1KY4
 
 | |
1KY5
 
 | | D244E mutant S-Adenosylhomocysteine hydrolase refined with noncrystallographic restraints | | 分子名称: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, 3'-OXO-ADENOSINE, S-adenosylhomocysteine hydrolase | | 著者 | Takata, Y, Takusagawa, F. | | 登録日 | 2002-02-03 | | 公開日 | 2002-09-25 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Catalytic Mechanism of S-adenosylhomocysteine hydrolase. Site-directed
mutagenesis of Asp-130, Lys-185, Asp-189, and Asn-190.
J.Biol.Chem., 277, 2002
|
|
7PY2
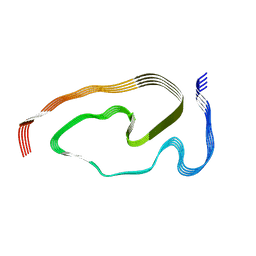 
 | | Structure of pathological TDP-43 filaments from ALS with FTLD | | 分子名称: | TAR DNA-binding protein 43 | | 著者 | Arseni, D, Hasegawa, H, Murzin, A.G, Kametani, F, Arai, M, Yoshida, M, Falcon, B. | | 登録日 | 2021-10-08 | | 公開日 | 2021-12-15 | | 最終更新日 | 2024-07-17 | | 実験手法 | ELECTRON MICROSCOPY (2.59 Å) | | 主引用文献 | Structure of pathological TDP-43 filaments from ALS with FTLD.
Nature, 601, 2022
|
|
1HJR
 
 | |
5JH9
 
 | | Crystal structure of prApe1 | | 分子名称: | CACODYLATE ION, Vacuolar aminopeptidase 1, ZINC ION | | 著者 | Noda, N.N, Adachi, W, Inagaki, F. | | 登録日 | 2016-04-20 | | 公開日 | 2016-06-29 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Structural Basis for Receptor-Mediated Selective Autophagy of Aminopeptidase I Aggregates
Cell Rep, 16, 2016
|
|
5JGF
 
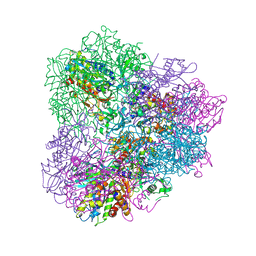 | | Crystal structure of mApe1 | | 分子名称: | Vacuolar aminopeptidase 1, ZINC ION | | 著者 | Noda, N.N, Adachi, W, Inagaki, F. | | 登録日 | 2016-04-20 | | 公開日 | 2016-06-29 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.83 Å) | | 主引用文献 | Structural Basis for Receptor-Mediated Selective Autophagy of Aminopeptidase I Aggregates
Cell Rep, 16, 2016
|
|
5JHC
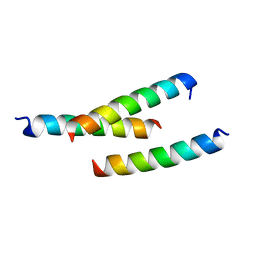 
 | |
5JGE
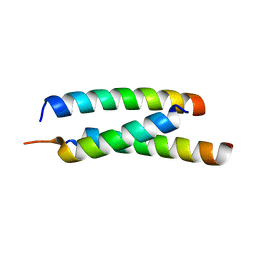 
 | |
1NBI
 
 | |
1KMF
 
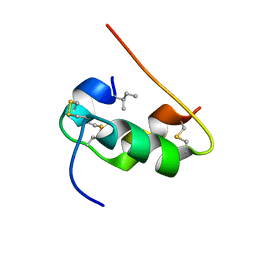 | | NMR STRUCTURE OF HUMAN INSULIN MUTANT ILE-A2-ALLO-ILE, HIS-B10-ASP, PRO-B28-LYS, LYS-B29-PRO, 15 STRUCTURES | | 分子名称: | Insulin | | 著者 | Xu, B, Hua, Q.X, Nakagawa, S.H, Jia, W, Chu, Y.C, Katsoyannis, P.G, Weiss, M.A. | | 登録日 | 2001-12-14 | | 公開日 | 2002-01-09 | | 最終更新日 | 2021-10-27 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Chiral mutagenesis of insulin's hidden receptor-binding surface: structure of an allo-isoleucine(A2) analogue.
J.Mol.Biol., 316, 2002
|
|
1NBH
 
