6GGZ
 
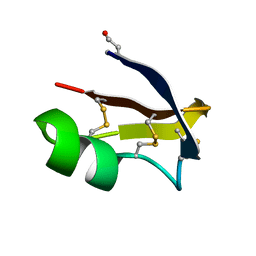 | | NMR structure of the scorpion toxin AmmTx3 | | Descriptor: | Potassium channel toxin alpha-KTx 15.3 | | Authors: | Landon, C, Meudal, H. | | Deposit date: | 2018-05-04 | | Release date: | 2019-01-30 | | Last modified: | 2020-03-11 | | Method: | SOLUTION NMR | | Cite: | Synthesis by native chemical ligation and characterization of the scorpion toxin AmmTx3.
Bioorg. Med. Chem., 27, 2019
|
|
4D0U
 
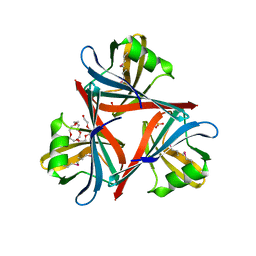 | | Crystal structure of the fiber head domain of the Atadenovirus snake adenovirus 1, selenomethionine-derivative | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, FIBER PROTEIN, SULFATE ION | | Authors: | Singh, A.K, van Raaij, M.J. | | Deposit date: | 2014-04-30 | | Release date: | 2014-12-17 | | Last modified: | 2018-12-19 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of the fibre head domain of the Atadenovirus Snake Adenovirus 1.
PLoS ONE, 9, 2014
|
|
7UCP
 
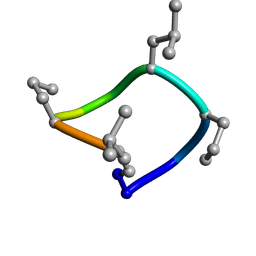 | | computationally designed macrocycle | | Descriptor: | computationally designed cyclic peptide D8.3.p2 | | Authors: | Bhardwaj, G, Baker, D, Rettie, S, Glynn, C, Sawaya, M. | | Deposit date: | 2022-03-17 | | Release date: | 2022-09-14 | | Last modified: | 2022-09-28 | | Method: | X-RAY DIFFRACTION (0.85 Å) | | Cite: | Accurate de novo design of membrane-traversing macrocycles.
Cell, 185, 2022
|
|
4PXJ
 
 | |
3O4N
 
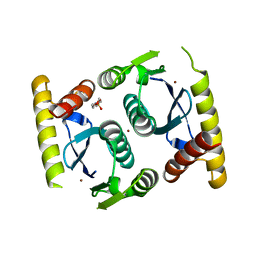 | |
3O4Q
 
 | |
4D0V
 
 | | Crystal structure of the fiber head domain of the Atadenovirus snake adenovirus 1, native, I213 crystal form | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, FIBER PROTEIN, SULFATE ION | | Authors: | Singh, A.K, van Raaij, M.J. | | Deposit date: | 2014-04-30 | | Release date: | 2014-12-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of the fibre head domain of the Atadenovirus Snake Adenovirus 1.
PLoS ONE, 9, 2014
|
|
4D1G
 
 | | Crystal structure of the fiber head domain of the Atadenovirus snake adenovirus 1, native, second P212121 crystal form | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, FIBER PROTEIN, SULFATE ION | | Authors: | Singh, A.K, van Raaij, M.J. | | Deposit date: | 2014-05-01 | | Release date: | 2014-12-17 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the fibre head domain of the Atadenovirus Snake Adenovirus 1.
PLoS ONE, 9, 2014
|
|
4D1F
 
 | |
7ZT2
 
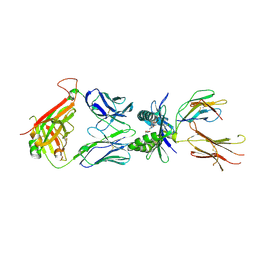 | | Structure of E8 TCR in complex with human MR1 bound to 5-OP-RU | | Descriptor: | 1,2-ETHANEDIOL, 1-deoxy-1-({2,6-dioxo-5-[(E)-propylideneamino]-1,2,3,6-tetrahydropyrimidin-4-yl}amino)-D-ribitol, Beta-2-microglobulin, ... | | Authors: | Karuppiah, V, Srikannathasan, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT4
 
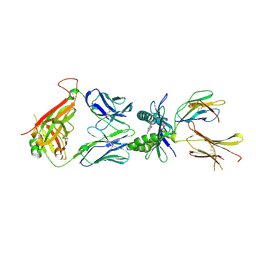 | | Structure of E8 TCR in complex with human MR1 bound to 6FP | | Descriptor: | 2-azanyl-6-methyl-3~{H}-pteridin-4-one, Beta-2-microglobulin, Major histocompatibility complex class I-related gene protein, ... | | Authors: | Karuppiah, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT9
 
