6EWW
 
 | |
6FEL
 
 | |
5O6V
 | |
5FS5
 
 | |
5FWR
 
 | |
5FX3
 | |
6GND
 
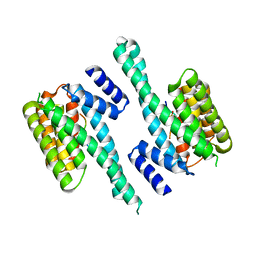
 | | Crystal structure of the complex of a Ferredoxin-Flavin Thioredoxin Reductase and a Thioredoxin from Clostridium acetobutylicum at 2.9 A resolution | | Descriptor: | 1,2-ETHANEDIOL, FLAVIN-ADENINE DINUCLEOTIDE, Thioredoxin, ... | | Authors: | Buey, R.M, Fernandez-Justel, D, Balsera, M. | | Deposit date: | 2018-05-30 | | Release date: | 2018-12-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.889 Å) | | Cite: | Ferredoxin-linked flavoenzyme defines a family of pyridine nucleotide-independent thioredoxin reductases.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6GKF
 
 | |
6GNB
 
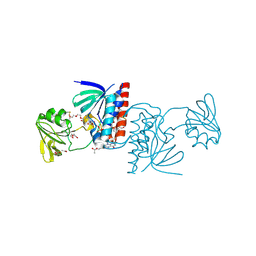 | | Crystal structure of a Ferredoxin-Flavin Thioredoxin Reductase from Clostridium acetobutylicum at 1.9 A resolution | | Descriptor: | ACETATE ION, DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Buey, R.M, Fernandez-Justel, D, Balsera, M. | | Deposit date: | 2018-05-30 | | Release date: | 2018-12-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.895 Å) | | Cite: | Ferredoxin-linked flavoenzyme defines a family of pyridine nucleotide-independent thioredoxin reductases.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6GN9
 
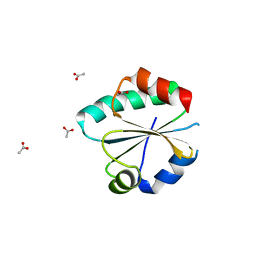 | |
6GNC
 
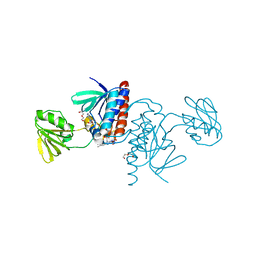 | |
6GKE
 
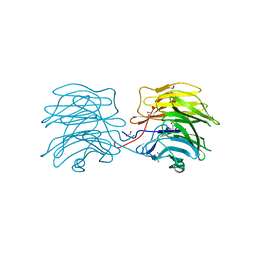 | | Aleuria aurantia lectin AAL N224Q mutant in complex with Fucalpha1-6GlcNAc | | Descriptor: | 1,2-ETHANEDIOL, Fucose-specific lectin, GLYCEROL, ... | | Authors: | Houser, J, Wimmerova, M, Herrera, H, Mehta, A.S. | | Deposit date: | 2018-05-19 | | Release date: | 2019-06-19 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | Structural dynamics and applications of a fucose-binding lectin with enhanced binding to alpha-1,6-linked core fucosylation
To Be Published
|
|
5O6A
 
 | |
6GNA
 
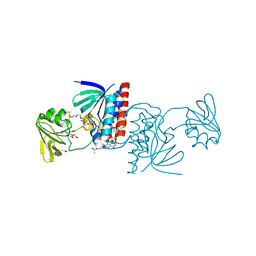 | | Crystal structure of a Ferredoxin-Flavin Thioredoxin Reductase from Clostridium acetobutylicum at 1.3 A resolution | | Descriptor: | ACETATE ION, DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Buey, R.M, Fernandez-Justel, D, Balsera, M. | | Deposit date: | 2018-05-30 | | Release date: | 2018-12-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.295 Å) | | Cite: | Ferredoxin-linked flavoenzyme defines a family of pyridine nucleotide-independent thioredoxin reductases.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6GKG
 