 | |
1KHH
 
 | |
7BRN
 
 | | Crystal structure of Atg40 AIM fused to Atg8 | | 分子名称: | 1,2-ETHANEDIOL, Autophagy-related protein 40,Autophagy-related protein 8, L-EPINEPHRINE | | 著者 | Yamasaki, A, Noda, N.N. | | 登録日 | 2020-03-29 | | 公開日 | 2020-07-08 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.231 Å) | | 主引用文献 | Super-assembly of ER-phagy receptor Atg40 induces local ER remodeling at contacts with forming autophagosomal membranes.
Nat Commun, 11, 2020
|
|
7BRQ
 
 | |
7BRT
 
 | | Crystal structure of Sec62 LIR fused to GABARAP | | 分子名称: | Translocation protein SEC62,Gamma-aminobutyric acid receptor-associated protein | | 著者 | Yamasaki, A, Noda, N.N. | | 登録日 | 2020-03-30 | | 公開日 | 2020-07-08 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.999 Å) | | 主引用文献 | Super-assembly of ER-phagy receptor Atg40 induces local ER remodeling at contacts with forming autophagosomal membranes.
Nat Commun, 11, 2020
|
|
7BRU
 
 | |
1D2C
 
 | | METHYLTRANSFERASE | | 分子名称: | PROTEIN (GLYCINE N-METHYLTRANSFERASE) | | 著者 | Huang, Y, Takusagawa, F. | | 登録日 | 1999-09-23 | | 公開日 | 1999-10-06 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Mechanisms for auto-inhibition and forced product release in glycine N-methyltransferase: crystal structures of wild-type, mutant R175K and S-adenosylhomocysteine-bound R175K enzymes.
J.Mol.Biol., 298, 2000
|
|
5H4S
 
 | |
1P1C
 
 | | Guanidinoacetate Methyltransferase with Gd ion | | 分子名称: | GADOLINIUM ION, Guanidinoacetate N-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | 著者 | Komoto, J, Takusagawa, F. | | 登録日 | 2003-04-12 | | 公開日 | 2003-04-29 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Monoclinic guanidinoacetate methyltransferase and gadolinium ion-binding characteristics.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1D2H
 
 | | CRYSTAL STRUCTURE OF R175K MUTANT GLYCINE N-METHYLTRANSFERASE COMPLEXED WITH S-ADENOSYLHOMOCYSTEINE | | 分子名称: | GLYCINE N-METHYLTRANSFERASE, S-ADENOSYL-L-HOMOCYSTEINE | | 著者 | Huang, Y, Komoto, J, Takusagawa, F, Konishi, K, Takata, Y. | | 登録日 | 1999-10-11 | | 公開日 | 1999-10-25 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Mechanisms for auto-inhibition and forced product release in glycine N-methyltransferase: crystal structures of wild-type, mutant R175K and S-adenosylhomocysteine-bound R175K enzymes.
J.Mol.Biol., 298, 2000
|
|
1D2G
 
 | | CRYSTAL STRUCTURE OF R175K MUTANT GLYCINE N-METHYLTRANSFERASE FROM RAT LIVER | | 分子名称: | GLYCINE N-METHYLTRANSFERASE | | 著者 | Huang, Y, Komoto, J, Takusagawa, F, Konishi, K, Takata, Y. | | 登録日 | 1999-10-08 | | 公開日 | 1999-10-25 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Mechanisms for auto-inhibition and forced product release in glycine N-methyltransferase: crystal structures of wild-type, mutant R175K and S-adenosylhomocysteine-bound R175K enzymes.
J.Mol.Biol., 298, 2000
|
|
1P1B
 
 | | Guanidinoacetate methyltransferase | | 分子名称: | Guanidinoacetate N-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | 著者 | Komoto, J, Takusagawa, F. | | 登録日 | 2003-04-12 | | 公開日 | 2003-04-29 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Monoclinic guanidinoacetate methyltransferase and gadolinium ion-binding characteristics.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
5F9E
 
 | | Structure of Protein Kinase C theta with compound 10: 2,2-dimethyl-7-(2-oxidanylidene-3~{H}-imidazo[4,5-b]pyridin-1-yl)-1-(phenylmethyl)-3~{H}-quinazolin-4-one | | 分子名称: | 2,2-dimethyl-7-(2-oxidanylidene-3~{H}-imidazo[4,5-b]pyridin-1-yl)-1-(phenylmethyl)-3~{H}-quinazolin-4-one, Protein kinase C theta type | | 著者 | Klein, M. | | 登録日 | 2015-12-09 | | 公開日 | 2016-05-11 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Discovery and optimization of 1,7-disubstituted-2,2-dimethyl-2,3-dihydroquinazolin-4(1H)-ones as potent and selective PKC theta inhibitors.
Bioorg.Med.Chem., 24, 2016
|
|
5F20
 
 | |
5F1Z
 
 | |
