6F1P
 
 | |
6F9Z
 
 | |
2F2W
 
 | | alpha-spectrin SH3 domain R21A mutant | | Descriptor: | SULFATE ION, Spectrin alpha chain, brain | | Authors: | Camara-Artigas, A, Conejero-Lara, F, Casares, S, Lopez-Mayorga, O, Vega, C. | | Deposit date: | 2005-11-18 | | Release date: | 2006-10-31 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Cooperative propagation of local stability changes from low-stability and high-stability regions in a SH3 domain
Proteins, 67, 2007
|
|
2F2X
 
 | | alpha-spectrin SH3 domain R21G mutant | | Descriptor: | SULFATE ION, Spectrin alpha chain, brain | | Authors: | Camara-Artigas, A, Conejero-Lara, F, Casares, S, Lopez-Mayorga, O, Vega, C. | | Deposit date: | 2005-11-18 | | Release date: | 2006-10-31 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Cooperative propagation of local stability changes from low-stability and high-stability regions in a SH3 domain
Proteins, 67, 2007
|
|
6F1O
 
 | | Orthorhombic Lysozyme crystallized at 298 K and pH 4.5 | | Descriptor: | CHLORIDE ION, Lysozyme C, PHOSPHATE ION | | Authors: | Camara-Artigas, A. | | Deposit date: | 2017-11-22 | | Release date: | 2018-05-09 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (0.96 Å) | | Cite: | Orthorhombic lysozyme crystallization at acidic pH values driven by phosphate binding.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
6F1M
 
 | |
6F1R
 
 | |
6FA0
 
 | |
6F9X
 
 | |
4OMO
 
 | | Crystal structure of the c-Src tyrosine kinase SH3 domain mutant Q128E | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, NICKEL (II) ION, Proto-oncogene tyrosine-protein kinase Src | | Authors: | Camara-Artigas, A, Bacarizo, J. | | Deposit date: | 2014-01-27 | | Release date: | 2014-12-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | Electrostatic Effects in the Folding of the SH3 Domain of the c-Src Tyrosine Kinase: pH-Dependence in 3D-Domain Swapping and Amyloid Formation.
Plos One, 9, 2014
|
|
4OML
 
 | |
4OMN
 
 | |
4OMP
 
 | |
4QT7
 
 | |
4R61
 
 | |
4REX
 
 | |
3HG6
 
 | | Crystal Structure of the Recombinant Onconase from Rana pipiens | | Descriptor: | GLYCEROL, Onconase, SULFATE ION | | Authors: | Camara-Artigas, A, Gavira, J.A, Casares-Atienza, S, Weininger, U, Balbach, J, Garcia-Mira, M.M. | | Deposit date: | 2009-05-13 | | Release date: | 2010-05-19 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Three-state thermal unfolding of onconase.
Biophys.Chem., 159, 2011
|
|
3M0R
 
 | |
4HVV
 
 | |
4HVW
 
 | |
4HVU
 | |
4JZ4
 
 | | Crystal structure of chicken c-Src-SH3 domain: monomeric form | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, NICKEL (II) ION, Proto-oncogene tyrosine-protein kinase Src | | Authors: | Camara-Artigas, A. | | Deposit date: | 2013-04-02 | | Release date: | 2014-04-23 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Electrostatic Effects in the Folding of the SH3 Domain of the c-Src Tyrosine Kinase: pH-Dependence in 3D-Domain Swapping and Amyloid Formation.
Plos One, 9, 2014
|
|
4JZ3
 
 | | Crystal structure of the chicken c-Src-SH3 domain intertwined dimer | | Descriptor: | DI(HYDROXYETHYL)ETHER, Proto-oncogene tyrosine-protein kinase Src, TRIETHYLENE GLYCOL | | Authors: | Camara-Artigas, A. | | Deposit date: | 2013-04-02 | | Release date: | 2014-04-23 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.852 Å) | | Cite: | Electrostatic Effects in the Folding of the SH3 Domain of the c-Src Tyrosine Kinase: pH-Dependence in 3D-Domain Swapping and Amyloid Formation.
Plos One, 9, 2014
|
|
2OLP
 
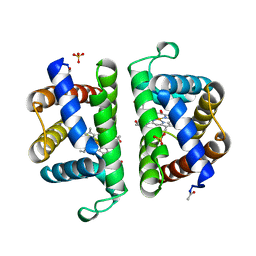 | | Structure and ligand selection of hemoglobin II from Lucina pectinata | | Descriptor: | Hemoglobin II, OXYGEN MOLECULE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Gavira, J.A, Camara-Artigas, A, de Jesus, W, Lopez-Garriga, J, Garcia-Ruiz, J.M. | | Deposit date: | 2007-01-19 | | Release date: | 2007-12-18 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.932 Å) | | Cite: | Structure and Ligand Selection of Hemoglobin II from Lucina pectinata
J.Biol.Chem., 283, 2008
|
|
2CDT
 | | alpha-SPECTRIN SH3 DOMAIN A56S MUTANT | | Descriptor: | SPECTRIN ALPHA CHAIN | | Authors: | Casares, S, Camara-Artigas, A, Vega, M.C, Lopez-Mayorga, O, Conejero-Lara, F. | | Deposit date: | 2006-01-27 | | Release date: | 2007-02-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Cooperative Propagation of Local Stability Changes from Low-Stability and High-Stability Regions in a SH3 Domain.
Proteins: Struct., Funct., Bioinf., 67, 2007
|
|
