9KWV
 
 | | Structure of a D98N/C135A/G136A mutant copper-containing nitrite reductase in complex with nitrite | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CHLORIDE ION, COPPER (II) ION, ... | | Authors: | Fukuda, Y, Lintuluoto, M, Hirano, Y, Kusaka, K, Inoue, T, Tamada, T. | | Deposit date: | 2024-12-06 | | Release date: | 2025-06-11 | | Last modified: | 2025-07-02 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Structural basis of cuproenzyme nitrite reduction at the level of a single hydrogen atom.
J.Biol.Chem., 301, 2025
|
|
9KVL
 
 | | Neutron and X-ray joint refined structure of a copper-containing nitrite reductase (C135A mutant) in complex with nitrite | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, COPPER (II) ION, ... | | Authors: | Fukuda, Y, Lintuluoto, M, Hirano, Y, Kusaka, K, Inoue, T, Tamada, T. | | Deposit date: | 2024-12-05 | | Release date: | 2025-06-11 | | Last modified: | 2025-07-02 | | Method: | NEUTRON DIFFRACTION (1.7 Å), X-RAY DIFFRACTION | | Cite: | Structural basis of cuproenzyme nitrite reduction at the level of a single hydrogen atom.
J.Biol.Chem., 301, 2025
|
|
9KWS
 
 | | D98N mutant of a copper-containing nitrite reductase from Geobacillus thermodenitrificans | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ACETIC ACID, CHLORIDE ION, ... | | Authors: | Fukuda, Y, Lintuluoto, M, Hirano, Y, Kusaka, K, Inoue, T. | | Deposit date: | 2024-12-06 | | Release date: | 2025-06-11 | | Last modified: | 2025-07-02 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Structural basis of cuproenzyme nitrite reduction at the level of a single hydrogen atom.
J.Biol.Chem., 301, 2025
|
|
9KWU
 
 | | D98N/C135A mutant of a copper-containing nitrite reductase in complex with nitrite | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, COPPER (II) ION, Copper-containing nitrite reductase, ... | | Authors: | Fukuda, Y, Lintuluoto, M, Hirano, Y, Kusaka, K, Inoue, T, Tamada, T. | | Deposit date: | 2024-12-06 | | Release date: | 2025-06-11 | | Last modified: | 2025-07-02 | | Method: | X-RAY DIFFRACTION (1.12 Å) | | Cite: | Structural basis of cuproenzyme nitrite reduction at the level of a single hydrogen atom.
J.Biol.Chem., 301, 2025
|
|
8K9N
 
 | | Subatomic resolution structure of Pseudoazurin from Alcaligenes faecalis | | Descriptor: | COPPER (II) ION, Pseudoazurin, SULFATE ION | | Authors: | Fukuda, Y, Lintuluoto, M, Kurihara, K, Hasegawa, K, Inoue, T, Tamada, T. | | Deposit date: | 2023-08-01 | | Release date: | 2024-02-14 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (0.86 Å) | | Cite: | Overlooked Hydrogen Bond in a Blue Copper Protein Uncovered by Neutron and Sub- angstrom ngstrom Resolution X-ray Crystallography.
Biochemistry, 63, 2024
|
|
5KF7
 
 | |
5KF6
 
 | |
4H6Q
 
 | |
8DW9
 
 | |
8DXS
 
 | |
8DWA
 
 | |
7YKJ
 
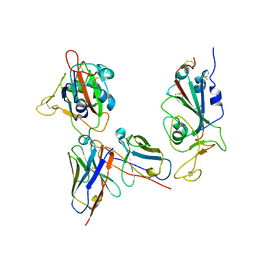 | | Omicron RBDs bound with P3E6 Fab (one up and one down) | | Descriptor: | P3E6 heavy chain, P3E6 light chain, Spike glycoprotein | | Authors: | Tang, B, Dang, S. | | Deposit date: | 2022-07-22 | | Release date: | 2022-12-28 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structural insights into broadly neutralizing antibodies elicited by hybrid immunity against SARS-CoV-2.
Emerg Microbes Infect, 12, 2023
|
|
4ZVW
 
