8Y85
 
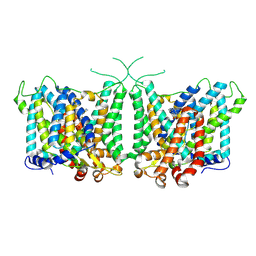 | | Human AE3 with NaHCO3- and DIDS | | 分子名称: | 2,2'-ethane-1,2-diylbis{5-[(sulfanylmethyl)amino]benzenesulfonic acid}, Anion exchange protein 3, BICARBONATE ION | | 著者 | Jian, L, Zhang, Q, Yao, D, Wang, Q, Xia, Y, Qin, A, Cao, Y. | | 登録日 | 2024-02-05 | | 公開日 | 2024-07-31 | | 最終更新日 | 2025-06-25 | | 実験手法 | ELECTRON MICROSCOPY (2.73 Å) | | 主引用文献 | The structural insight into the functional modulation of human anion exchanger 3.
Nat Commun, 15, 2024
|
|
7D6X
 
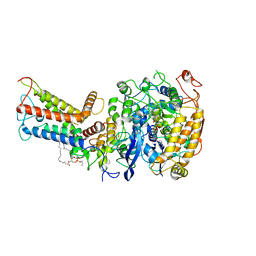 | | Mycobacterium smegmatis Sdh1 complex in the apo form | | 分子名称: | (1S)-2-{[(2-AMINOETHOXY)(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL STEARATE, FE2/S2 (INORGANIC) CLUSTER, FE3-S4 CLUSTER, ... | | 著者 | Zhou, X, Gao, Y, Wang, Q, Gong, H, Rao, Z. | | 登録日 | 2020-10-02 | | 公開日 | 2021-04-07 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (2.88 Å) | | 主引用文献 | Architecture of the mycobacterial succinate dehydrogenase with a membrane-embedded Rieske FeS cluster.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7D6V
 
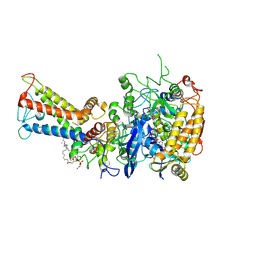 | | Mycobacterium smegmatis Sdh1 in complex with UQ1 | | 分子名称: | (1S)-2-{[(2-AMINOETHOXY)(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL STEARATE, FE2/S2 (INORGANIC) CLUSTER, FE3-S4 CLUSTER, ... | | 著者 | Zhou, X, Gao, Y, Wang, Q, Gong, H, Rao, Z. | | 登録日 | 2020-10-02 | | 公開日 | 2021-04-07 | | 最終更新日 | 2025-07-02 | | 実験手法 | ELECTRON MICROSCOPY (2.53 Å) | | 主引用文献 | Architecture of the mycobacterial succinate dehydrogenase with a membrane-embedded Rieske FeS cluster.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
8ZLE
 
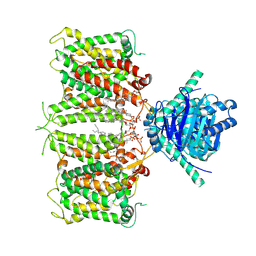 | | hAE3NTD2TMD with PT5,CLR, and Y01 | | 分子名称: | CHOLESTEROL, CHOLESTEROL HEMISUCCINATE, [(2R)-1-octadecanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(oxidanyl)-4,5-diphosphonooxy-cyclohexyl]oxy-phospho ryl]oxy-propan-2-yl] (8Z)-icosa-5,8,11,14-tetraenoate, ... | | 著者 | Jian, L, Zhang, Q, Yao, D, Wang, Q, Xia, Y, Qin, A, Cao, Y. | | 登録日 | 2024-05-19 | | 公開日 | 2024-07-31 | | 最終更新日 | 2025-07-02 | | 実験手法 | ELECTRON MICROSCOPY (3.35 Å) | | 主引用文献 | The structural insight into the functional modulation of human anion exchanger 3.
Nat Commun, 15, 2024
|
|
8K0M
 
 | | Human collagen prolyl processing enzyme complex, P3H1/CRTAP/PPIB heterotrimer, bound to 2-oxoglutarate | | 分子名称: | 2-OXOGLUTARIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, Cartilage-associated protein, ... | | 著者 | Li, W, Peng, J, Yao, D, Rao, B, Xia, Y, Wang, Q, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Zhao, J, Cao, Y. | | 登録日 | 2023-07-09 | | 公開日 | 2024-09-18 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (3.17 Å) | | 主引用文献 | The structural basis for the collagen processing by human P3H1/CRTAP/PPIB ternary complex.
Nat Commun, 15, 2024
|
|
8K0I
 