 | |
6F45
 
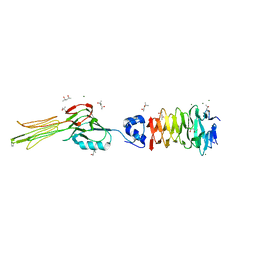 | | Crystal structure of the gp37-gp38 adhesin tip complex of the bacteriophage S16 long tail fiber | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, IMIDAZOLE, ... | | Authors: | Dunne, M, Leiman, P, Klumpp, J, Loessner, M.J. | | Deposit date: | 2017-11-29 | | Release date: | 2018-08-01 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.70355821 Å) | | Cite: | Salmonella Phage S16 Tail Fiber Adhesin Features a Rare Polyglycine Rich Domain for Host Recognition.
Structure, 26, 2018
|
|
5TOZ
 
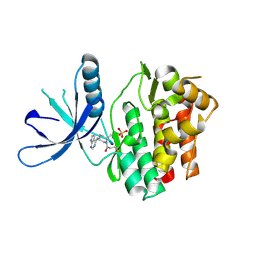 | | JAK3 with covalent inhibitor PF-06651600 | | Descriptor: | 1-{(2S,5R)-2-methyl-5-[(7H-pyrrolo[2,3-d]pyrimidin-4-yl)amino]piperidin-1-yl}propan-1-one, SULFATE ION, Tyrosine-protein kinase JAK3 | | Authors: | Vajdos, F.F. | | Deposit date: | 2016-10-19 | | Release date: | 2016-11-09 | | Last modified: | 2017-11-22 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Discovery of a JAK3-Selective Inhibitor: Functional Differentiation of JAK3-Selective Inhibition over pan-JAK or JAK1-Selective Inhibition.
ACS Chem. Biol., 11, 2016
|
|
7K0S
 | |
4CA4
 | | Crystal structure of FimH lectin domain with the Tyr48Ala mutation, in complex with heptyl alpha-D-mannopyrannoside | | Descriptor: | FIMH, heptyl alpha-D-mannopyranoside | | Authors: | Rabbani, S, Bouckaert, J, Zalewski, A, Preston, R, Eid, S, Thompson, A, Puorger, C, Glockshuber, R, Ernst, B. | | Deposit date: | 2013-10-06 | | Release date: | 2014-10-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Mutation of Tyr137 of the universal Escherichia coli fimbrial adhesin FimH relaxes the tyrosine gate prior to mannose binding.
IUCrJ, 4, 2017
|
|
4BYJ
 
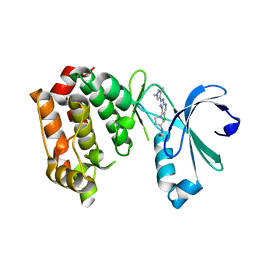 | | Aurora A kinase bound to a highly selective imidazopyridine inhibitor | | Descriptor: | (S)-N-(1-(6-chloro-2-(1,3-dimethyl-1H-pyrazol-4-yl)-3H-imidazo[4,5-b]pyridin-7-yl)pyrrolidin-3-yl)acetamide, AURORA KINASE A | | Authors: | Joshi, A, Kosmopoulou, M, Bayliss, R. | | Deposit date: | 2013-07-19 | | Release date: | 2013-11-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Aurora Isoform Selectivity: Design and Synthesis of Imidazo[4,5-B]Pyridine Derivatives as Highly Selective Inhibitors of Aurora-A Kinase in Cells.
J.Med.Chem., 56, 2013
|
|
4BYI
 
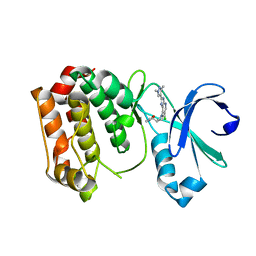 | | Aurora A kinase bound to a highly selective imidazopyridine inhibitor | | Descriptor: | (S)-N-((1-(6-chloro-2-(1,3-dimethyl-1H-pyrazol-4-yl)-3H-imidazo[4,5-b]pyridin-7-yl)pyrrolidin-3-yl)methyl)acetamide, AURORA KINASE A | | Authors: | Joshi, A, Kosmopoulou, M, Bayliss, R. | | Deposit date: | 2013-07-19 | | Release date: | 2013-11-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Aurora Isoform Selectivity: Design and Synthesis of Imidazo[4,5-B]Pyridine Derivatives as Highly Selective Inhibitors of Aurora-A Kinase in Cells.
J.Med.Chem., 56, 2013
|
|
6AN6
 
