1S7C
 
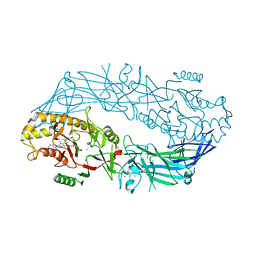
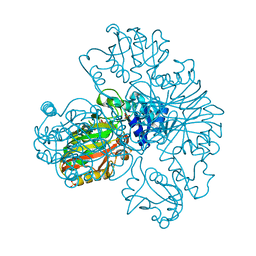 | | Crystal structure of MES buffer bound form of glyceraldehyde 3-phosphate dehydrogenase from Escherichia coli | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Glyceraldehyde 3-phosphate dehydrogenase A, SULFATE ION | | Authors: | Shin, D.H, Thor, J, Yokota, H, Kim, R, Kim, S.H, Berkeley Structural Genomics Center (BSGC) | | Deposit date: | 2004-01-29 | | Release date: | 2004-08-10 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Crystal structure of MES buffer bound form of glyceraldehyde 3-phosphate dehydrogenase from Escherichia coli
To be Published
|
|
4X8G
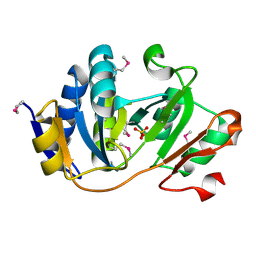
 | | Crystal structure of human peptidylarginine deiminase type4 (PAD4) in complex with GSK199 | | Descriptor: | CALCIUM ION, Protein-arginine deiminase type-4, [(3R)-3-aminopiperidin-1-yl][2-(1-ethyl-1H-pyrrolo[2,3-b]pyridin-2-yl)-7-methoxy-1-methyl-1H-benzimidazol-5-yl]methanone | | Authors: | Lewis, H.D, Bax, B.D, Chung, C.-W, Polyakova, O, Thorpe, J. | | Deposit date: | 2014-12-10 | | Release date: | 2015-01-28 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.29 Å) | | Cite: | Inhibition of PAD4 activity is sufficient to disrupt mouse and human NET formation.
Nat.Chem.Biol., 11, 2015
|
|
4X8C
 
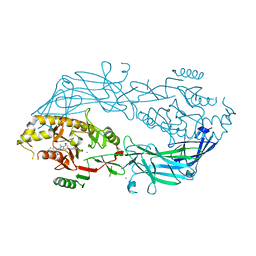 | | Crystal structure of human peptidylarginine deiminase type4 (PAD4) in complex with GSK147 | | Descriptor: | CALCIUM ION, Protein-arginine deiminase type-4, [(3S,4R)-3-amino-4-hydroxypiperidin-1-yl]{2-[1-(cyclopropylmethyl)-1H-pyrrolo[2,3-b]pyridin-2-yl]-7-methoxy-1-methyl-1H-benzimidazol-5-yl}methanone | | Authors: | Lewis, H.D, Bax, B.D, Chung, C.-W, Polyakova, O, Thorpe, J. | | Deposit date: | 2014-12-10 | | Release date: | 2015-01-28 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Inhibition of PAD4 activity is sufficient to disrupt mouse and human NET formation.
Nat.Chem.Biol., 11, 2015
|
|
1FPW
 
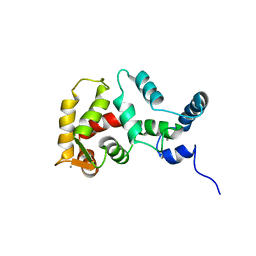 | | STRUCTURE OF YEAST FREQUENIN | | Descriptor: | CALCIUM ION, CALCIUM-BINDING PROTEIN NCS-1 | | Authors: | Ames, J.B, Hendricks, K.B, Strahl, T, Huttner, I.G, Thorner, J. | | Deposit date: | 2000-08-31 | | Release date: | 2000-10-18 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure and calcium-binding properties of Frq1, a novel calcium sensor in the yeast Saccharomyces cerevisiae.
Biochemistry, 39, 2000
|
|
1K7J
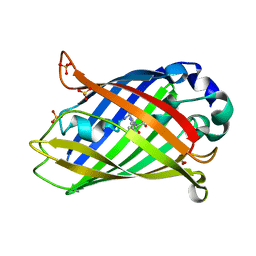
 | | Structural Genomics, protein TF1 | | Descriptor: | Protein yciO, SULFATE ION | | Authors: | Zhang, R, Dementieva, I, Thorn, J, Donnelly, M, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2001-10-19 | | Release date: | 2002-08-14 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural Genomics, protein TF1
To be Published
|
|
2KX6
 
