidazole-4-acetic acid sodium salt, a substrate homologue | | Descriptor: | 2H-IMIDAZOL-4-YLACETIC ACID, Imidazolonepropionase, ZINC ION | | Authors: | Yu, Y, Liang, Y.H, Su, X.D. | | Deposit date: | 2006-02-19 | | Release date: | 2006-09-26 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A catalytic mechanism revealed by the crystal structures of the imidazolonepropionase from Bacillus subtilis
J.Biol.Chem., 281, 2006
|
|
3Q5M
 
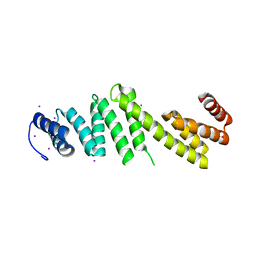 | | Crystal structure of Escherichia coli BamD | | Descriptor: | IODIDE ION, UPF0169 lipoprotein yfiO | | Authors: | Dong, C, Hou, H, Yang, X, Dong, Y, Shen, Y. | | Deposit date: | 2010-12-28 | | Release date: | 2011-12-28 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.604 Å) | | Cite: | Structure of Escherichia coli BamD and its functional implications in outer membrane protein assembly
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4NSR
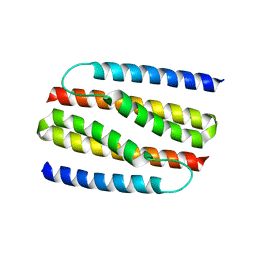 
 | |
3Q54
 
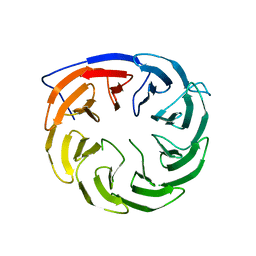 | | Crystal structure of Escherichia coli BamB | | Descriptor: | Outer membrane assembly lipoprotein YfgL | | Authors: | Dong, C, Hou, H, Yang, X. | | Deposit date: | 2010-12-27 | | Release date: | 2012-05-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.004 Å) | | Cite: | Structure of Escherichia coli BamB and its interaction with POTRA domains of BamA.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4XO2
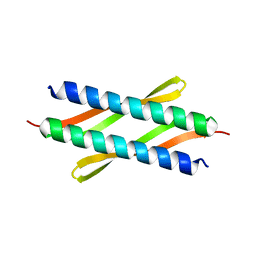 
 | |
4NSO
 
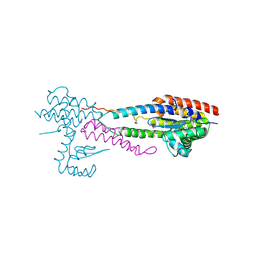 | | Crystal structure of the effector-immunity protein complex | | Descriptor: | Effector protein, Immunity protein | | Authors: | Dong, C. | | Deposit date: | 2013-11-28 | | Release date: | 2014-04-16 | | Last modified: | 2014-06-11 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for recognition of the type VI spike protein VgrG3 by a cognate immunity protein.
Febs Lett., 588, 2014
|
|
6JDP
 
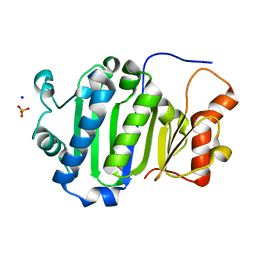 | |
8GOT
 
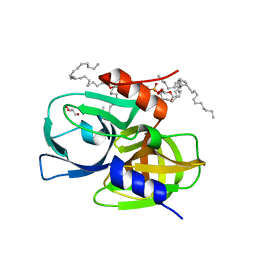 | | Crystal structure of wild-type protease 3C from Seneca Valley Virus | | Descriptor: | GLYCEROL, Peptidase C3, [(2~{S})-2-hexadecanoyloxy-3-[[(2~{R})-3-[[(2~{S})-3-[(5~{E},8~{E},11~{Z},14~{E})-icosa-5,8,11,14-tetraenoyl]oxy-2-[(9~{E},12~{Z})-octadeca-9,12-dienoyl]oxy-propoxy]-oxidanyl-phosphoryl]oxy-2-oxidanyl-propoxy]-oxidanyl-phosphoryl]oxy-propyl] icosanoate | | Authors: | Zhao, H.F, Zhang, H. | | Deposit date: | 2022-08-25 | | Release date: | 2023-05-24 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.989 Å) | | Cite: | Allosteric regulation of Senecavirus A 3Cpro proteolytic activity by an endogenous phospholipid.
Plos Pathog., 19, 2023
|
|
8GPH
 
