6MEP
 
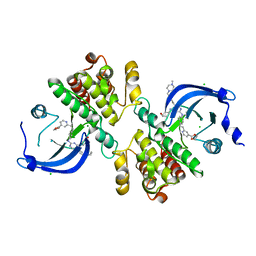 | | Crystal structure of the catalytic domain of the proto-oncogene tyrosine-protein kinase MER in complex with inhibitor UNC3437 | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, Tyrosine-protein kinase Mer, ... | | Authors: | Da, C, Zhang, D, Stashko, M.A, Cheng, A, Hunter, D, Norris-Drouin, J, Graves, L, Machius, M, Miley, M.J, DeRyckere, D, Earp, H.S, Graham, D.K, Frye, S.V, Wang, X, Kireev, D. | | Deposit date: | 2018-09-06 | | Release date: | 2019-09-11 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.893 Å) | | Cite: | Data-Driven Construction of Antitumor Agents with Controlled Polypharmacology.
J.Am.Chem.Soc., 141, 2019
|
|
2ETL
 
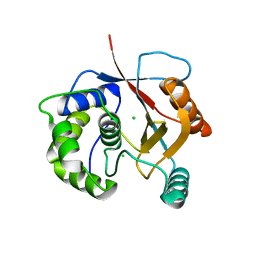 | | Crystal Structure of Ubiquitin Carboxy-terminal Hydrolase L1 (UCH-L1) | | Descriptor: | CHLORIDE ION, Ubiquitin carboxyl-terminal hydrolase isozyme L1 | | Authors: | Das, C, Hoang, Q.Q, Kreinbring, C.A, Luchansky, S.J, Meray, R.K, Ray, S.S, Lansbury, P.T, Ringe, D, Petsko, G.A. | | Deposit date: | 2005-10-27 | | Release date: | 2006-03-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for conformational plasticity of the Parkinson's disease-associated ubiquitin hydrolase UCH-L1.
Proc.Natl.Acad.Sci.USA, 103, 2006
|
|
4OUF
 
 | | Crystal Structure of CBP bromodomain | | Descriptor: | 1,2-ETHANEDIOL, CREB-binding protein, DI(HYDROXYETHYL)ETHER | | Authors: | Roy, S, Das, C, Tyler, J.K, Kutateladze, T.G. | | Deposit date: | 2014-02-17 | | Release date: | 2014-03-12 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Binding of the histone chaperone ASF1 to the CBP bromodomain promotes histone acetylation.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
3RII
 
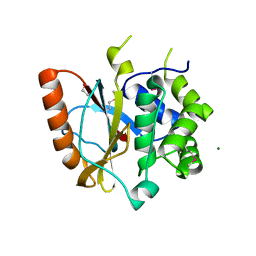 | | Crystal structure of the catalytic domain of UCHL5, a proteasome-associated human deubiquitinating enzyme, reveals an unproductive form of the enzyme | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, Ubiquitin carboxyl-terminal hydrolase isozyme L5 | | Authors: | Das, C. | | Deposit date: | 2011-04-13 | | Release date: | 2011-11-09 | | Last modified: | 2011-12-14 | | Method: | X-RAY DIFFRACTION (2.0008 Å) | | Cite: | Crystal structure of the catalytic domain of UCHL5, a proteasome-associated human deubiquitinating enzyme, reveals an unproductive form of the enzyme.
Febs J., 278, 2011
|
|
3RIS
 
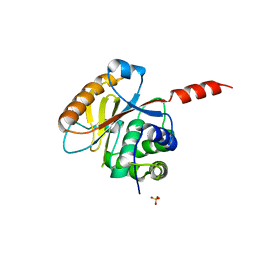 | | Crystal structure of the catalytic domain of UCHL5, a proteasome-associated human deubiquitinating enzyme, reveals an unproductive form of the enzyme | | Descriptor: | GLYCEROL, SULFATE ION, Ubiquitin carboxyl-terminal hydrolase isozyme L5 | | Authors: | Das, C, Permaul, M, Maiti, T.K. | | Deposit date: | 2011-04-14 | | Release date: | 2011-11-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.398 Å) | | Cite: | Crystal structure of the catalytic domain of UCHL5, a proteasome-associated human deubiquitinating enzyme, reveals an unproductive form of the enzyme.
Febs J., 278, 2011
|
|
4IG7
 
