1MAE
 
 | |
1MAF
 
 | |
1NOP
 
 | | Crystal structure of human tyrosyl-DNA phosphodiesterase (Tdp1) in complex with vanadate, DNA and a human topoisomerase I-derived peptide | | Descriptor: | 5'-D(*AP*GP*AP*GP*TP*T)-3', VANADATE ION, topoisomerase I-derived peptide, ... | | Authors: | Davies, D.R, Interthal, H, Champoux, J.J, Hol, W.G.J. | | Deposit date: | 2003-01-16 | | Release date: | 2003-03-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of a transition state mimic for Tdp1 assembled from vanadate, DNA, and a topoisomerase I-derived peptide
Chem.Biol., 10, 2003
|
|
1RL4
 
 | | Plasmodium falciparum peptide deformylase complex with inhibitor | | Descriptor: | (2R)-2-{[FORMYL(HYDROXY)AMINO]METHYL}HEXANOIC ACID, 2-{N'-[2-(5-AMINO-1-PHENYLCARBAMOYL-PENTYLCARBAMOYL)-HEXYL]-HYDRAZINOMETHYL}-HEXANOIC ACID(5-AMINO-1-PHENYLCARBAMOYL-PENTYL)-AMIDE, COBALT (II) ION, ... | | Authors: | Robien, M.A, Nguyen, K.T, Kumar, A, Hirsh, I, Turley, S, Pei, D, Hol, W.G.J. | | Deposit date: | 2003-11-24 | | Release date: | 2003-12-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | An improved crystal form of Plasmodium falciparum peptide deformylase.
Protein Sci., 13, 2004
|
|
1HC1
 
 | |
1HCY
 
 | |
1QCB
 
 | |
1P2P
 
 | | STRUCTURE OF PORCINE PANCREATIC PHOSPHOLIPASE A2 AT 2.6 ANGSTROMS RESOLUTION AND COMPARISON WITH BOVINE PHOSPHOLIPASE A2 | | Descriptor: | CALCIUM ION, PHOSPHOLIPASE A2 | | Authors: | Dijkstra, B.W, Renetseder, R, Kalk, K.H, Hol, W.G.J, Drenth, J. | | Deposit date: | 1983-06-27 | | Release date: | 1983-09-15 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of porcine pancreatic phospholipase A2 at 2.6 A resolution and comparison with bovine phospholipase A2.
J.Mol.Biol., 168, 1983
|
|
1CHQ
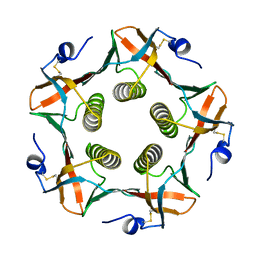 
 | |
1CHP
 
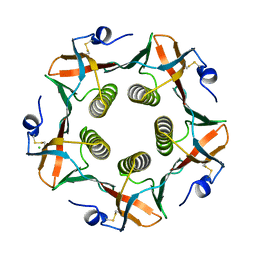 | |
1MU9
 
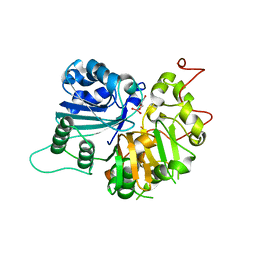 | | Crystal Structure of a Human Tyrosyl-DNA Phosphodiesterase (Tdp1)-Vanadate Complex | | Descriptor: | GLYCEROL, Tyrosyl-DNA Phosphodiesterase, VANADATE ION | | Authors: | Davies, D.R, Interthal, H, Champoux, J.J, Hol, W.G.J. | | Deposit date: | 2002-09-23 | | Release date: | 2003-01-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Insights Into Substrate Binding and Catalytic Mechanism of Human Tyrosyl-DNA Phosphodiesterase (Tdp1) from Vanadate- and Tungstate-Inhibited Structures
J.Mol.Biol., 324, 2002
|
|
1MU7
 
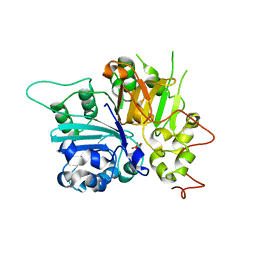 | | Crystal Structure of a Human Tyrosyl-DNA Phosphodiesterase (Tdp1)-Tungstate Complex | | Descriptor: | GLYCEROL, TUNGSTATE(VI)ION, Tyrosyl-DNA Phosphodiesterase | | Authors: | Davies, D.R, Interthal, H, Champoux, J.J, Hol, W.G.J. | | Deposit date: | 2002-09-23 | | Release date: | 2003-01-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Insights Into Substrate Binding and Catalytic Mechanism of Human Tyrosyl-DNA Phosphodiesterase (Tdp1) from Vanadate- and Tungstate-Inhibited Structures
J.Mol.Biol., 324, 2003
|
|
1EEI
 
