7ES2
 
 | | a mutant of glycosyktransferase in complex with UDP and Reb D | | 分子名称: | Glycosyltransferase, URIDINE-5'-DIPHOSPHATE, rebaudioside D | | 著者 | Zhu, X. | | 登録日 | 2021-05-08 | | 公開日 | 2021-12-08 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (2.34 Å) | | 主引用文献 | Catalytic flexibility of rice glycosyltransferase OsUGT91C1 for the production of palatable steviol glycosides.
Nat Commun, 12, 2021
|
|
7ERX
 
 | | Glycosyltransferase in complex with UDP and STB | | 分子名称: | GLYCEROL, Glycosyltransferase, Steviolbioside, ... | | 著者 | Zhu, X. | | 登録日 | 2021-05-08 | | 公開日 | 2021-12-08 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.92 Å) | | 主引用文献 | Catalytic flexibility of rice glycosyltransferase OsUGT91C1 for the production of palatable steviol glycosides.
Nat Commun, 12, 2021
|
|
7ES1
 
 | | glycosyltransferase in complex with UDP and ST | | 分子名称: | Glycosyltransferase, URIDINE-5'-DIPHOSPHATE, steviol-19-o-glucoside | | 著者 | Zhu, X. | | 登録日 | 2021-05-08 | | 公開日 | 2021-12-08 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (1.66 Å) | | 主引用文献 | Catalytic flexibility of rice glycosyltransferase OsUGT91C1 for the production of palatable steviol glycosides.
Nat Commun, 12, 2021
|
|
7ES0
 
 | | a rice glycosyltransferase in complex with UDP and REX | | 分子名称: | 1,2-ETHANEDIOL, 3[N-MORPHOLINO]PROPANE SULFONIC ACID, GLYCEROL, ... | | 著者 | Zhu, X. | | 登録日 | 2021-05-08 | | 公開日 | 2021-12-08 | | 最終更新日 | 2024-05-29 | | 実験手法 | X-RAY DIFFRACTION (1.395 Å) | | 主引用文献 | Catalytic flexibility of rice glycosyltransferase OsUGT91C1 for the production of palatable steviol glycosides.
Nat Commun, 12, 2021
|
|
6J1O
 
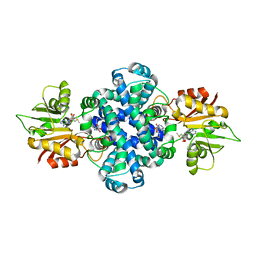 | |
6J46
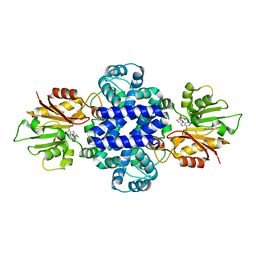 
 | | LepI-SAH complex structure | | 分子名称: | O-methyltransferase lepI, S-ADENOSYL-L-HOMOCYSTEINE | | 著者 | Qiu, S, Wei, C. | | 登録日 | 2019-01-08 | | 公開日 | 2019-05-01 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (2.621 Å) | | 主引用文献 | Deciphering the regulatory and catalytic mechanisms of an unusual SAM-dependent enzyme.
Signal Transduct Target Ther, 4, 2019
|
|
6J9E
 