 | |
3IFW
 
 | |
6WTG
 
 | | SdeA DUB Domain in complex with Ubiquitin | | Descriptor: | Ubiquitin, Ubiquitinating/deubiquitinating enzyme SdeA | | Authors: | Kenny, S, Sheedlo, M, Das, C. | | Deposit date: | 2020-05-02 | | Release date: | 2021-03-03 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Insights into Ubiquitin Product Release in Hydrolysis Catalyzed by the Bacterial Deubiquitinase SdeA.
Biochemistry, 60, 2021
|
|
8DMU
 
 | |
8DMR
 
 | |
8EFW
 
 | |
8EFX
 
 | |
8FEK
 
 | |
8DMP
 
 | |
8DMS
 
 | |
8DMQ
 
 | |
8DMT
 
 | | Crystal structure of macrodomain CG2909 from Drosophila melanogaster in complex with ADP-ribose | | Descriptor: | RE54994p, [(2R,3S,4R,5R)-5-(6-AMINOPURIN-9-YL)-3,4-DIHYDROXY-OXOLAN-2-YL]METHYL [HYDROXY-[[(2R,3S,4R,5S)-3,4,5-TRIHYDROXYOXOLAN-2-YL]METHOXY]PHOSPHORYL] HYDROGEN PHOSPHATE | | Authors: | Zhang, Z, Das, C. | | Deposit date: | 2022-07-08 | | Release date: | 2023-08-02 | | Last modified: | 2024-07-24 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Legionella metaeffector MavL reverses ubiquitin ADP-ribosylation via a conserved arginine-specific macrodomain.
Nat Commun, 15, 2024
|
|
6OAM
 
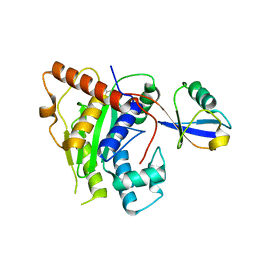 | | Crystal Structure of ChlaDUB2 DUB domain | | Descriptor: | Deubiquitinase and deneddylase Dub2, Ubiquitin | | Authors: | Hausman, J.M, Das, C. | | Deposit date: | 2019-03-17 | | Release date: | 2020-04-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.503 Å) | | Cite: | The Two Deubiquitinating Enzymes fromChlamydia trachomatisHave Distinct Ubiquitin Recognition Properties.
Biochemistry, 59, 2020
|
|
6OV1
 
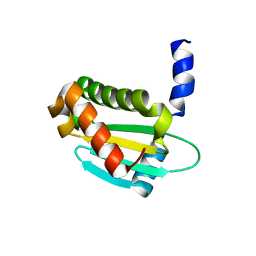 | | Structure of Staphylococcus aureus RNase P protein mutant with defective mRNA degradation activity | | Descriptor: | Ribonuclease P protein component | | Authors: | Ha, L, Colquhoun, J, Noinaj, N, Das, C, Dunman, P, Flaherty, D.P. | | Deposit date: | 2019-05-06 | | Release date: | 2020-12-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Genetic and biochemical characterization of Staphylococcus aureus RnpA
To Be Published
|
|
6P5H
 
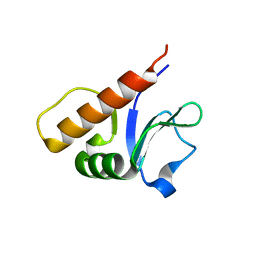 | |
6P5B
 
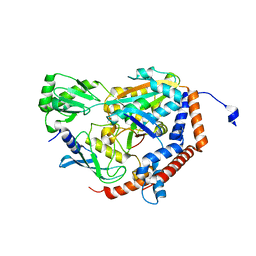 | | Crystal Structure of MavC in Complex with Ub-UbE2N | | Descriptor: | MavC, Ubiquitin, Ubiquitin-conjugating enzyme E2 N | | Authors: | Puvar, K, Iyer, S, Negron Teron, K.I, Das, C. | | Deposit date: | 2019-05-30 | | Release date: | 2020-05-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.099 Å) | | Cite: | Legionella effector MavC targets the Ube2N~Ub conjugate for noncanonical ubiquitination.
Nat Commun, 11, 2020
|
|
4Q3Y
 
 | |
4Q3Z
 
 | |
4Q3W
 
 | | Crystal structure of C. violaceum phenylalanine hydroxylase D139E mutation | | Descriptor: | 1,2-ETHANEDIOL, COBALT (II) ION, Phenylalanine-4-hydroxylase | | Authors: | Ronau, J.A, Abu-Omar, M.M, Das, C. | | Deposit date: | 2014-04-12 | | Release date: | 2015-02-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | A conserved acidic residue in phenylalanine hydroxylase contributes to cofactor affinity and catalysis.
Biochemistry, 53, 2014
|
|
4Q3X
 
 | |
