7VGI
 
 | |
7VGH
 
 | |
7XG7
 
 | | Crystal structure of PstS protein from cyanophage Syn19 | | Descriptor: | GLYCEROL, PHOSPHATE ION, Phosphate transporter subunit | | Authors: | Cai, K, Jiang, Y.L. | | Deposit date: | 2022-04-03 | | Release date: | 2023-02-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Biochemical and structural characterization of the cyanophage-encoded phosphate-binding protein: implications for enhanced phosphate uptake of infected cyanobacteria
Environ.Microbiol., 24, 2022
|
|
7XG8
 
 | | Crystal structure of PstS protein from cyanophage P-SSM2 | | Descriptor: | ABC transporter, substrate binding protein, phosphate, ... | | Authors: | Cai, K, Jiang, Y.L. | | Deposit date: | 2022-04-03 | | Release date: | 2023-02-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Biochemical and structural characterization of the cyanophage-encoded phosphate-binding protein: implications for enhanced phosphate uptake of infected cyanobacteria
Environ.Microbiol., 24, 2022
|
|
8JBR
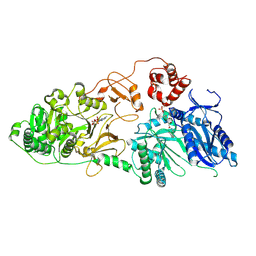 
 | | Structure of McyA2-CAPCP | | Descriptor: | 4'-PHOSPHOPANTETHEINE, ADENOSINE-5'-TRIPHOSPHATE, McyA protein | | Authors: | Peng, Y.J. | | Deposit date: | 2023-05-09 | | Release date: | 2024-01-24 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Modular catalytic activity of nonribosomal peptide synthetases depends on the dynamic interaction between adenylation and condensation domains.
Structure, 32, 2024
|
|
5Y46
 
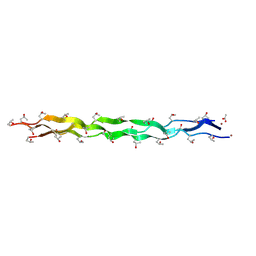 | |
5YFB
 
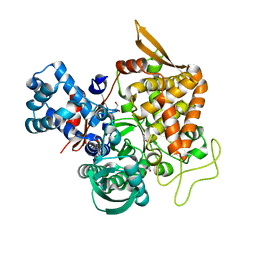 | | Crystal structure of a new DPP III family member | | Descriptor: | 1,2-ETHANEDIOL, Dipeptidyl peptidase 3, GLYCEROL, ... | | Authors: | Xu, T, Sun, Z, Liu, J. | | Deposit date: | 2017-09-20 | | Release date: | 2018-01-03 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structures of Aflatoxin-oxidase from Armillariella tabescens reveal a dual activity enzyme.
Biochem. Biophys. Res. Commun., 494, 2017
|
|
5Y45
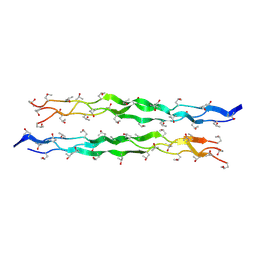 
 | |
5YFC
 
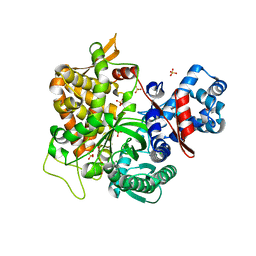 | | Crystal structure of a new DPP III family member | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, Dipeptidyl peptidase 3, ... | | Authors: | Xu, T, Liu, J. | | Deposit date: | 2017-09-20 | | Release date: | 2018-01-03 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Crystal structures of Aflatoxin-oxidase from Armillariella tabescens reveal a dual activity enzyme.
Biochem. Biophys. Res. Commun., 494, 2017
|
|
5YFD
 
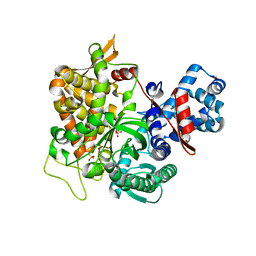 | | Crystal structure of a new DPP III family member | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, COPPER (II) ION, Dipeptidyl peptidase 3, ... | | Authors: | Xu, T, Liu, J. | | Deposit date: | 2017-09-20 | | Release date: | 2018-01-03 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Crystal structures of Aflatoxin-oxidase from Armillariella tabescens reveal a dual activity enzyme.
Biochem. Biophys. Res. Commun., 494, 2017
|
|
8HLK
 
 | | Structure of McyB-C1A1 complexed with L-Leu and AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, LEUCINE, Microcystin synthetase B (Fragment) | | Authors: | Peng, Y.J. | | Deposit date: | 2022-11-30 | | Release date: | 2023-12-13 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Modular catalytic activity of nonribosomal peptide synthetases depends on the dynamic interaction between adenylation and condensation domains.
Structure, 32, 2024
|
|
6L5D
 
 | |
6LNW
 
 | | Crystal structure of accessory secretory protein 1,2 and 3 in Streptococcus pneumoniae | | Descriptor: | Accessory secretory protein Asp1, Accessory secretory protein Asp2, Accessory secretory protein Asp3 | | Authors: | Guo, C, Feng, Z, Zuo, G. | | Deposit date: | 2020-01-02 | | Release date: | 2020-02-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural and functional insights into the Asp1/2/3 complex mediated secretion of pneumococcal serine-rich repeat protein PsrP.
Biochem.Biophys.Res.Commun., 524, 2020
|
|
7DV5
 
 | |
7E1A
 
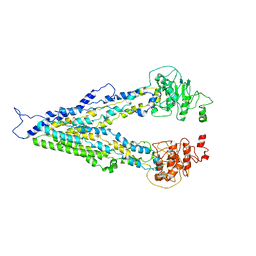 | |
4PVC
 
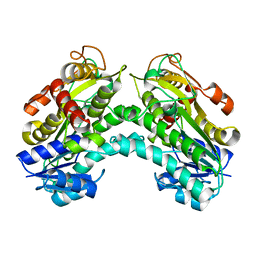 | |
4PVD
 
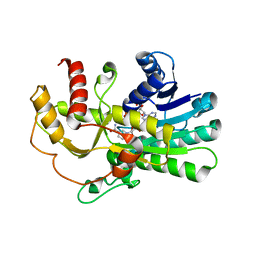 | | Crystal structure of yeast methylglyoxal/isovaleraldehyde reductase Gre2 complexed with NADPH | | Descriptor: | NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NADPH-dependent methylglyoxal reductase GRE2 | | Authors: | Guo, P.C, Bao, Z.Z, Li, W.F, Zhou, C.Z. | | Deposit date: | 2014-03-17 | | Release date: | 2014-10-22 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural insights into the cofactor-assisted substrate recognition of yeast methylglyoxal/isovaleraldehyde reductase Gre2
Biochim.Biophys.Acta, 1844, 2014
|
|