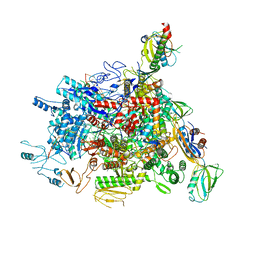 | |
6J24
 
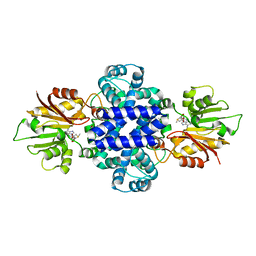 | | Crystal structure of a SAM-dependent methyltransferase LepI in complex with its substrate | | 分子名称: | (3~{S},4'~{R},4'~{a}~{S},6'~{R},8'~{a}~{S})-4',6'-dimethyl-5-phenyl-spiro[1~{H}-pyridine-3,5'-2,3,4,4~{a},6,8~{a}-hexahydro-1~{H}-naphthalene]-2,4-dione, O-methyltransferase, S-ADENOSYLMETHIONINE | | 著者 | Qiu, S, Wei, C. | | 登録日 | 2018-12-30 | | 公開日 | 2019-05-01 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.244 Å) | | 主引用文献 | Deciphering the regulatory and catalytic mechanisms of an unusual SAM-dependent enzyme.
Signal Transduct Target Ther, 4, 2019
|
|
6J3P
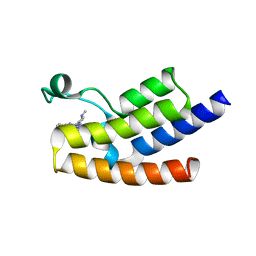 
 | | Crystal structure of the human GCN5 bromodomain in complex with compound (R,R)-36n | | 分子名称: | 2-{[(3R,5R)-5-(2,3-dihydro-1,4-benzodioxin-6-yl)-1-methylpiperidin-3-yl]amino}-3-methyl-3,5-dihydro-4H-pyrrolo[3,2-d]pyrimidin-4-one, Histone acetyltransferase KAT2A | | 著者 | Huang, L.Y, Li, H, Niu, L, Wu, C.Y, Yu, Y.M, Li, L.L, Yang, S.Y. | | 登録日 | 2019-01-05 | | 公開日 | 2019-05-01 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.598 Å) | | 主引用文献 | Discovery of Pyrrolo[3,2- d]pyrimidin-4-one Derivatives as a New Class of Potent and Cell-Active Inhibitors of P300/CBP-Associated Factor Bromodomain.
J.Med.Chem., 62, 2019
|
|
6J3O
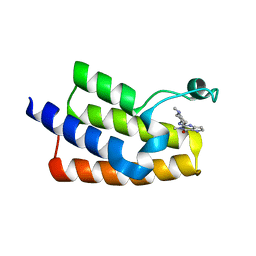 
 | | Crystal structure of the human PCAF bromodomain in complex with compound 12 | | 分子名称: | 3-methyl-2-[[(3~{R})-1-methylpiperidin-3-yl]amino]-5~{H}-pyrrolo[3,2-d]pyrimidin-4-one, Histone acetyltransferase KAT2B | | 著者 | Huang, L.Y, Li, H, Li, L.L, Niu, L, Seupel, R, Wu, C.Y, Li, G.B, Yu, Y.M, Brennan, P.E, Yang, S.Y. | | 登録日 | 2019-01-05 | | 公開日 | 2019-05-01 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.11 Å) | | 主引用文献 | Discovery of Pyrrolo[3,2- d]pyrimidin-4-one Derivatives as a New Class of Potent and Cell-Active Inhibitors of P300/CBP-Associated Factor Bromodomain.
J.Med.Chem., 62, 2019
|
|
6J9F
 
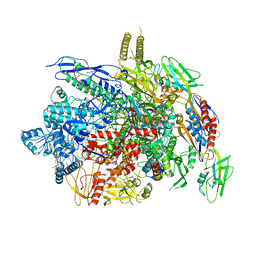 | |
6KYB
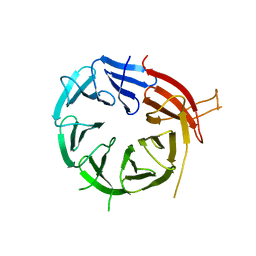 
 | | Crystal structure of Atg18 from Saccharomyces cerevisiae | | 分子名称: | Autophagy-related protein 18 | | 著者 | Tang, D, Lei, Y, Liao, G, Chen, Q, Xu, L, Lu, K, Qi, S. | | 登録日 | 2019-09-17 | | 公開日 | 2020-09-09 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | The crystal structure of Atg18 reveals a new binding site for Atg2 in Saccharomyces cerevisiae.
Cell.Mol.Life Sci., 78, 2021
|
|
8JYD
 
 | |
7EGV
 
 | | Acetolactate Synthase from Trichoderma harzianum with inhibitor harzianic acid | | 分子名称: | (2S)-3-methyl-2-[[(2S,4R)-1-methyl-4-[(2E,4E)-octa-2,4-dienoyl]-3,5-bis(oxidanylidene)pyrrolidin-2-yl]methyl]-2-oxidanyl-butanoic acid, 1,2-ETHANEDIOL, 2-{3-[(4-AMINO-2-METHYLPYRIMIDIN-5-YL)METHYL]-4-METHYL-2-OXO-2,3-DIHYDRO-1,3-THIAZOL-5-YL}ETHYL TRIHYDROGEN DIPHOSPHATE, ... | | 著者 | Zang, X, Xie, L, Chen, M, Tang, Y, Zhou, J. | | 登録日 | 2021-03-26 | | 公開日 | 2021-06-30 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.54 Å) | | 主引用文献 | Harzianic Acid from Trichoderma afroharzianum Is a Natural Product Inhibitor of Acetohydroxyacid Synthase.
J.Am.Chem.Soc., 2021
|
|
7EHE
 
 | | Acetolactate Synthase from Trichoderma harzianum | | 分子名称: | 2-{3-[(4-AMINO-2-METHYLPYRIMIDIN-5-YL)METHYL]-4-METHYL-2-OXO-2,3-DIHYDRO-1,3-THIAZOL-5-YL}ETHYL TRIHYDROGEN DIPHOSPHATE, Acetolactate synthase, FLAVIN-ADENINE DINUCLEOTIDE, ... | | 著者 | Zang, X, Tang, Y, Zhou, J. | | 登録日 | 2021-03-29 | | 公開日 | 2021-06-30 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.28 Å) | | 主引用文献 | Harzianic Acid from Trichoderma afroharzianum Is a Natural Product Inhibitor of Acetohydroxyacid Synthase.
J.Am.Chem.Soc., 2021
|
|
8ZT3
 | |
8ZT4
 