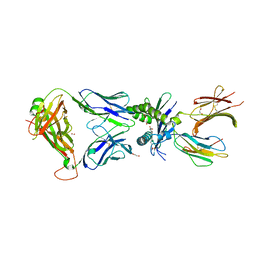 | | Structure of E8 TCR in complex in human MR1 bound to 4FBA | | Descriptor: | 1,2-ETHANEDIOL, 4-METHYLBENZOIC ACID, Beta-2-microglobulin, ... | | Authors: | Karuppiah, V, Srikannathasan, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT8
 
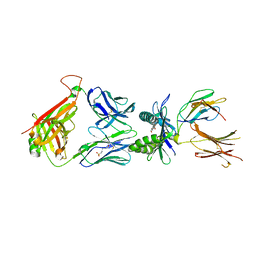 | | Structure of E8 TCR in complex in human MR1 bound to 3FBA | | Descriptor: | 1,2-ETHANEDIOL, 3-methylbenzoic acid, 3[N-MORPHOLINO]PROPANE SULFONIC ACID, ... | | Authors: | Karuppiah, V, Srikannathasan, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT3
 
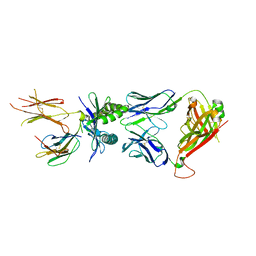 | | Structure of E8 TCR in complex in human MR1 K43A | | Descriptor: | Beta-2-microglobulin, Major histocompatibility complex class I-related gene protein, TCR alpha, ... | | Authors: | Karuppiah, V, Srikannathasan, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT5
 
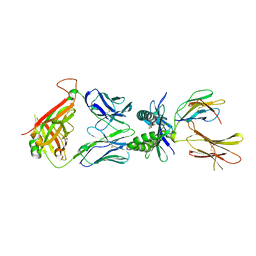 | | Structure of E8 TCR in complex in human MR1 bound to 3FSA | | Descriptor: | 3-methanoyl-2-oxidanyl-benzoic acid, Beta-2-microglobulin, Major histocompatibility complex class I-related gene protein, ... | | Authors: | Karuppiah, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT7
 
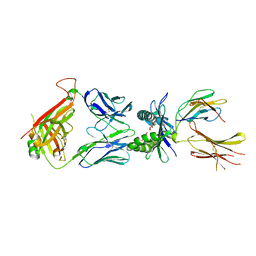 | |
8PEP
 
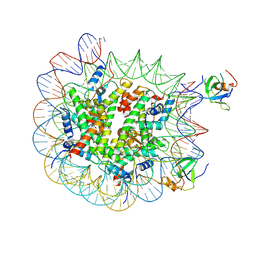 | | H3K36me2 nucleosome-LEDGF/p75 PWWP domain complex - pose 2 | | Descriptor: | Histone H2A, Histone H2B 1.1, Histone H3, ... | | Authors: | Koutna, E, Kouba, T, Veverka, V. | | Deposit date: | 2023-06-14 | | Release date: | 2023-08-16 | | Last modified: | 2023-11-15 | | Method: | ELECTRON MICROSCOPY (3.33 Å) | | Cite: | Multivalency of nucleosome recognition by LEDGF.
Nucleic Acids Res., 51, 2023
|
|
8PEO
 
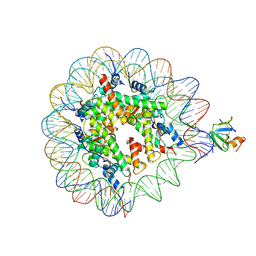 | | H3K36me2 nucleosome-LEDGF/p75 PWWP domain complex | | Descriptor: | (2R)-2-amino-3-(2-dimethylaminoethylsulfanyl)propanoic acid, Histone H2A, Histone H2B 1.1, ... | | Authors: | Koutna, E, Kouba, T, Veverka, V. | | Deposit date: | 2023-06-14 | | Release date: | 2023-08-16 | | Last modified: | 2023-11-15 | | Method: | ELECTRON MICROSCOPY (2.69 Å) | | Cite: | Multivalency of nucleosome recognition by LEDGF.
Nucleic Acids Res., 51, 2023
|
|
8PC5
 
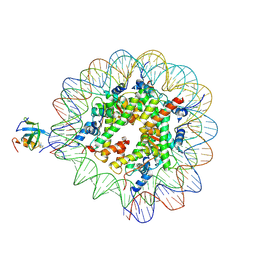 | | H3K36me3 nucleosome-LEDGF/p75 PWWP domain complex | | Descriptor: | Histone H2A, Histone H2B 1.1, Histone H3, ... | | Authors: | Koutna, E, Kouba, T, Veverka, V. | | Deposit date: | 2023-06-09 | | Release date: | 2023-08-16 | | Last modified: | 2023-11-15 | | Method: | ELECTRON MICROSCOPY (3.02 Å) | | Cite: | Multivalency of nucleosome recognition by LEDGF.
Nucleic Acids Res., 51, 2023
|
|
8PC6
 
