7EX0
 
 | |
7EX6
 
 | |
7EX2
 
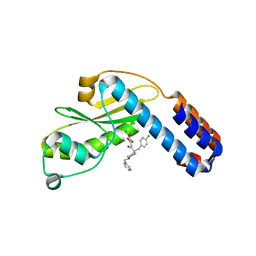 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with L-742,001 (methyl ester form) | | Descriptor: | MANGANESE (II) ION, Replicase, methyl (Z)-4-[4-[(4-chlorophenyl)methyl]-1-(phenylmethyl)piperidin-4-yl]-2-oxidanyl-4-oxidanylidene-but-2-enoate | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
7EX4
 
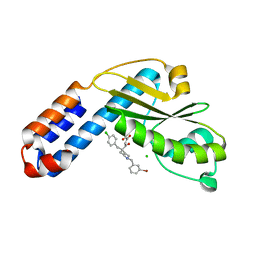 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with inhibitor 5 | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, Replicase, ... | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
7EX7
 
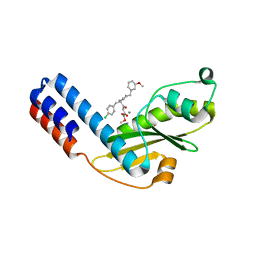 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with inhibitor 16 | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, Replicase, ... | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
7EX3
 
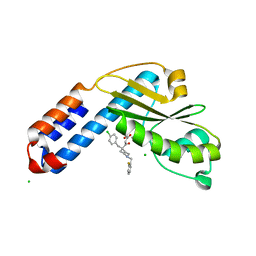 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with inhibitor 3 | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, Replicase, ... | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
7EX8
 
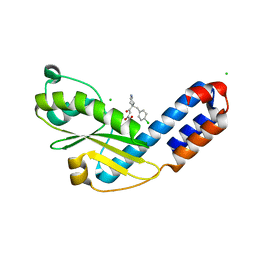 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with inhibitor 27 | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, Replicase, ... | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
9D11
 
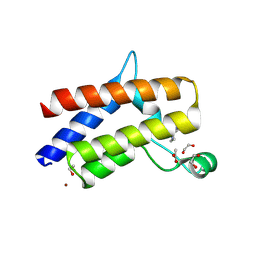 | | Smarca2 Bromodomain in complex with compound 22 | | Descriptor: | (12'S)-4'-chloro-10'-(piperidin-4-yl)-5'H-spiro[cyclohexane-1,7'-indolo[1,2-a]quinazolin]-5'-one, 1,2-ETHANEDIOL, ACETATE ION, ... | | Authors: | Meagher, J.L, Stuckey, J.A. | | Deposit date: | 2024-08-07 | | Release date: | 2025-01-15 | | Last modified: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (1.684 Å) | | Cite: | Discovery of High-Affinity SMARCA2/4 Bromodomain Ligands and Development of Potent and Exceptionally Selective SMARCA2 PROTAC Degraders.
J.Med.Chem., 68, 2025
|
|
9D12
 
 | | Smarca2 Bromodomain in complex with compound 15 | | Descriptor: | (12'R)-4'-chloro-9'-(piperidin-4-yl)-5'H-spiro[cyclohexane-1,7'-indolo[1,2-a]quinazolin]-5'-one, ACETATE ION, Isoform Short of Probable global transcription activator SNF2L2, ... | | Authors: | Meagher, J.L, Stuckey, J.A. | | Deposit date: | 2024-08-07 | | Release date: | 2025-01-15 | | Last modified: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (2.099 Å) | | Cite: | Discovery of High-Affinity SMARCA2/4 Bromodomain Ligands and Development of Potent and Exceptionally Selective SMARCA2 PROTAC Degraders.
J.Med.Chem., 68, 2025
|
|
2W42
 
