6UPV
 
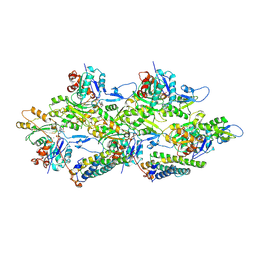 | | Alpha-E-catenin ABD-F-actin complex | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Actin, alpha skeletal muscle, ... | | Authors: | Mei, L, Alushin, G.M. | | Deposit date: | 2019-10-18 | | Release date: | 2020-09-30 | | Last modified: | 2020-10-07 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Molecular mechanism for direct actin force-sensing by alpha-catenin.
Elife, 9, 2020
|
|
6UPW
 
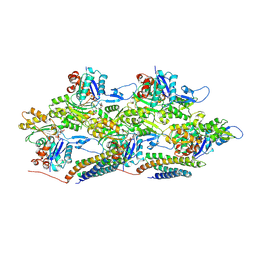 | | Metavinculin ABD-F-actin complex | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Actin, alpha skeletal muscle, ... | | Authors: | Mei, L, Alushin, G.M. | | Deposit date: | 2019-10-18 | | Release date: | 2020-09-30 | | Last modified: | 2020-10-07 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Molecular mechanism for direct actin force-sensing by alpha-catenin.
Elife, 9, 2020
|
|
6L5Z
 | |
6LJM
 
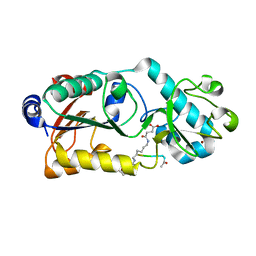 | |
6L4Z
 
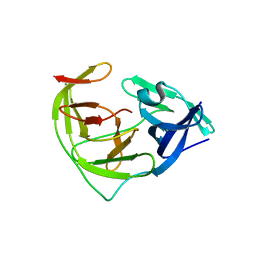 | | Crystal structure of Zika NS2B-NS3 protease with compound 6 | | Descriptor: | 4-(hydroxymethyl)benzoic acid, Genome polyprotein | | Authors: | Quek, J.P. | | Deposit date: | 2019-10-21 | | Release date: | 2020-07-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Identification and structural characterization of small molecule fragments targeting Zika virus NS2B-NS3 protease.
Antiviral Res., 175, 2020
|
|
6LJN
 
 | |
6L50
 
 | | Crystal structure of Zika NS2B-NS3 protease with compound 16 | | Descriptor: | 2-sulfanylidene-1,3-thiazolidin-4-one, NS3 protease, Serine protease subunit NS2B | | Authors: | Quek, J.P. | | Deposit date: | 2019-10-21 | | Release date: | 2020-07-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Identification and structural characterization of small molecule fragments targeting Zika virus NS2B-NS3 protease.
Antiviral Res., 175, 2020
|
|
6LJK
 
 | |
7U2L
 
 | | C5guano-uOR-Gi-scFv16 | | Descriptor: | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, Guanine nucleotide-binding protein G(i) subunit alpha-1, ... | | Authors: | Wang, H, Qu, Q, Skiniotis, G, Kobilka, B. | | Deposit date: | 2022-02-24 | | Release date: | 2022-05-04 | | Last modified: | 2023-02-08 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structure-based design of bitopic ligands for the μ-opioid receptor.
Nature, 613, 2023
|
|
7U2K
 
 | | C6-guano bound Mu Opioid Receptor-Gi Protein Complex | | Descriptor: | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, Guanine nucleotide-binding protein G(i) subunit alpha-1, ... | | Authors: | Wang, H, Kobilka, B. | | Deposit date: | 2022-02-24 | | Release date: | 2022-12-07 | | Last modified: | 2023-02-08 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Structure-based design of bitopic ligands for the μ-opioid receptor.
Nature, 613, 2023
|
|
3LM5
 