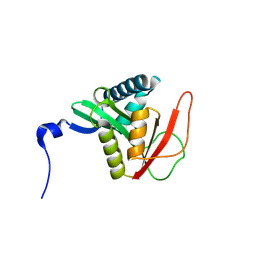
 | | Complex structure of N-acetyltransferase SbzI and carrier protein SbzG in the biosynthesis of altemicidin | | 分子名称: | BICARBONATE ION, CHLORIDE ION, Carrier protein,GNAT family transferase, ... | | 著者 | Zhu, Y, Mori, T, Abe, I. | | 登録日 | 2024-06-06 | | 公開日 | 2025-02-05 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Structure-function analysis of carrier protein-dependent 2-sulfamoylacetyl transferase in the biosynthesis of altemicidin.
Nat Commun, 15, 2024
|
|
7C8U
 
 | | The crystal structure of COVID-19 main protease in complex with GC376 | | 分子名称: | (1S,2S)-2-({N-[(benzyloxy)carbonyl]-L-leucyl}amino)-1-hydroxy-3-[(3S)-2-oxopyrrolidin-3-yl]propane-1-sulfonic acid, 3C-like proteinase | | 著者 | Luan, X, Shang, W, Wang, Y, Yin, W, Jiang, Y, Feng, S, Wang, Y, Liu, M, Zhou, R, Zhang, Z, Wang, F, Cheng, W, Gao, M, Wang, H, Wu, W, Tian, R, Tian, Z, Jin, Y, Jiang, H.W, Zhang, L, Xu, H.E, Zhang, S. | | 登録日 | 2020-06-03 | | 公開日 | 2020-06-24 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (2.35 Å) | | 主引用文献 | The crystal structure of COVID-19 main protease in complex with GC376
To Be Published
|
|
7XR0
 
 | | Crystal structure of T2R-TTL-27a complex | | 分子名称: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-chloranyl-5-fluoranyl-N-(4-methoxyphenyl)-N-methyl-quinazolin-4-amine, CALCIUM ION, ... | | 著者 | Lun, T, Wu, C.Y. | | 登録日 | 2022-05-09 | | 公開日 | 2023-05-17 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | Crystal structure of T2R-TTL-27a complex
To Be Published
|
|
7XR1
 
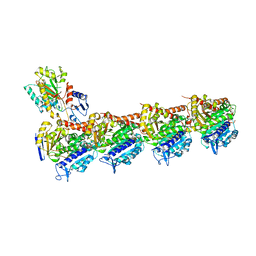 | | Crystal structure of T2R-TTL-3a complex | | 分子名称: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-chloranyl-6-fluoranyl-N-(4-methoxyphenyl)-N-methyl-quinazolin-4-amine, CALCIUM ION, ... | | 著者 | Lun, T, Wu, C.Y. | | 登録日 | 2022-05-09 | | 公開日 | 2023-05-17 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.81 Å) | | 主引用文献 | Crystal structure of T2R-TTL-3a complex
To Be Published
|
|
8WJB
 
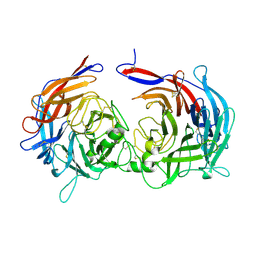 | |
6AIC
 
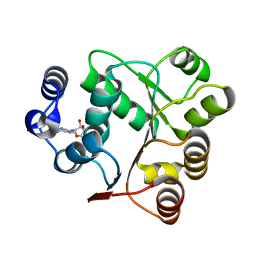 | | Crystal structures of the N-terminal domain of Staphylococcus aureus DEAD-box Cold shock RNA helicase CshA in complex with AMP | | 分子名称: | ADENOSINE MONOPHOSPHATE, DEAD-box ATP-dependent RNA helicase CshA | | 著者 | Tian, T, Chengliang, W, Xiaobao, C, Xuan, Z, Jianye, Z. | | 登録日 | 2018-08-22 | | 公開日 | 2018-11-21 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Crystal structures of the N-terminal domain of the Staphylococcus aureus DEAD-box RNA helicase CshA and its complex with AMP
Acta Crystallogr F Struct Biol Commun, 74, 2018
|
|
7CAJ
 
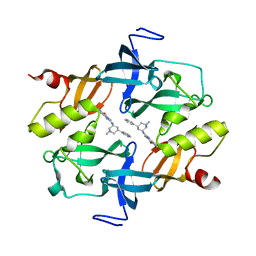 | | Crystal structure of SETDB1 Tudor domain in complexed with Compound 2. | | 分子名称: | 3-methyl-2-[[(3R,5R)-1-methyl-5-phenyl-piperidin-3-yl]amino]-5H-pyrrolo[3,2-d]pyrimidin-4-one, Histone-lysine N-methyltransferase SETDB1 | | 著者 | Guo, Y.P, Liang, X, Xin, M, Luyi, H, Chengyong, W, Yang, S.Y. | | 登録日 | 2020-06-08 | | 公開日 | 2021-04-07 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.198 Å) | | 主引用文献 | Structure-Guided Discovery of a Potent and Selective Cell-Active Inhibitor of SETDB1 Tudor Domain.
Angew.Chem.Int.Ed.Engl., 60, 2021
|
|