 | | Human collagen prolyl processing enzyme complex, P3H1/CRTAP/PPIB heterotrimer, in its dual-ternary state | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Cartilage-associated protein, Peptidyl-prolyl cis-trans isomerase B, ... | | 著者 | Li, W, Peng, J, Yao, D, Rao, B, Xia, Y, Wang, Q, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Zhao, J, Cao, Y. | | 登録日 | 2023-07-09 | | 公開日 | 2024-09-18 | | 最終更新日 | 2024-10-30 | | 実験手法 | ELECTRON MICROSCOPY (3.62 Å) | | 主引用文献 | The structural basis for the collagen processing by human P3H1/CRTAP/PPIB ternary complex.
Nat Commun, 15, 2024
|
|
8K0F
 
 | | Human collagen prolyl processing enzyme complex, P3H1/CRTAP/PPIB heterotrimer, in its apo state | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Cartilage-associated protein, FE (III) ION, ... | | 著者 | Li, W, Peng, J, Yao, D, Rao, B, Xia, Y, Wang, Q, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Zhao, J, Cao, Y. | | 登録日 | 2023-07-08 | | 公開日 | 2024-09-18 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (3.37 Å) | | 主引用文献 | The structural basis for the collagen processing by human P3H1/CRTAP/PPIB ternary complex.
Nat Commun, 15, 2024
|
|
8K17
 
 | | Human collagen prolyl processing enzyme complex, P3H1/CRTAP/PPIB heterotrimer, bound to collagen alpha-1(I) chain | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Cartilage-associated protein, FE (III) ION, ... | | 著者 | Li, W, Peng, J, Yao, D, Rao, B, Xia, Y, Wang, Q, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Zhao, J, Cao, Y. | | 登録日 | 2023-07-10 | | 公開日 | 2024-09-18 | | 最終更新日 | 2024-11-13 | | 実験手法 | ELECTRON MICROSCOPY (3.18 Å) | | 主引用文献 | The structural basis for the collagen processing by human P3H1/CRTAP/PPIB ternary complex.
Nat Commun, 15, 2024
|
|
8K0E
 
 | | Human collagen prolyl processing enzyme complex, P3H1/CRTAP heterodimer | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Cartilage-associated protein, FE (III) ION, ... | | 著者 | Li, W, Peng, J, Yao, D, Rao, B, Xia, Y, Wang, Q, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Zhao, J, Cao, Y. | | 登録日 | 2023-07-08 | | 公開日 | 2024-09-18 | | 最終更新日 | 2024-11-13 | | 実験手法 | ELECTRON MICROSCOPY (3.65 Å) | | 主引用文献 | The structural basis for the collagen processing by human P3H1/CRTAP/PPIB ternary complex.
Nat Commun, 15, 2024
|
|
8KC9
 | | Human collagen prolyl processing enzyme complex, P3H1/CRTAP/PPIB heterotrimer, bound to cyclosporin A | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Cartilage-associated protein, DSN-MLE-MLE-MVA-BMT-ABA-SAR-MLE-VAL-MLE-ALA, ... | | 著者 | Li, W, Peng, J, Yao, D, Rao, B, Xia, Y, Wang, Q, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Zhao, J, Cao, Y. | | 登録日 | 2023-08-06 | | 公開日 | 2024-09-18 | | 実験手法 | ELECTRON MICROSCOPY (3.75 Å) | | 主引用文献 | The structural basis for the collagen processing by human P3H1/CRTAP/PPIB ternary complex.
Nat Commun, 15, 2024
|
|
7U4K
 
 | | Crystal structure of human GPX4-U46C-R152H in complex with ML162 | | 分子名称: | 1,2-ETHANEDIOL, 2-chloro-N-(3-chloro-4-methoxyphenyl)-N-[(1R)-2-oxo-2-[(2-phenylethyl)amino]-1-(thiophen-2-yl)ethyl]acetamide, Phospholipid hydroperoxide glutathione peroxidase | | 著者 | Forouhar, F, Liu, H, Lin, A.J, Wang, Q, Xia, X, Soni, R.K, Stockwell, B.R. | | 登録日 | 2022-02-28 | | 公開日 | 2022-12-07 | | 最終更新日 | 2024-10-09 | | 実験手法 | X-RAY DIFFRACTION (1.69 Å) | | 主引用文献 | Small-molecule allosteric inhibitors of GPX4.
Cell Chem Biol, 29, 2022
|
|
7U4L
 