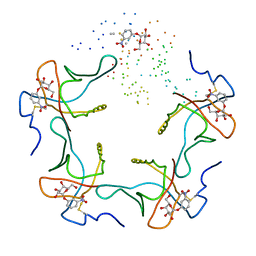 | |
1U8R
 
 | | Crystal Structure of an IdeR-DNA Complex Reveals a Conformational Change in Activated IdeR for Base-specific Interactions | | Descriptor: | COBALT (II) ION, Iron-dependent repressor ideR, SODIUM ION, ... | | Authors: | Wisedchaisri, G, Holmes, R.K, Hol, W.G.J. | | Deposit date: | 2004-08-06 | | Release date: | 2004-10-05 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Crystal Structure of an IdeR-DNA Complex Reveals a Conformational Change in Activated IdeR for Base-specific Interactions.
J.Mol.Biol., 342, 2004
|
|
1LTS
 
 | |
1LTA
 
 | | 2.2 ANGSTROMS CRYSTAL STRUCTURE OF E. COLI HEAT-LABILE ENTEROTOXIN (LT) WITH BOUND GALACTOSE | | Descriptor: | HEAT-LABILE ENTEROTOXIN, SUBUNIT A, SUBUNIT B, ... | | Authors: | Merritt, E.A, Sixma, T.K, Kalk, K.H, Van Zanten, B.A.M, Hol, W.G.J. | | Deposit date: | 1993-09-15 | | Release date: | 1994-01-31 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Galactose-binding site in Escherichia coli heat-labile enterotoxin (LT) and cholera toxin (CT).
Mol.Microbiol., 13, 1994
|
|
1SYR
 
 | |
1TC5
 
 | |
1UV7
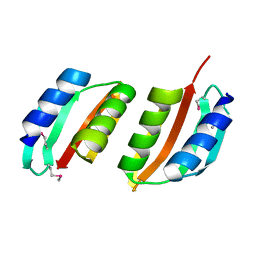 
 | | periplasmic domain of EpsM from Vibrio cholerae | | Descriptor: | GENERAL SECRETION PATHWAY PROTEIN M | | Authors: | Abendroth, J, Hol, W.G.J. | | Deposit date: | 2004-01-15 | | Release date: | 2004-04-23 | | Last modified: | 2019-05-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Crystal Structure of the Periplasmic Domain of the Type II Secretion System Protein Epsm from Vibrio Cholerae: The Simplest Version of the Ferredoxin Fold
J.Mol.Biol., 338, 2004
|
|
1LVL
 
 | |
1NOL
 
 | | OXYGENATED HEMOCYANIN (SUBUNIT TYPE II) | | Descriptor: | CALCIUM ION, COPPER (II) ION, HEMOCYANIN (SUBUNIT TYPE II), ... | | Authors: | Hazes, B, Hol, W.G.J. | | Deposit date: | 1995-10-17 | | Release date: | 1996-03-08 | | Last modified: | 2018-04-18 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of deoxygenated Limulus polyphemus subunit II hemocyanin at 2.18 A resolution: clues for a mechanism for allosteric regulation.
Protein Sci., 2, 1993
|
|
1HTL
 
 | |
1LTG
 
 | |
1P9W
 
 | | Crystal Structure of Vibrio cholerae putative NTPase EpsE | | Descriptor: | General secretion pathway protein E, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ZINC ION | | Authors: | Robien, M.A, Krumm, B.E, Sandkvist, M, Hol, W.G.J. | | Deposit date: | 2003-05-12 | | Release date: | 2003-10-14 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the extracellular protein secretion NTPase EpsE of Vibrio cholerae
J.Mol.Biol., 333, 2003
|
|
1M66
 
 | | Crystal Structure of Leishmania mexicana GPDH Complexed with Inhibitor 2-bromo-6-chloro-purine | | Descriptor: | 2-BROMO-6-CHLORO-PURINE, Glycerol-3-phosphate dehydrogenase, PALMITIC ACID | | Authors: | Choe, J, Suresh, S, Wisedchaisri, G, Kennedy, K.J, Gelb, M.H, Hol, W.G.J. | | Deposit date: | 2002-07-12 | | Release date: | 2002-12-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Anomalous differences of light elements in determining precise binding modes of ligands to
glycerol-3-phosphate dehydrogenase
Chem.Biol., 9, 2002
|
|