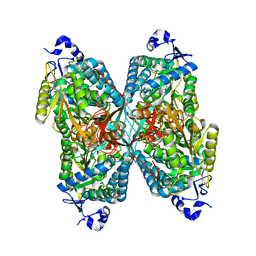 | | Structure of apo human ALDH7A1 in space group C2 | | Descriptor: | Alpha-aminoadipic semialdehyde dehydrogenase | | Authors: | Tanner, J.J. | | Deposit date: | 2015-05-18 | | Release date: | 2015-08-26 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural Basis of Substrate Recognition by Aldehyde Dehydrogenase 7A1.
Biochemistry, 54, 2015
|
|
4ZVX
 
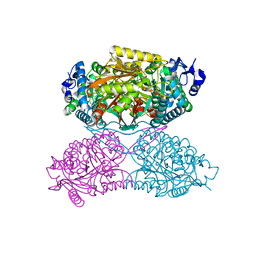 | | Structure of apo human ALDH7A1 in space group P4212 | | Descriptor: | Alpha-aminoadipic semialdehyde dehydrogenase | | Authors: | Tanner, J.J. | | Deposit date: | 2015-05-18 | | Release date: | 2015-08-26 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Basis of Substrate Recognition by Aldehyde Dehydrogenase 7A1.
Biochemistry, 54, 2015
|
|
4ZVY
 
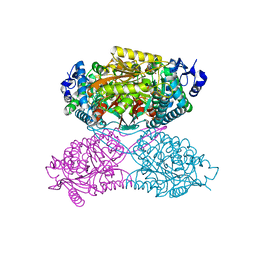 | |
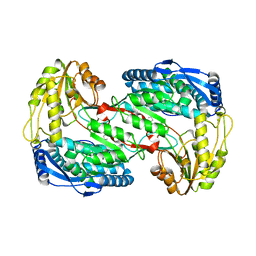
4K57
 
 | |
8K19
 
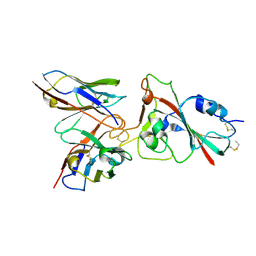 | |
4H6R
 
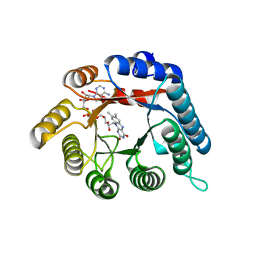 | | Structure of reduced Deinococcus radiodurans proline dehydrogenase | | Descriptor: | ACETATE ION, DIHYDROFLAVINE-ADENINE DINUCLEOTIDE, Proline dehydrogenase | | Authors: | Min, L, Tanner, J.J. | | Deposit date: | 2012-09-19 | | Release date: | 2012-11-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structures and kinetics of monofunctional proline dehydrogenase provide insight into substrate recognition and conformational changes associated with flavin reduction and product release.
Biochemistry, 51, 2012
|
|
2HWC
 
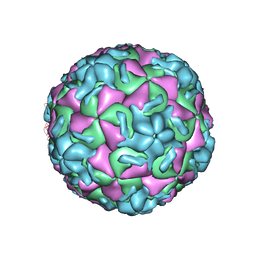 | | A COMPARISON OF THE ANTI-RHINOVIRAL DRUG BINDING POCKET IN HRV14 AND HRV1A | | Descriptor: | 5-(5-(2,6-DICHLORO-4-(4,5-DIHYDRO-2-OXAZOLY)PHENOXY)PENTYL)-3-METHYL ISOXAZOLE, HUMAN RHINOVIRUS 14 COAT PROTEIN (SUBUNIT VP1), HUMAN RHINOVIRUS 14 COAT PROTEIN (SUBUNIT VP2), ... | | Authors: | Kim, K.H, Rossmann, M.G. | | Deposit date: | 1994-01-25 | | Release date: | 1994-11-01 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | A comparison of the anti-rhinoviral drug binding pocket in HRV14 and HRV1A.
J.Mol.Biol., 230, 1993
|
|
2HWD
 
