2QKU
 
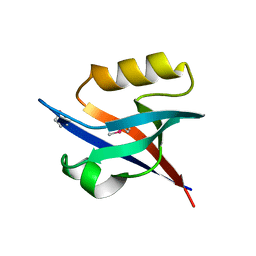 | |
2QKV
 
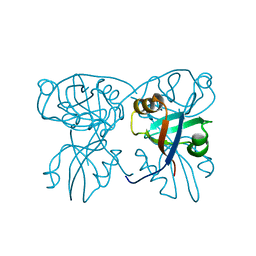 | |
3SOW
 
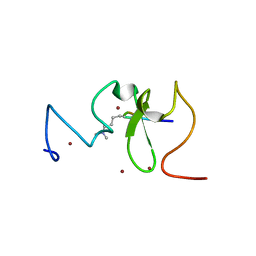 | |
7WH8
 
 | |
3RY2
 
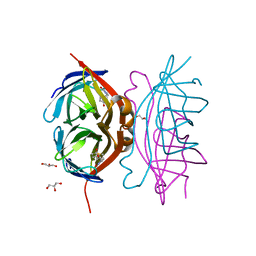 | | Wild-type core streptavidin-biotin complex at atomic resolution | | 分子名称: | BIOTIN, GLYCEROL, Streptavidin | | 著者 | Stenkamp, R.E, Le Trong, I, Stayton, P.S, Lybrand, T.P. | | 登録日 | 2011-05-10 | | 公開日 | 2011-08-24 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (0.95 Å) | | 主引用文献 | Streptavidin and its biotin complex at atomic resolution.
Acta Crystallogr.,Sect.D, 67, 2011
|
|
7YC9
 
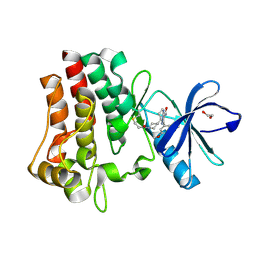 | | Co-crystal structure of BTK kinase domain with inhibitor | | 分子名称: | (7~{S})-2-(4-bromanyl-3,5-dimethoxy-phenyl)-7-(1-propanoylpiperidin-4-yl)-4,5,6,7-tetrahydropyrazolo[1,5-a]pyrimidine-3-carboxamide, 1,2-ETHANEDIOL, Tyrosine-protein kinase BTK | | 著者 | Zhou, X. | | 登録日 | 2022-07-01 | | 公開日 | 2023-05-17 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Discovery of BGB-8035, a Highly Selective Covalent Inhibitor of Bruton's Tyrosine Kinase for B-Cell Malignancies and Autoimmune Diseases.
J.Med.Chem., 66, 2023
|
|
3SOU
 
 | |
3SV1
 
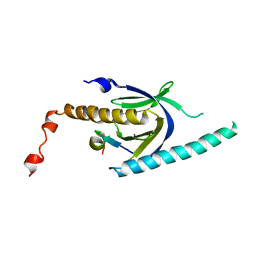 | | Crystal structure of APP peptide bound rat Mint2 PARM | | 分子名称: | Amyloid beta A4 precursor protein-binding family A member 2, Amyloid beta A4 protein | | 著者 | Shen, Y, Long, J, Yan, X, Xie, X. | | 登録日 | 2011-07-12 | | 公開日 | 2012-07-11 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (3.3 Å) | | 主引用文献 | Open-closed motion of Mint2 regulates APP metabolism
J Mol Cell Biol, 5, 2013
|
|
3SOX
 
 | |
3SUZ
 
 | |
6AKY
 
 | | The Crystal structure of Human Chemokine Receptor CCR5 in complex with compound 34 | | 分子名称: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 4,4-difluoro-N-[(1S)-3-{(3-exo)-3-[3-methyl-5-(propan-2-yl)-4H-1,2,4-triazol-4-yl]-8-azabicyclo[3.2.1]octan-8-yl}-1-(thiophen-3-yl)propyl]cyclohexane-1-carboxamide, C-C chemokine receptor type 5,Rubredoxin,C-C chemokine receptor type 5, ... | | 著者 | Zhu, Y, Zhao, Q, Wu, B. | | 登録日 | 2018-09-04 | | 公開日 | 2018-10-24 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Structure-Based Design of 1-Heteroaryl-1,3-propanediamine Derivatives as a Novel Series of CC-Chemokine Receptor 5 Antagonists.
J. Med. Chem., 61, 2018
|
|
3UIT
 
 | | Overall structure of Patj/Pals1/Mals complex | | 分子名称: | ACETATE ION, InaD-like protein, MAGUK p55 subfamily member 5, ... | | 著者 | Zhang, J, Yang, X, Long, J, Shen, Y. | | 登録日 | 2011-11-06 | | 公開日 | 2012-02-22 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (2.05 Å) | | 主引用文献 | Structure of an L27 domain heterotrimer from cell polarity complex Patj/Pals1/Mals2 reveals mutually independent L27 domain assembly mode
J.Biol.Chem., 287, 2012
|
|
3UIM
 
 | |
6KZ5
 
 | | Crystal Structure Analysis of the Csn-B-bounded NUR77 Ligand binding Domain | | 分子名称: | Nuclear receptor subfamily 4 group A member 1, ethyl 2-[2-octanoyl-3,5-bis(oxidanyl)phenyl]ethanoate | | 著者 | Hong, W, Chen, H, Wu, Q, Lin, T. | | 登録日 | 2019-09-23 | | 公開日 | 2020-10-14 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (4.45 Å) | | 主引用文献 | Blocking PPAR gamma interaction facilitates Nur77 interdiction of fatty acid uptake and suppresses breast cancer progression.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
2GZM
 
