8IHU
 
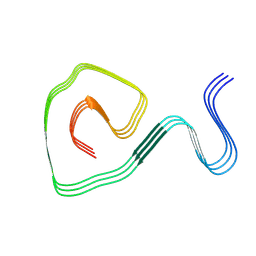 | | Cryo-EM structure of an amyloid fibril formed by ALS-causing SOD1 mutation G85R | | Descriptor: | Superoxide dismutase [Cu-Zn] | | Authors: | Wang, L.Q, Ma, Y.Y, Zhang, M.Y, Yuan, H.Y, Li, X.N, Zhao, K, Chen, J, Li, D, Wang, Z.Z, Le, W.D, Liu, C, Liang, Y. | | Deposit date: | 2023-02-23 | | Release date: | 2024-10-30 | | Method: | ELECTRON MICROSCOPY (2.97 Å) | | Cite: | Amyloid fibril structures and ferroptosis activation induced by ALS-causing SOD1 mutations.
Sci Adv, 2024
|
|
8IHV
 
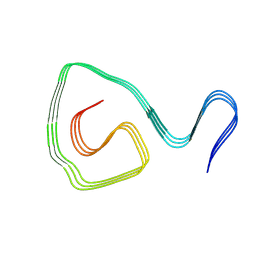 | | Cryo-EM structure of an amyloid fibril formed by ALS-causing SOD1 mutation H46R | | Descriptor: | Superoxide dismutase [Cu-Zn] | | Authors: | Wang, L.Q, Ma, Y.Y, Zhang, M.Y, Yuan, H.Y, Li, X.N, Zhao, K, Chen, J, Li, D, Wang, Z.Z, Le, W.D, Liu, C, Liang, Y. | | Deposit date: | 2023-02-23 | | Release date: | 2024-10-30 | | Method: | ELECTRON MICROSCOPY (3.11 Å) | | Cite: | Amyloid fibril structures and ferroptosis activation induced by ALS-causing SOD1 mutations.
Sci Adv, 2024
|
|
9J6I
 
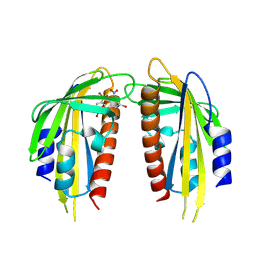 | | Crystal structure of the ABA receptor PYL1 in complex with DBSA compound | | Descriptor: | 5-bromanyl-N2-[(4-bromophenyl)-bis(oxidanyl)-$l^4-sulfanyl]-N1,N1,N3,N3-tetrakis(oxidanyl)benzene-1,2,3-triamine, Abscisic acid receptor PYL1 | | Authors: | Yan, J. | | Deposit date: | 2024-08-16 | | Release date: | 2024-09-04 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Stabilization of dimeric PYR/PYL/RCAR family members relieves abscisic acid-induced inhibition of seed germination.
Nat Commun, 15, 2024
|
|
3O6Y
 
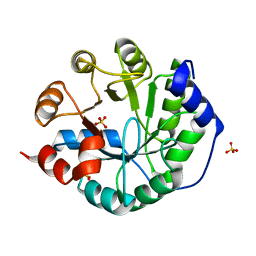 | | Robust computational design, optimization, and structural characterization of retroaldol enzymes | | Descriptor: | Retro-Aldolase, SULFATE ION | | Authors: | Althoff, E.A, Wang, L, Jiang, L, Moody, J, Bolduc, J, Lassila, J.K, Wang, Z.Z, Smith, M, Hari, S, Herschlag, D, Stoddard, B.L, Baker, D. | | Deposit date: | 2010-07-29 | | Release date: | 2011-06-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.091 Å) | | Cite: | Structural analyses of covalent enzyme-substrate analog complexes reveal strengths and limitations of de novo enzyme design.
J.Mol.Biol., 415, 2012
|
|
3NXF
 
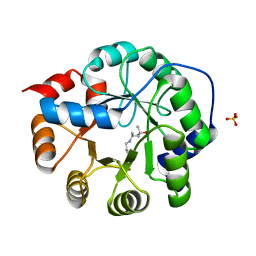 | | Robust computational design, optimization, and structural characterization of retroaldol enzymes | | Descriptor: | Retro-Aldolase, SULFATE ION | | Authors: | Althoff, E.A, Jiang, L, Wang, L, Lassila, J.K, Moody, J, Bolduc, J, Wang, Z.Z, Smith, M, Hari, S, Herschlag, D, Stoddard, B.L, Baker, D. | | Deposit date: | 2010-07-13 | | Release date: | 2011-06-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural analyses of covalent enzyme-substrate analog complexes reveal strengths and limitations of de novo enzyme design.
J.Mol.Biol., 415, 2012
|
|
2QC1
 
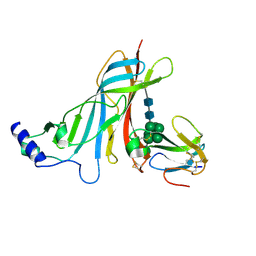 | | Crystal structure of the extracellular domain of the nicotinic acetylcholine receptor 1 subunit bound to alpha-bungarotoxin at 1.9 A resolution | | Descriptor: | Acetylcholine receptor subunit alpha, Alpha-bungarotoxin, alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-6)-[alpha-D-mannopyranose-(1-3)]alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Dellisanti, C.D, Yao, Y, Stroud, J.C, Wang, Z, Chen, L. | | Deposit date: | 2007-06-18 | | Release date: | 2007-08-07 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Crystal structure of the extracellular domain of nAChR alpha1 bound to alpha-bungarotoxin at 1.94 A resolution.
Nat.Neurosci., 10, 2007
|
|
4KK1
 
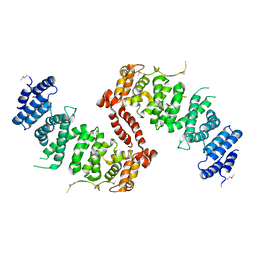 | | Crystal Structure of TSC1 core domain from S. pombe | | Descriptor: | Tuberous sclerosis 1 protein homolog | | Authors: | Sun, W, Zhu, Y, Wang, Z.Z, Zhong, Q, Gao, F, Lou, J.Z, Gong, W.M, Xu, W.Q. | | Deposit date: | 2013-05-05 | | Release date: | 2013-07-17 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Crystal structure of the yeast TSC1 core domain and implications for tuberous sclerosis pathological mutations.
Nat Commun, 4, 2013
|
|
4KK0
 
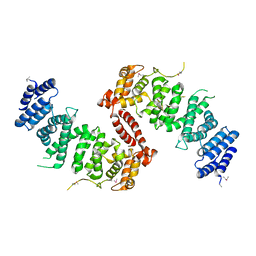 | | Crystal Structure of TSC1 core domain from S. pombe | | Descriptor: | Tuberous sclerosis 1 protein homolog | | Authors: | Sun, W, Zhu, Y, Wang, Z.Z, Zhong, Q, Gao, F, Lou, J.Z, Gong, W.M, Xu, W.Q. | | Deposit date: | 2013-05-05 | | Release date: | 2013-07-17 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of the yeast TSC1 core domain and implications for tuberous sclerosis pathological mutations.
Nat Commun, 4, 2013
|
|