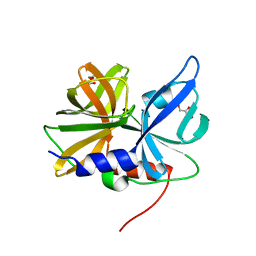 | |
5IFG
 
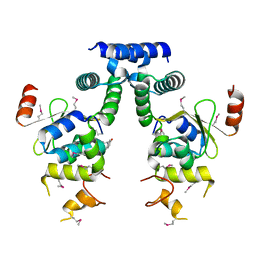 | | Crystal structure of HigA-HigB complex from E. Coli | | Descriptor: | Antitoxin HigA, mRNA interferase HigB | | Authors: | Yang, J.S, Zhou, K, Gao, z.Q, Liu, Q.S, Dong, Y.H. | | Deposit date: | 2016-02-26 | | Release date: | 2017-03-01 | | Method: | X-RAY DIFFRACTION (2.702 Å) | | Cite: | Structural insight into the E. coli HigBA complex
Biochem. Biophys. Res. Commun., 478, 2016
|
|
4X36
 
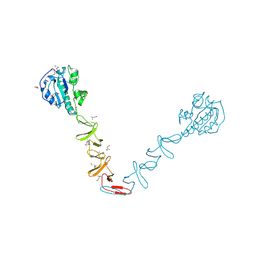 | | Crystal structure of the autolysin LytA from Streptococcus pneumoniae TIGR4 | | Descriptor: | Autolysin, CHOLINE ION, GLYCEROL, ... | | Authors: | Cheng, W, Li, Q, Zhou, C.Z, Chen, Y.X. | | Deposit date: | 2014-11-28 | | Release date: | 2015-05-27 | | Last modified: | 2015-06-24 | | Method: | X-RAY DIFFRACTION (2.101 Å) | | Cite: | Full-length structure of the major autolysin LytA.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
7D8U
 
 | | Crystal structure of the C-terminal domain of pNP868R from African swine fever virus | | Descriptor: | 1,2-ETHANEDIOL, FORMIC ACID, GTP--RNA guanylyltransferase, ... | | Authors: | Du, X, Geng, Z, Zhang, H. | | Deposit date: | 2020-10-10 | | Release date: | 2020-12-02 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure and Biochemical Characteristic of the Methyltransferase (MTase) Domain of RNA Capping Enzyme from African Swine Fever Virus.
J.Virol., 2020
|
|
4JKZ
 
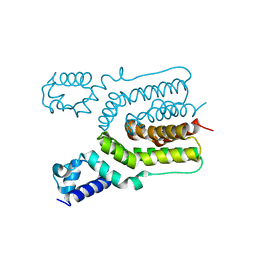 | | Crystal structure of ms6564 from mycobacterium smegmatis | | Descriptor: | Transcriptional regulator, TetR family | | Authors: | Yang, S.F, Gao, Z.Q, He, Z.G, Dong, Y.H. | | Deposit date: | 2013-03-12 | | Release date: | 2013-06-26 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for interaction between Mycobacterium smegmatis Ms6564, a TetR family master regulator, and its target DNA.
J.Biol.Chem., 288, 2013
|
|
4JL3
 
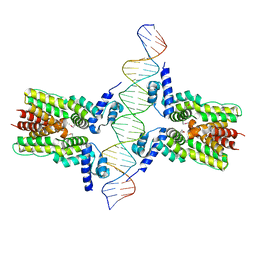 | | Crystal structure of ms6564-dna complex | | Descriptor: | DNA (31-MER), Transcriptional regulator, TetR family | | Authors: | Yang, S.F, Gao, Z.Q, He, Z.G, Dong, Y.H. | | Deposit date: | 2013-03-12 | | Release date: | 2013-06-26 | | Last modified: | 2022-08-24 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis for interaction between Mycobacterium smegmatis Ms6564, a TetR family master regulator, and its target DNA.
J.Biol.Chem., 288, 2013
|
|
3OIX
 