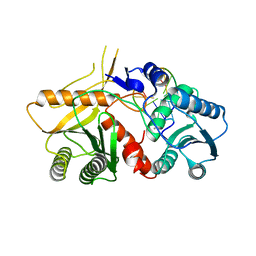 | | THE STRUCTURE OF A PIWI PROTEIN FROM ARCHAEOGLOBUS FULGIDUS COMPLEXED WITH A 16NT DNA DUPLEX. | | Descriptor: | 5'-D(*GP*TP*CP*GP*AP*AP*TP*TP)-3', 5'-D(*TP*TP*CP*GP*AP*CP*GP*CP)-3', MANGANESE (II) ION, ... | | Authors: | Parker, J.S, Roe, S.M, Barford, D. | | Deposit date: | 2008-11-19 | | Release date: | 2008-12-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Enhancement of the Seed-Target Recognition Step in RNA Silencing by a Piwi-Mid Domain Protein
Mol.Cell, 33, 2009
|
|
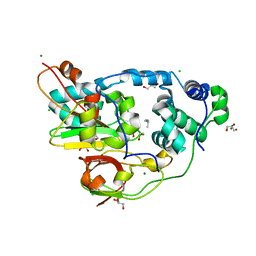
2QDY
 
 | | Crystal Structure of Fe-type NHase from Rhodococcus erythropolis AJ270 | | Descriptor: | 2-METHYLPROPAN-1-AMINE, CHLORIDE ION, FE (III) ION, ... | | Authors: | Song, L, Shi, J, Xue, Z, Wang, M.-X, Qian, S. | | Deposit date: | 2007-06-22 | | Release date: | 2008-05-13 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | High resolution X-ray molecular structure of the nitrile hydratase from Rhodococcus erythropolis AJ270 reveals posttranslational oxidation of two cysteines into sulfinic acids and a novel biocatalytic nitrile hydration mechanism
Biochem.Biophys.Res.Commun., 362, 2007
|
|
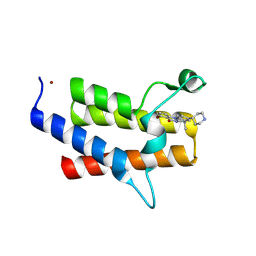
9E1K
 
 | | Discovery of Potent, Highly Selective and Efficacious SMARCA2 Degraders - Compound 11 | | Descriptor: | (12S)-4-bromo-7,7-dimethyl-9-(piperidin-4-yl)indolo[1,2-a]quinazolin-5(7H)-one, Isoform Short of Probable global transcription activator SNF2L2, ZINC ION | | Authors: | Strickland, C, Rice, C. | | Deposit date: | 2024-10-21 | | Release date: | 2024-12-18 | | Last modified: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Discovery of Potent, Highly Selective, and Efficacious SMARCA2 Degraders.
J.Med.Chem., 68, 2025
|
|
9E31
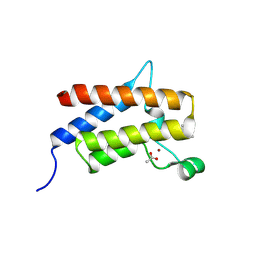 
 | | Discovery of Potent, Highly Selective and Efficacious SMARCA2 Degraders - Compound 6 | | Descriptor: | (12'R)-4'-chloro-9'-(piperidin-4-yl)-5'H-spiro[cyclohexane-1,7'-indolo[1,2-a]quinazolin]-5'-one, ACETATE ION, Isoform Short of Probable global transcription activator SNF2L2, ... | | Authors: | Strickland, C, Rice, C. | | Deposit date: | 2024-10-23 | | Release date: | 2024-12-18 | | Last modified: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Discovery of Potent, Highly Selective, and Efficacious SMARCA2 Degraders.
J.Med.Chem., 68, 2025
|
|
9E30
 
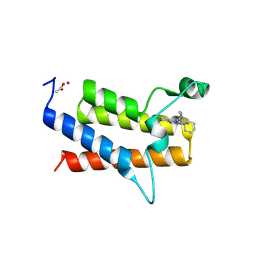 | | Discovery of Potent, Highly Selective and Efficacious SMARCA2 Degraders - Compound 13 | | Descriptor: | (12'S)-4'-chloro-10'-(piperidin-4-yl)-5'H-spiro[cyclohexane-1,7'-indolo[1,2-a]quinazolin]-5'-one, ACETATE ION, Isoform Short of Probable global transcription activator SNF2L2, ... | | Authors: | Strickland, C, Rice, C. | | Deposit date: | 2024-10-23 | | Release date: | 2024-12-18 | | Last modified: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Discovery of Potent, Highly Selective, and Efficacious SMARCA2 Degraders.
J.Med.Chem., 68, 2025
|
|
9DOM
 