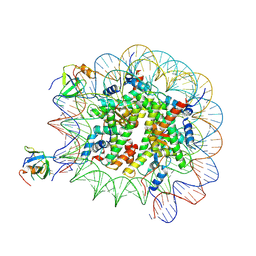 | | H3K36me3 nucleosome-LEDGF/p75 PWWP domain complex - pose 2 | | Descriptor: | Histone H2A, Histone H2B 1.1, Histone H3, ... | | Authors: | Koutna, E, Kouba, T, Veverka, V. | | Deposit date: | 2023-06-09 | | Release date: | 2023-08-16 | | Last modified: | 2023-11-15 | | Method: | ELECTRON MICROSCOPY (3.04 Å) | | Cite: | Multivalency of nucleosome recognition by LEDGF.
Nucleic Acids Res., 51, 2023
|
|
2JLB
 
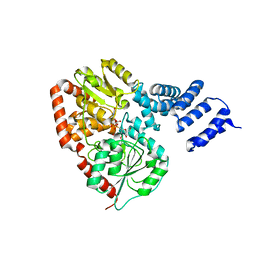 | | Xanthomonas campestris putative OGT (XCC0866), complex with UDP- GlcNAc phosphonate analogue | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, URIDINE-DIPHOSPHATE-METHYLENE-N-ACETYL-GLUCOSAMINE, ... | | Authors: | Schuettelkopf, A.W, Clarke, A.J, van Aalten, D.M.F. | | Deposit date: | 2008-09-07 | | Release date: | 2008-11-25 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural Insights Into Mechanism and Specificity of O-Glcnac Transferase.
Embo J., 27, 2008
|
|
1VI7
 
 | | Crystal structure of an hypothetical protein | | Descriptor: | Hypothetical protein yigZ | | Authors: | Structural GenomiX | | Deposit date: | 2003-12-01 | | Release date: | 2003-12-30 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of YIGZ, a conserved hypothetical protein from Escherichia coli k12 with a novel fold
Proteins, 55, 2004
|
|
8BI2
 
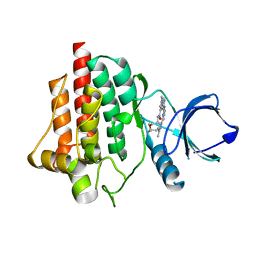 | | Syk kinase domain in complex with macrocyclic inhibitor 20a | | Descriptor: | 10,13,23-trimethyl-16-oxa-2,4,8,9,13,19,23,30-octazapentacyclo[19.5.2.1^{3,7}.1^{8,11}.0^{24,28}]triaconta-1(27),3,5,7(30),9,11(29),21,24(28),25-nonaen-20-one, Tyrosine-protein kinase SYK | | Authors: | Read, J.A, Patel, J. | | Deposit date: | 2022-11-01 | | Release date: | 2023-06-14 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.508 Å) | | Cite: | Optimization of a series of novel, potent and selective Macrocyclic SYK inhibitors.
Bioorg.Med.Chem.Lett., 91, 2023
|
|
8CIQ
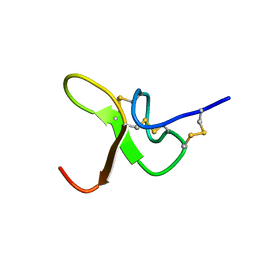 
 | | JzTx-34 toxin peptide | | Descriptor: | Mu-theraphotoxin-Cg1a | | Authors: | Landon, C, Meudal, H. | | Deposit date: | 2023-02-10 | | Release date: | 2023-07-26 | | Method: | SOLUTION NMR | | Cite: | Structure-function relationship of new peptides activating human Na v 1.1.
Biomed Pharmacother, 165, 2023
|
|
8CJP
 
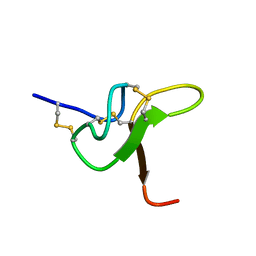 | | JzTx-34 toxin peptide H18A mutant | | Descriptor: | Mu-theraphotoxin-Cg1a | | Authors: | Landon, C, Meudal, H. | | Deposit date: | 2023-02-13 | | Release date: | 2023-07-26 | | Method: | SOLUTION NMR | | Cite: | Structure-function relationship of new peptides activating human Na v 1.1.
Biomed Pharmacother, 165, 2023
|
|