 | | Crystal Structure of human Serine/Threonine Kinase 17B (STK17B) in complex with Quercetin | | Descriptor: | 3,5,7,3',4'-PENTAHYDROXYFLAVONE, Serine/threonine-protein kinase 17B | | Authors: | Ugochukwu, E, Soundararajan, M, Rellos, P, Fedorov, O, Phillips, C, Wang, J, Hapka, E, Filippakopoulos, P, Chaikuad, A, Pike, A.C.W, Carpenter, L, Vollmar, M, von Delft, F, Bountra, C, Arrowsmith, C.H, Weigelt, J, Edwards, A, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-01-29 | | Release date: | 2010-03-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | A Chemical Probe for Dark Kinase STK17B Derives Its Potency and High Selectivity through a Unique P-Loop Conformation.
J.Med.Chem., 63, 2020
|
|
7TW3
 
 | | Cryo-EM structure of human ankyrin complex (B2P1A1) from red blood cell | | Descriptor: | Ankyrin-1, Band 3 anion transport protein, Protein 4.2 | | Authors: | Xia, X, Liu, S.H, Zhou, Z.H. | | Deposit date: | 2022-02-06 | | Release date: | 2022-06-08 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | Structure, dynamics and assembly of the ankyrin complex on human red blood cell membrane.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7TVZ
 
 | | Cryo-EM structure of human band 3-protein 4.2 complex in diagonal conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Band 3 anion transport protein, CHOLESTEROL, ... | | Authors: | Xia, X, Liu, S.H, Zhou, Z.H. | | Deposit date: | 2022-02-06 | | Release date: | 2022-06-08 | | Last modified: | 2022-07-27 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structure, dynamics and assembly of the ankyrin complex on human red blood cell membrane.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7TW2
 
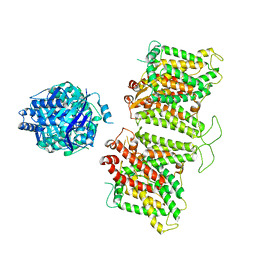 | |
7TW0
 
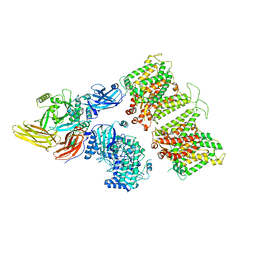 | | Cryo-EM structure of human band 3-protein 4.2 complex in vertical conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Band 3 anion transport protein, Protein 4.2 | | Authors: | Xia, X, Liu, S.H, Zhou, Z.H. | | Deposit date: | 2022-02-06 | | Release date: | 2022-06-08 | | Last modified: | 2022-07-27 | | Method: | ELECTRON MICROSCOPY (4.6 Å) | | Cite: | Structure, dynamics and assembly of the ankyrin complex on human red blood cell membrane.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7TW1
 
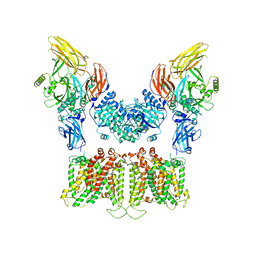 | |
7TW5
 
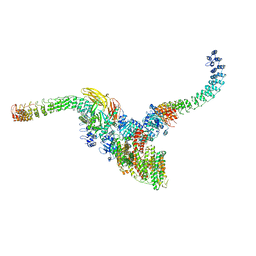 | | Cryo-EM structure of human ankyrin complex (B2P1A2) from red blood cell | | Descriptor: | Ankyrin-1, Band 3 anion transport protein, Protein 4.2 | | Authors: | Xia, X, Liu, S.H, Zhou, Z.H. | | Deposit date: | 2022-02-06 | | Release date: | 2022-06-08 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (5.7 Å) | | Cite: | Structure, dynamics and assembly of the ankyrin complex on human red blood cell membrane.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7TW6
 