 | | Crystal structure of human GPX4-U46C in complex with MAC-5576 | | 分子名称: | Phospholipid hydroperoxide glutathione peroxidase, thiophene-2-carbaldehyde | | 著者 | Forouhar, F, Liu, H, Lin, A.J, Wang, Q, Polychronidou, V, Soni, R.K, Xia, X, Stockwell, B.R. | | 登録日 | 2022-02-28 | | 公開日 | 2022-12-07 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (2.25 Å) | | 主引用文献 | Small-molecule allosteric inhibitors of GPX4.
Cell Chem Biol, 29, 2022
|
|
7U4N
 
 | | Crystal structure of human GPX4-U46C in complex with RSL3 | | 分子名称: | Phospholipid hydroperoxide glutathione peroxidase, methyl (1S,3R)-2-(chloroacetyl)-1-[4-(methoxycarbonyl)phenyl]-2,3,4,9-tetrahydro-1H-pyrido[3,4-b]indole-3-carboxylate | | 著者 | Forouhar, F, Liu, H, Lin, A.J, Wang, Q, Xia, X, Soni, R.K, Stockwell, B.R. | | 登録日 | 2022-02-28 | | 公開日 | 2022-12-07 | | 最終更新日 | 2024-10-09 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Small-molecule allosteric inhibitors of GPX4.
Cell Chem Biol, 29, 2022
|
|
7U4M
 
 | | Crystal structure of human GPX4-U46C in complex with LOC1886 | | 分子名称: | 1,2-ETHANEDIOL, 4-methoxy-1H-indole-2-carbaldehyde, Phospholipid hydroperoxide glutathione peroxidase | | 著者 | Forouhar, F, Liu, H, Lin, A.J, Wang, Q, Polychronidou, V, Soni, R.K, Xia, X, Stockwell, B.R. | | 登録日 | 2022-02-28 | | 公開日 | 2022-12-07 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (1.93 Å) | | 主引用文献 | Small-molecule allosteric inhibitors of GPX4.
Cell Chem Biol, 29, 2022
|
|
7U4J
 
 | | Crystal structure of human GPX4-U46C-R152H in complex with TMT10 | | 分子名称: | Phospholipid hydroperoxide glutathione peroxidase, THIOCYANATE ION, ~{N}-(3-chloranyl-4-methoxy-phenyl)ethanamide | | 著者 | Forouhar, F, Liu, H, Lin, A.J, Wang, Q, Polychronidou, V, Soni, R.K, Xia, X, Stockwell, B.R. | | 登録日 | 2022-02-28 | | 公開日 | 2022-12-07 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (1.81 Å) | | 主引用文献 | Small-molecule allosteric inhibitors of GPX4.
Cell Chem Biol, 29, 2022
|
|
7U4I
 
 | | Crystal structure of human GPX4-U46C-R152H in complex with CDS9 | | 分子名称: | 2-bromo-N-[(thiophen-2-yl)methyl]acetamide, Phospholipid hydroperoxide glutathione peroxidase, THIOCYANATE ION | | 著者 | Forouhar, F, Liu, H, Lin, A.J, Wang, Q, Polychronidou, V, Soni, R.K, Xia, X, Stockwell, B.R. | | 登録日 | 2022-02-28 | | 公開日 | 2022-12-07 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (1.97 Å) | | 主引用文献 | Small-molecule allosteric inhibitors of GPX4.
Cell Chem Biol, 29, 2022
|
|
8ZGH
 
 | | Human lysine O-link glycosylation complex, LH3/ColGalT1 in its apo state | | 分子名称: | 2-OXOGLUTARIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | 著者 | Peng, J, Li, W, Yao, D, Xia, Y, Wang, Q, Cai, Y, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Cao, Y. | | 登録日 | 2024-05-09 | | 公開日 | 2025-03-19 | | 最終更新日 | 2025-03-26 | | 実験手法 | ELECTRON MICROSCOPY (3.93 Å) | | 主引用文献 | The structural basis for the human procollagen lysine hydroxylation and dual-glycosylation.
Nat Commun, 16, 2025
|
|
8ZGE
 
 | | Human lysine O-link glycosylation complex, LH3/ColGalT1 tetramer with bound UDP-galactose | | 分子名称: | 2-OXOGLUTARIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | 著者 | Peng, J, Li, W, Yao, D, Xia, Y, Wang, Q, Cai, Y, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Cao, Y. | | 登録日 | 2024-05-09 | | 公開日 | 2025-03-19 | | 最終更新日 | 2025-03-26 | | 実験手法 | ELECTRON MICROSCOPY (3.4 Å) | | 主引用文献 | The structural basis for the human procollagen lysine hydroxylation and dual-glycosylation.
Nat Commun, 16, 2025
|
|
8ZGG
 