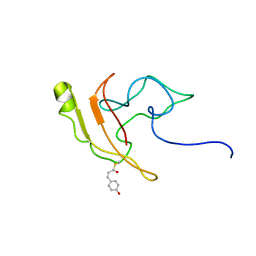 | | Signaling state of Photoactive Yellow Protein | | Descriptor: | 4'-HYDROXYCINNAMIC ACID, Photoactive yellow protein | | Authors: | Ramachandran, P.L, Lovett, J.E, Carl, P.J, Cammarata, M, Lee, J.H, Yang, J.O, Ihee, H, Timmel, C.R, van Thor, J. | | Deposit date: | 2010-04-27 | | Release date: | 2011-06-15 | | Last modified: | 2012-07-18 | | Method: | SOLUTION NMR, SOLUTION SCATTERING | | Cite: | The short-lived signaling state of the photoactive yellow protein photoreceptor revealed by combined structural probes.
J.Am.Chem.Soc., 133, 2011
|
|
8A83
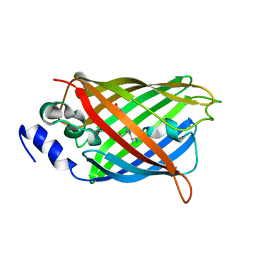
 | |
8A6O
 
 | |
8A6R
 
 | |
8A6S
 
 | |
8A6G
 
 | |
8A6N
 
 | |
8A6P
 
 | |
8A6Q
 
 | |
8A7V
 | |
8AM4
 
 | | Cl-rsEGFP2 Long Wavelength Structure | | Descriptor: | Green fluorescent protein | | Authors: | Orr, C.M, Fadini, A, van Thor, J. | | Deposit date: | 2022-08-02 | | Release date: | 2023-08-02 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Serial Femtosecond Crystallography Reveals that Photoactivation in a Fluorescent Protein Proceeds via the Hula Twist Mechanism.
J.Am.Chem.Soc., 2023
|
|
6T2K
 
 | | Furano[2,3-d]prymidine amides as Notum inhibitors | | Descriptor: | 1,2-ETHANEDIOL, 2-(6-chloranyl-7-cyclopropyl-thieno[3,2-d]pyrimidin-4-yl)sulfanylethanoic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Zhao, Y, Jones, E.Y. | | Deposit date: | 2019-10-08 | | Release date: | 2020-01-01 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Scaffold-hopping identifies furano[2,3-d]pyrimidine amides as potent Notum inhibitors.
Bioorg.Med.Chem.Lett., 30, 2020
|
|
7PK3
 
 | | Notum_ARUK3001185 | | Descriptor: | 1-[2,4-bis(chloranyl)-3-(trifluoromethyl)phenyl]-1,2,3-triazole, 2-acetamido-2-deoxy-beta-D-glucopyranose, DIMETHYL SULFOXIDE, ... | | Authors: | Vecchia, L, Hillier, J, Zhao, Y, Fish, P, Jones, E.Y. | | Deposit date: | 2021-08-25 | | Release date: | 2022-09-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.41 Å) | | Cite: | Design of a Potent, Selective, and Brain-Penetrant Inhibitor of Wnt-Deactivating Enzyme Notum by Optimization of a Crystallographic Fragment Hit.
J.Med.Chem., 65, 2022
|
|
7A16
 
 | |
7A15
 
 | |
7A14
 
 | |
7A12
 
 | |
7A13
 
 | |
2AD1
 
 | | Human Sulfotransferase SULT1C2 | | Descriptor: | Sulfotransferase 1C2 | | Authors: | Dong, A, Dombrovski, L, Loppnau, P, Edwards, A.M, Arrowsmith, C.H, Weigelt, J, Sundstrom, M, Bochkarev, A, Plotnikov, A.N, Structural Genomics Consortium (SGC) | | Deposit date: | 2005-07-19 | | Release date: | 2005-08-02 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and chemical profiling of the human cytosolic sulfotransferases.
Plos Biol., 5, 2007
|
|
6ZVL
 
 | | ARUK3000263 complex with Notum | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 5-[4-chloranyl-3-(trifluoromethyl)phenyl]-3~{H}-1,3,4-oxadiazol-2-one, ... | | Authors: | Zhao, Y, Ruza, R. | | Deposit date: | 2020-07-24 | | Release date: | 2020-11-11 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | 5-Phenyl-1,3,4-oxadiazol-2(3 H )-ones Are Potent Inhibitors of Notum Carboxylesterase Activity Identified by the Optimization of a Crystallographic Fragment Screening Hit.
J.Med.Chem., 63, 2020
|
|