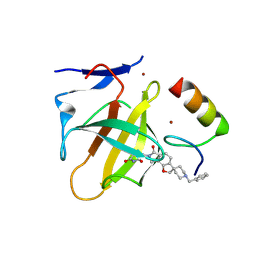 | | PVTX-405: A Potent, Highly Selective, and Orally Efficacious Molecular Glue Degrader of IKZF2 for Cancer Immunotherapy | | Descriptor: | (3S)-3-(1'-benzyl-6-oxo-6,8-dihydro-2H,7H-spiro[furo[2,3-e]isoindole-3,4'-piperidin]-7-yl)piperidine-2,6-dione, Protein cereblon, ZINC ION, ... | | Authors: | Strickland, C.O, Rice, C.T. | | Deposit date: | 2024-09-19 | | Release date: | 2025-05-14 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Development of PVTX-405 as a potent and highly selective molecular glue degrader of IKZF2 for cancer immunotherapy.
Nat Commun, 16, 2025
|
|
9JDZ
 
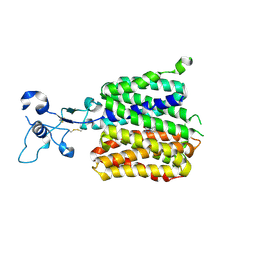 | | Human URAT1 bound to lesinurad | | Descriptor: | Solute carrier family 22 member 12, lesinurad | | Authors: | Wu, C, Xu, H.E. | | Deposit date: | 2024-09-01 | | Release date: | 2024-10-16 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Molecular mechanisms of urate transport by the native human URAT1 and its inhibition by anti-gout drugs.
Cell Discov, 11, 2025
|
|
2XTX
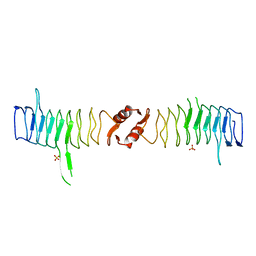 
 | | Structure of QnrB1 (M102R-Trypsin Treated), a plasmid-mediated fluoroquinolone resistance protein | | Descriptor: | QNRB1, SULFATE ION | | Authors: | Vetting, M.W, Hegde, S.S, Park, C.H, Jacoby, G.A, Hooper, D.C, Blanchard, J.S. | | Deposit date: | 2010-10-12 | | Release date: | 2010-10-20 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of Qnrb1, a Plasmid-Mediated Fluoroquinolone Resistance Factor.
J.Biol.Chem., 286, 2011
|
|
8WKJ
 
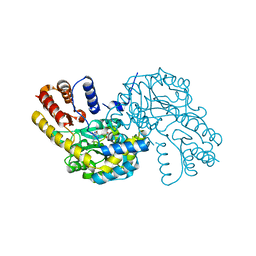 | |
8WOU
 
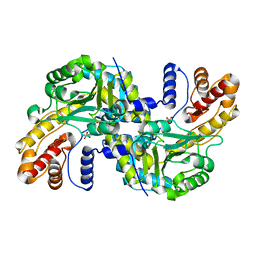 | |
2R6W
 
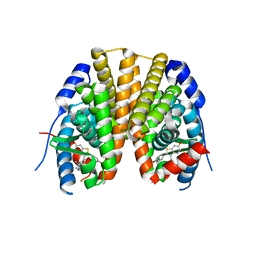 | | Estrogen receptor alpha ligand-binding domain complexed to a SERM | | Descriptor: | Estrogen receptor, [6-hydroxy-2-(4-hydroxyphenyl)-1-benzothien-3-yl]{4-[2-(4-methylpiperidin-1-yl)ethoxy]phenyl}methanone | | Authors: | Wang, Y. | | Deposit date: | 2007-09-06 | | Release date: | 2008-04-08 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Prediction of the tissue-specificity of selective estrogen receptor modulators by using a single biochemical method.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
2XTY
 