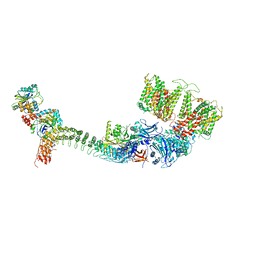 | | Cryo-EM structure of human ankyrin complex (B4P1A1) from red blood cell | | Descriptor: | Ankyrin-1, Band 3 anion transport protein, Protein 4.2 | | Authors: | Xia, X, Liu, S.H, Zhou, Z.H. | | Deposit date: | 2022-02-06 | | Release date: | 2022-06-08 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (5.6 Å) | | Cite: | Structure, dynamics and assembly of the ankyrin complex on human red blood cell membrane.
Nat.Struct.Mol.Biol., 29, 2022
|
|
6LW2
 
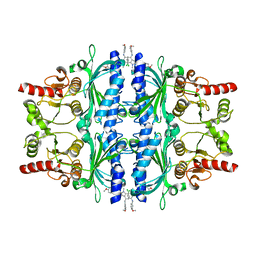 | | The N-arylsulfonyl-indole-2-carboxamide-based inhibitors against fructose-1,6-bisphosphatase | | Descriptor: | 7-chloranyl-4-[(3-methoxyphenyl)amino]-N-(4-methoxyphenyl)sulfonyl-1-methyl-indole-2-carboxamide, Fructose-1,6-bisphosphatase 1 | | Authors: | Wang, X.Y, Zhou, J, Xu, B.L. | | Deposit date: | 2020-02-07 | | Release date: | 2020-05-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Discovery of N -Arylsulfonyl-Indole-2-Carboxamide Derivatives as Potent, Selective, and Orally Bioavailable Fructose-1,6-Bisphosphatase Inhibitors-Design, Synthesis, In Vivo Glucose Lowering Effects, and X-ray Crystal Complex Analysis.
J.Med.Chem., 63, 2020
|
|
7E3O
 
 | |
4J51
 
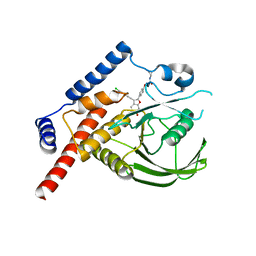 | |
4B4A
 
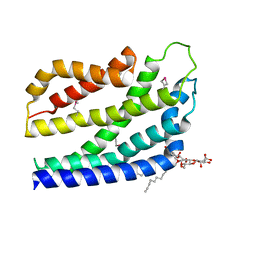 | | Structure of the TatC core of the twin arginine protein translocation system | | Descriptor: | Lauryl Maltose Neopentyl Glycol, SEC-INDEPENDENT PROTEIN TRANSLOCASE PROTEIN TATC | | Authors: | Rollauer, S.E, Tarry, M.J, Jaaskelainen, M, Graham, J.E, Jaeger, F, Krehenbrink, M, Roversi, P, McDowell, M.A, Stansfeld, P.J, Johnson, S, Liu, S.M, Lukey, M.J, Marcoux, J, Robinson, C.V, Sansom, M.S, Palmer, T, Hogbom, M, Berks, B.C, Lea, S.M. | | Deposit date: | 2012-07-30 | | Release date: | 2012-12-05 | | Last modified: | 2012-12-19 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structure of the Tatc Core of the Twin-Arginine Protein Transport System.
Nature, 49, 2012
|
|
4G8B
 
 | |
4G9E
 
 | |
4GA7
 
 | | Crystal structure of human serpinB1 mutant | | Descriptor: | Leukocyte elastase inhibitor | | Authors: | Wang, L, Li, Q, Wu, L, Zhang, K, Tong, L, Sun, F, Fan, Z. | | Deposit date: | 2012-07-25 | | Release date: | 2013-01-16 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Identification of SERPINB1 as a physiological inhibitor of human granzyme H
J.Immunol., 190, 2013
|
|