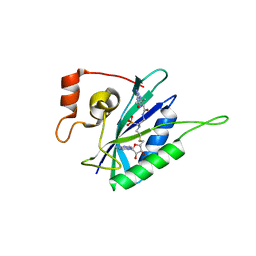 | | Crystal structure of Escherichia coli HPPK in complex with bisubstrate analogue inhibitor HP-72 | | Descriptor: | 2-amino-4-hydroxy-6-hydroxymethyldihydropteridine pyrophosphokinase, 5'-S-{2-[{2-[(2-amino-7,7-dimethyl-4-oxo-3,4,7,8-tetrahydropteridine-6-carbonyl)amino]ethyl}(phosphonomethyl)amino]ethyl}-5'-thioadenosine | | Authors: | Shaw, G.X, Shi, G, Ji, X. | | Deposit date: | 2017-08-12 | | Release date: | 2018-08-15 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Bisubstrate analogue inhibitors of HPPK: Transition state mimetics
to be published
|
|
4B0G
 
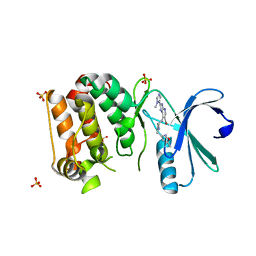 | | Complex of Aurora-A bound to an Imidazopyridine-based inhibitor | | Descriptor: | 6-bromo-2-(1-methyl-1H-imidazol-5-yl)-7-{4-[(5-methyl-1,2-oxazol-3-yl)methyl]piperazin-1-yl}-1H-imidazo[4,5-b]pyridine, AURORA KINASE A, SULFATE ION | | Authors: | Kosmopoulou, M, Bayliss, R. | | Deposit date: | 2012-07-02 | | Release date: | 2013-03-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Optimization of Imidazo[4,5-B]Pyridine-Based Kinase Inhibitors: Identification of a Dual Flt3/Aurora Kinase Inhibitor as an Orally Bioavailable Preclinical Development Candidate for the Treatment of Acute Myeloid Leukemia.
J.Med.Chem., 55, 2012
|
|
5GSU
 
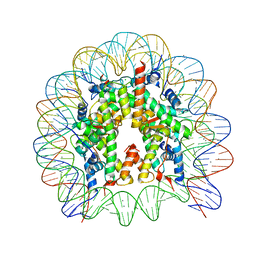 | | Crystal structure of nucleosome core particle consisting of human testis-specific histone variants, Th2A and Th2B | | Descriptor: | CHLORIDE ION, DNA (146-MER), Histone H2A type 1-A, ... | | Authors: | Kumarevel, T, Sivaraman, P. | | Deposit date: | 2016-08-17 | | Release date: | 2017-02-15 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural analyses of the nucleosome complexes with human testis-specific histone variants, hTh2a and hTh2b
Biophys. Chem., 221, 2017
|
|
6AN4
 
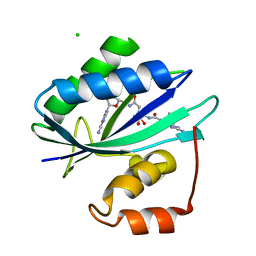 | | Crystal structure of Escherichia coli HPPK in complex with bisubstrate analogue inhibitor HP-39 (J1F) | | Descriptor: | ((2-(2-amino-7,7-dimethyl-4-oxo-3,4,7,8-tetrahydropteridine-6-carboxamido)-N-(2-((((2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxytetrahydrofuran-2-yl)methyl)amino)-2-oxoethyl)acetamido)methyl)phosphonic acid, 2-amino-4-hydroxy-6-hydroxymethyldihydropteridine pyrophosphokinase, CHLORIDE ION | | Authors: | Shaw, G.X, Shi, G, Ji, X. | | Deposit date: | 2017-08-12 | | Release date: | 2018-08-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Bisubstrate analog inhibitors of HPPK: Transition state mimetics
to be published
|
|