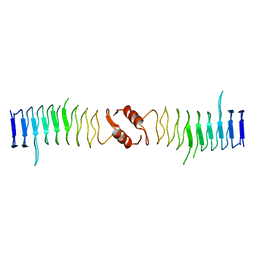 | | Structure of QnrB1 (R167E-Trypsin Treated), a plasmid-mediated fluoroquinolone resistance protein | | Descriptor: | QNRB1 | | Authors: | Vetting, M.W, Hegde, S.S, Park, C.H, Jacoby, G.A, Hooper, D.C, Blanchard, J.S. | | Deposit date: | 2010-10-13 | | Release date: | 2010-10-27 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of Qnrb1, a Plasmid-Mediated Fluoroquinolone Resistance Factor.
J.Biol.Chem., 286, 2011
|
|
2XTW
 
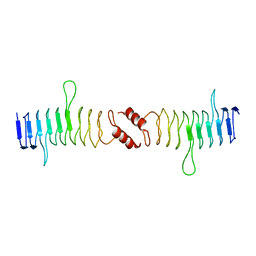 | | Structure of QnrB1 (Full length), a plasmid-mediated fluoroquinolone resistance protein | | Descriptor: | QNRB1 | | Authors: | Vetting, M.W, Hegde, S.S, Park, C.H, Jacoby, G.A, Hooper, D.C, Blanchard, J.S. | | Deposit date: | 2010-10-12 | | Release date: | 2010-10-20 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.803 Å) | | Cite: | Structure of Qnrb1, a Plasmid-Mediated Fluoroquinolone Resistance Factor.
J.Biol.Chem., 286, 2011
|
|
2R6Y
 
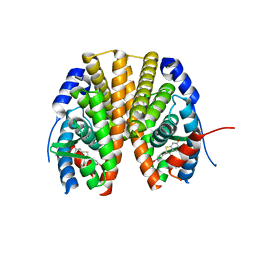 | | Estrogen receptor alpha ligand-binding domain in complex with a SERM | | Descriptor: | Estrogen receptor, [6-hydroxy-2-(4-hydroxyphenyl)-1-benzothien-3-yl][4-(2-pyrrolidin-1-ylethoxy)phenyl]methanone | | Authors: | Wang, Y. | | Deposit date: | 2007-09-06 | | Release date: | 2008-04-08 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Prediction of the tissue-specificity of selective estrogen receptor modulators by using a single biochemical method.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
3LAL
 
 | | Crystal structure of HIV-1 reverse transcriptase in complex with N1-ethyl pyrimidinedione non-nucleoside inhibitor | | Descriptor: | 3-{[3-ethyl-5-(1-methylethyl)-2,6-dioxo-1,2,3,6-tetrahydropyrimidin-4-yl]carbonyl}-5-methylbenzonitrile, HIV Reverse transcriptase, SULFATE ION | | Authors: | Lansdon, E.B, Mitchell, M.L. | | Deposit date: | 2010-01-06 | | Release date: | 2010-02-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | N1-Alkyl pyrimidinediones as non-nucleoside inhibitors of HIV-1 reverse transcriptase.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
3LAK
 
 | | Crystal structure of HIV-1 reverse transcriptase in complex with N1-heterocycle pyrimidinedione non-nucleoside inhibitor | | Descriptor: | 3-({3-[(2-amino-6-fluoropyridin-4-yl)methyl]-5-(1-methylethyl)-2,6-dioxo-1,2,3,6-tetrahydropyrimidin-4-yl}carbonyl)-5-methylbenzonitrile, CHLORIDE ION, HIV Reverse transcriptase, ... | | Authors: | Lansdon, E.B, Mitchell, M.L. | | Deposit date: | 2010-01-06 | | Release date: | 2010-02-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | N1-Heterocyclic pyrimidinediones as non-nucleoside inhibitors of HIV-1 reverse transcriptase.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