 | | Crystal Structure of the Glutamate Racemase from Bacillus anthracis | | 分子名称: | D-GLUTAMIC ACID, Glutamate racemase | | 著者 | May, M, Santarsiero, B.D, Johnson, M.E, Mesecar, A.D. | | 登録日 | 2006-05-11 | | 公開日 | 2007-05-29 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (1.99 Å) | | 主引用文献 | Structural and functional analysis of two glutamate racemase isozymes from Bacillus anthracis and implications for inhibitor design.
J.Mol.Biol., 371, 2007
|
|
4M6T
 
 | | Structure of human Paf1 and Leo1 complex | | 分子名称: | RNA polymerase II-associated factor 1 homolog, Linker, RNA polymerase-associated protein LEO1, ... | | 著者 | Shen, Y, Qin, X. | | 登録日 | 2013-08-11 | | 公開日 | 2013-10-02 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (2.498 Å) | | 主引用文献 | Structural insights into Paf1 complex assembly and histone binding
Nucleic Acids Res., 41, 2013
|
|
4L8D
 
 | |
4L8B
 
 | |
7XKG
 
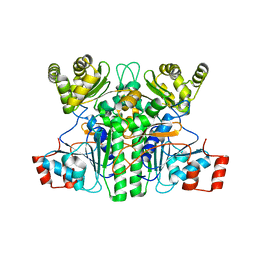 | | Crystal structure of an intramolecular mesacyl-CoA transferase from the 3-hydroxypropionic acid cycle of Roseiflexus castenholzii | | 分子名称: | Acyl-CoA transferase/carnitine dehydratase-like protein | | 著者 | Min, Z.Z, Fan, C.P, Wu, W.P, Xin, Y.Y, Liu, M.H, Zhang, X, Wang, Z.G, Xu, X.L. | | 登録日 | 2022-04-19 | | 公開日 | 2022-06-15 | | 最終更新日 | 2024-05-29 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Crystal Structure of an Intramolecular Mesaconyl-Coenzyme A Transferase From the 3-Hydroxypropionic Acid Cycle of Roseiflexus castenholzii .
Front Microbiol, 13, 2022
|
|
4NWU
 
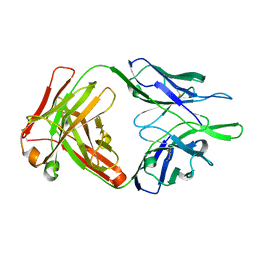 | |
7Y78
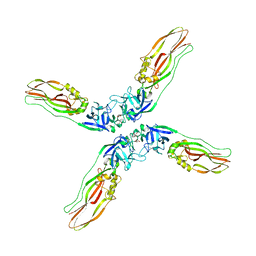 
 | | Crystal structure of Cry78Aa | | 分子名称: | 1,2-ETHANEDIOL, AMMONIUM ION, Toxin | | 著者 | Cao, B.B, Nie, Y.F, Wang, N.C, Guan, Z.Y, Zhang, D.L, Zhang, J. | | 登録日 | 2022-06-21 | | 公開日 | 2022-08-31 | | 最終更新日 | 2024-05-29 | | 実験手法 | X-RAY DIFFRACTION (2.9 Å) | | 主引用文献 | The crystal structure of Cry78Aa from Bacillus thuringiensis provides insights into its insecticidal activity.
Commun Biol, 5, 2022
|
|
7Y79
 
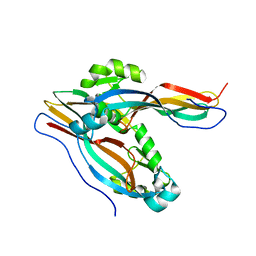 | | Crystal structure of Cry78Aa | | 分子名称: | Toxin | | 著者 | Cao, B.B, Nie, Y.F, Wang, N.C, Guan, Z.Y, Zhang, D.L, Zhang, J. | | 登録日 | 2022-06-21 | | 公開日 | 2022-08-31 | | 最終更新日 | 2024-05-29 | | 実験手法 | X-RAY DIFFRACTION (2.32 Å) | | 主引用文献 | The crystal structure of Cry78Aa from Bacillus thuringiensis provides insights into its insecticidal activity.
Commun Biol, 5, 2022
|
|
4NWT
 
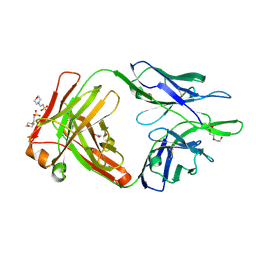 | | Crystal structure of the anti-human NGF Fab APE1531 | | 分子名称: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ACETATE ION, APE1531 Ab Fab heavy chain, ... | | 著者 | Verdino, P, Stanfield, R.L, Wilson, I.A. | | 登録日 | 2013-12-06 | | 公開日 | 2014-10-22 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (1.75 Å) | | 主引用文献 | Nucleotide insertions and deletions complement point mutations to massively expand the diversity created by somatic hypermutation of antibodies.
J.Biol.Chem., 289, 2014
|
|
4OE1
 
 | | Crystal structure of the pentatricopeptide repeat protein PPR10 (C256S/C430S/C449S) in complex with an 18-nt PSAJ rna element | | 分子名称: | Chloroplast pentatricopeptide repeat protein 10, PHOSPHATE ION, psaJ RNA | | 著者 | Li, Q, Yan, C, Wu, J, Yin, P, Yan, N. | | 登録日 | 2014-01-11 | | 公開日 | 2014-09-24 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Examination of the dimerization states of the single-stranded RNA recognition protein pentatricopeptide repeat 10 (PPR10).
J.Biol.Chem., 289, 2014
|
|
1OIB
 
 | |