 | | A COMPARISON OF THE ANTI-RHINOVIRAL DRUG BINDING POCKET IN HRV14 AND HRV1A | | Descriptor: | 5-(3-(2,6-DICHLORO-4-(4,5-DIHYDRO-2-OXAZOLYL)PHENOXY)PROPYL)-3-METHYL ISOXAZOLE, HUMAN RHINOVIRUS 1A COAT PROTEIN (SUBUNIT VP1), HUMAN RHINOVIRUS 1A COAT PROTEIN (SUBUNIT VP2), ... | | Authors: | Kim, K.H, Rossmann, M.G. | | Deposit date: | 1994-01-25 | | Release date: | 1994-09-30 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | A comparison of the anti-rhinoviral drug binding pocket in HRV14 and HRV1A.
J.Mol.Biol., 230, 1993
|
|
2HWB
 
 | | A comparison of the anti-rhinoviral drug binding pocket in hrv14 and hrv1a | | Descriptor: | 5-(3-(2,6-DICHLORO-4-(4,5-DIHYDRO-2-OXAZOLYL)PHENOXY)PROPYL)-3-METHYL ISOXAZOLE, HUMAN RHINOVIRUS 14 COAT PROTEIN (SUBUNIT VP1), HUMAN RHINOVIRUS 14 COAT PROTEIN (SUBUNIT VP2), ... | | Authors: | Kim, K.H, Rossmann, M.G. | | Deposit date: | 1994-01-25 | | Release date: | 1994-11-01 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | A comparison of the anti-rhinoviral drug binding pocket in HRV14 and HRV1A.
J.Mol.Biol., 230, 1993
|
|
2HWF
 
 | |
2HWE
 
 | | A COMPARISON OF THE ANTI-RHINOVIRAL DRUG BINDING POCKET IN HRV14 AND HRV1A | | Descriptor: | 5-(5-(2,6-DICHLORO-4-(4,5-DIHYDRO-2-OXAZOLY)PHENOXY)PENTYL)-3-METHYL ISOXAZOLE, HUMAN RHINOVIRUS 1A COAT PROTEIN (SUBUNIT VP1), HUMAN RHINOVIRUS 1A COAT PROTEIN (SUBUNIT VP2), ... | | Authors: | Kim, K.H, Rossmann, M.G. | | Deposit date: | 1994-01-25 | | Release date: | 1994-09-30 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | A comparison of the anti-rhinoviral drug binding pocket in HRV14 and HRV1A.
J.Mol.Biol., 230, 1993
|
|
1MEC
 
 | |
1R08
 
 | | STRUCTURAL ANALYSIS OF ANTIVIRAL AGENTS THAT INTERACT WITH THE CAPSID OF HUMAN RHINOVIRUSES | | Descriptor: | 5-(5-(2,6-DICHLORO-4-(4,5-DIHYDRO-2-OXAZOLYL)PHENOXY)PENTYL)-3-(HYDROXYETHYL OXYMETHYLENEOXYMETHYL) ISOXAZOLE, HUMAN RHINOVIRUS 14 COAT PROTEIN (SUBUNIT VP1), HUMAN RHINOVIRUS 14 COAT PROTEIN (SUBUNIT VP2), ... | | Authors: | Badger, J, Smith, T.J, Rossmann, M.G. | | Deposit date: | 1988-10-03 | | Release date: | 1990-01-15 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural analysis of antiviral agents that interact with the capsid of human rhinoviruses.
Proteins, 6, 1989
|
|