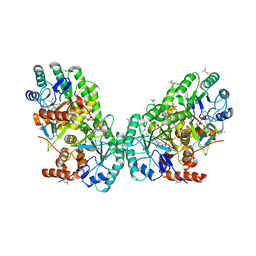 | | Crystal structure of the putative dihydroorotate dehydrogenase from Streptococcus mutans | | Descriptor: | FLAVIN MONONUCLEOTIDE, GLYCEROL, Putative dihydroorotate dehydrogenase; dihydroorotate oxidase | | Authors: | Liu, Y, Gao, Z.Q, Liu, C.P, Dong, Y.H. | | Deposit date: | 2010-08-20 | | Release date: | 2010-09-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.399 Å) | | Cite: | Structure of the putative dihydroorotate dehydrogenase from Streptococcus mutans
Acta Crystallogr.,Sect.F, 67, 2011
|
|
7AER
 
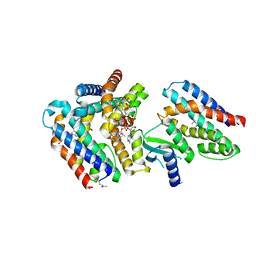 | | Rebuilt and re-refined PDB entry 5yep: tri-AMPylated Shewanella oneidensis HEPN toxin in complex with MNT antitoxin | | Descriptor: | ADENOSINE MONOPHOSPHATE, Toxin-antitoxin system antidote Mnt family, Toxin-antitoxin system toxin HepN family | | Authors: | Tamulaitiene, G, Sasnauskas, G, Songailiene, I, Juozapaitis, J, Siksnys, V. | | Deposit date: | 2020-09-18 | | Release date: | 2020-12-30 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | HEPN-MNT Toxin-Antitoxin System: The HEPN Ribonuclease Is Neutralized by OligoAMPylation.
Mol.Cell, 80, 2020
|
|
6A2H
 
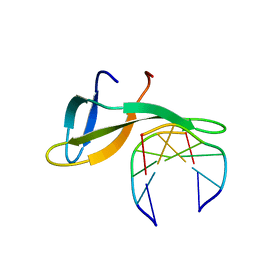 | | Architectural roles of Cren7 in folding crenarchaeal chromatin filament | | Descriptor: | Chromatin protein Cren7, DNA (5'-D(P*AP*AP*TP*TP*AP*C)-3'), DNA (5'-D(P*GP*TP*AP*AP*TP*T)-3') | | Authors: | Zhang, Z.F, Zhao, M.H, Chen, Y.Y, Wang, L, Dong, Y.H, Gong, Y, Huang, L. | | Deposit date: | 2018-06-11 | | Release date: | 2019-01-16 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Architectural roles of Cren7 in folding crenarchaeal chromatin filament.
Mol. Microbiol., 111, 2019
|
|
6A2I
 
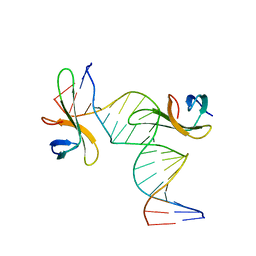 | | Architectural roles of Cren7 in folding crenarchaeal chromatin filament | | Descriptor: | Chromatin protein Cren7, DNA (5'-D(*CP*GP*TP*AP*GP*CP*TP*AP*AP*TP*TP*AP*GP*CP*TP*AP*CP*G)-3') | | Authors: | Zhang, Z.F, Zhao, M.H, Chen, Y.Y, Wang, L, Dong, Y.H, Gong, Y, Huang, L. | | Deposit date: | 2018-06-11 | | Release date: | 2019-01-16 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Architectural roles of Cren7 in folding crenarchaeal chromatin filament.
Mol. Microbiol., 111, 2019
|
|