 | | Human lysine O-link glycosylation complex, LH3/ColGalT1 with bound UDP-glucose | | 分子名称: | 2-OXOGLUTARIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | 著者 | Peng, J, Li, W, Yao, D, Xia, Y, Wang, Q, Cai, Y, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Cao, Y. | | 登録日 | 2024-05-09 | | 公開日 | 2025-03-19 | | 最終更新日 | 2025-03-26 | | 実験手法 | ELECTRON MICROSCOPY (3.75 Å) | | 主引用文献 | The structural basis for the human procollagen lysine hydroxylation and dual-glycosylation.
Nat Commun, 16, 2025
|
|
8ZGC
 
 | | Human lysine O-link glycosylation complex, LH3/ColGalT1 with bound UDP-galactose | | 分子名称: | 2-OXOGLUTARIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | 著者 | Peng, J, Li, W, Yao, D, Xia, Y, Wang, Q, Cai, Y, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Cao, Y. | | 登録日 | 2024-05-09 | | 公開日 | 2025-03-19 | | 最終更新日 | 2025-03-26 | | 実験手法 | ELECTRON MICROSCOPY (3.58 Å) | | 主引用文献 | The structural basis for the human procollagen lysine hydroxylation and dual-glycosylation.
Nat Commun, 16, 2025
|
|
7BX8
 
 | | Mycobacterium smegmatis arabinosyltransferase complex EmbB2-AcpM2 in symmetric "resting state" | | 分子名称: | Integral membrane indolylacetylinositol arabinosyltransferase EmbB, Meromycolate extension acyl carrier protein | | 著者 | Gao, R.G, Zhang, L, Wang, Q, Rao, Z.H. | | 登録日 | 2020-04-17 | | 公開日 | 2020-05-27 | | 最終更新日 | 2024-03-27 | | 実験手法 | ELECTRON MICROSCOPY (3.6 Å) | | 主引用文献 | Cryo-EM snapshots of mycobacterial arabinosyltransferase complex EmbB2-AcpM2.
Protein Cell, 11, 2020
|
|
7CP9
 | | Cryo-EM structure of human mitochondrial translocase TOM complex at 3.0 angstrom. | | 分子名称: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, Mitochondrial import receptor subunit TOM22 homolog, Mitochondrial import receptor subunit TOM40 homolog, ... | | 著者 | Guan, Z, Yan, L, Wang, Q, Yan, C, Yin, P. | | 登録日 | 2020-08-06 | | 公開日 | 2021-04-28 | | 最終更新日 | 2024-03-27 | | 実験手法 | ELECTRON MICROSCOPY (3 Å) | | 主引用文献 | Structural insights into assembly of human mitochondrial translocase TOM complex.
Cell Discov, 7, 2021
|
|
7CZB
 
 | | The cryo-EM structure of the ERAD retrotranslocation channel formed by human Derlin-1 | | 分子名称: | Derlin-1 | | 著者 | Rao, B, Li, S, Yao, D, Wang, Q, Xia, Y, Jia, Y, Shen, Y, Cao, Y. | | 登録日 | 2020-09-07 | | 公開日 | 2021-03-17 | | 最終更新日 | 2024-03-27 | | 実験手法 | ELECTRON MICROSCOPY (3.8 Å) | | 主引用文献 | The cryo-EM structure of an ERAD protein channel formed by tetrameric human Derlin-1.
Sci Adv, 7, 2021
|
|
7BWR
 
 | | Mycobacterium smegmatis arabinosyltransferase complex EmbB2-AcpM2 in substrate DPA bound asymmetric "active state" | | 分子名称: | CALCIUM ION, Integral membrane indolylacetylinositol arabinosyltransferase EmbB, Meromycolate extension acyl carrier protein, ... | | 著者 | Gao, R.G, Zhang, L, Wang, Q, Rao, Z.H. | | 登録日 | 2020-04-15 | | 公開日 | 2020-05-27 | | 最終更新日 | 2024-03-27 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | Cryo-EM snapshots of mycobacterial arabinosyltransferase complex EmbB2-AcpM2.
Protein Cell, 11, 2020
|
|
7CGP
 
 | | Cryo-EM structure of the human mitochondrial translocase TIM22 complex at 3.7 angstrom. | | 分子名称: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, Acylglycerol kinase, mitochondrial, ... | | 著者 | Qi, L, Wang, Q, Guan, Z, Yan, C, Yin, P. | | 登録日 | 2020-07-01 | | 公開日 | 2020-10-14 | | 最終更新日 | 2025-06-18 | | 実験手法 | ELECTRON MICROSCOPY (3.7 Å) | | 主引用文献 | Cryo-EM structure of the human mitochondrial translocase TIM22 complex.
Cell Res., 31, 2021
|
|
