6S5V
 
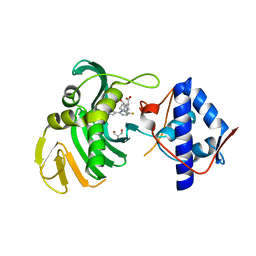 | | Crystal structure of the Cap-Midlink region of the H5N1 Influenza A virus polymerase in complex with a Cap-domain binding analogue | | Descriptor: | (1~{S},2~{S},3~{S},6~{R})-2-[[2-[5,7-bis(fluoranyl)-1~{H}-indol-3-yl]-5-fluoranyl-pyrimidin-4-yl]amino]-3,6-dimethyl-cyclohexane-1-carboxylic acid, GLYCEROL, POTASSIUM ION, ... | | Authors: | Keown, J.R, Fodor, E, Grimes, J.M. | | Deposit date: | 2019-07-02 | | Release date: | 2019-11-06 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Design, Synthesis, and Biological Evaluation of Novel Indoles Targeting the Influenza PB2 Cap Binding Region.
J.Med.Chem., 62, 2019
|
|
4R3B
 
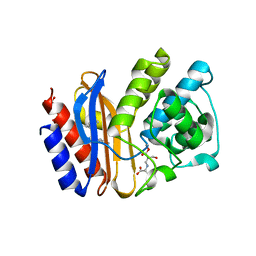 | |
3KD5
 
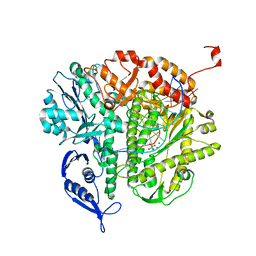 | |
4R5P
 
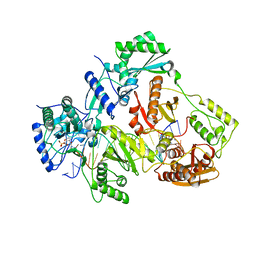 | | Crystal structure of HIV-1 reverse transcriptase (RT) with DNA and a nucleoside triphosphate mimic alpha-carboxy nucleoside phosphonate inhibitor | | Descriptor: | 5'-D(*CP*AP*GP*TP*CP*CP*CP*TP*GP*TP*TP*CP*GP*GP*(MRG)P*CP*GP*CP*CP*G)-3', 5'-D(*TP*GP*GP*AP*CP*GP*GP*CP*GP*CP*CP*CP*GP*AP*AP*CP*AP*GP*GP*GP*AP*CP*TP*G)-3', HIV-1 reverse transcriptase, ... | | Authors: | Das, K, Martinez, S.E, Arnold, E. | | Deposit date: | 2014-08-21 | | Release date: | 2015-03-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.894 Å) | | Cite: | Alpha-carboxy nucleoside phosphonates as universal nucleoside triphosphate mimics.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
3KD1
 
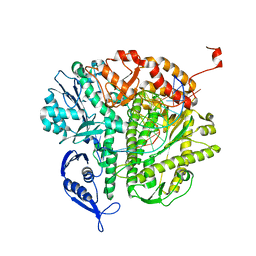 | |
6RG0
 
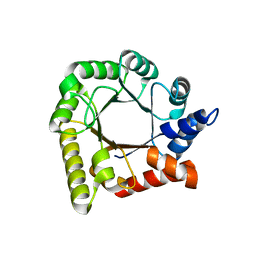 | | Structure of pdxj | | Descriptor: | Pyridoxine 5'-phosphate synthase | | Authors: | Rohweder, B, Rajendran, C, Sterner, R. | | Deposit date: | 2019-04-16 | | Release date: | 2020-05-13 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.074 Å) | | Cite: | Library Selection with a Randomized Repertoire of ( beta alpha )8-Barrel Enzymes Results in Unexpected Induction of Gene Expression.
Biochemistry, 58, 2019
|
|
1R63
 
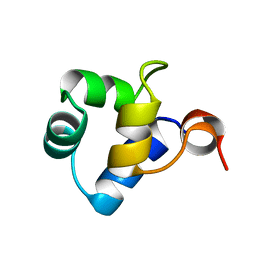 | | STRUCTURAL ROLE OF A BURIED SALT BRIDGE IN THE 434 REPRESSOR DNA-BINDING DOMAIN, NMR, 20 STRUCTURES | | Descriptor: | REPRESSOR PROTEIN FROM BACTERIOPHAGE 434 | | Authors: | Pervushin, K.V, Billeter, M, Siegal, G, Wuthrich, K. | | Deposit date: | 1996-11-08 | | Release date: | 1997-06-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural role of a buried salt bridge in the 434 repressor DNA-binding domain.
J.Mol.Biol., 264, 1996
|
|
4N0S
 
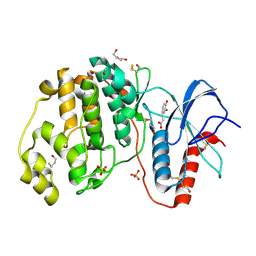 | | Complex of ERK2 with caffeic acid | | Descriptor: | CAFFEIC ACID, DI(HYDROXYETHYL)ETHER, DIMETHYL SULFOXIDE, ... | | Authors: | Kurinov, I, Malakhova, M. | | Deposit date: | 2013-10-02 | | Release date: | 2014-08-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7992 Å) | | Cite: | Caffeic Acid Directly Targets ERK1/2 to Attenuate Solar UV-Induced Skin Carcinogenesis.
Cancer Prev Res (Phila), 7, 2014
|
|
1TRH
 
 | |
3HBR
 
 | | Crystal structure of OXA-48 beta-lactamase | | Descriptor: | 1,2-ETHANEDIOL, OXA-48 | | Authors: | Calderone, V, Mangani, S, Benvenuti, M, Rossolini, G.M, Docquier, J.D. | | Deposit date: | 2009-05-05 | | Release date: | 2009-06-23 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the OXA-48 beta-lactamase reveals mechanistic diversity among class D carbapenemases.
Chem.Biol., 16, 2009
|
|
4ZH3
 
 | |
4ZH4
 
 | | Crystal structure of Escherichia coli RNA polymerase in complex with CBRP18 | | Descriptor: | 5-(4-fluorophenyl)-4-[4-fluoro-3-(trifluoromethyl)phenyl]-1H-pyrazole, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | Authors: | Feng, Y, Ebright, R.H. | | Deposit date: | 2015-04-24 | | Release date: | 2015-08-05 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.993 Å) | | Cite: | Structural Basis of Transcription Inhibition by CBR Hydroxamidines and CBR Pyrazoles.
Structure, 23, 2015
|
|
7MXJ
 
 | |
7MXB
 
 | |
7MXK
 
 | |
5SAV
 
 | | DDR1, N-[2-[3-(2-aminopyrimidin-5-yl)oxyphenyl]ethyl]-3-(trifluoromethoxy)benzamide, 1.760A, P212121, Rfree=23.5% | | Descriptor: | Epithelial discoidin domain-containing receptor 1, IODIDE ION, N-(2-{3-[(2-aminopyrimidin-5-yl)oxy]phenyl}ethyl)-3-(trifluoromethoxy)benzamide | | Authors: | Stihle, M, Richter, H, Benz, J, Hochstrasser, R, Rudolph, M.G. | | Deposit date: | 2021-06-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Crystal Structure of a DDR1 complex
To be published
|
|
5SAY
 
 | | DDR1, N-[2-[3-(2-aminopyrimidin-5-yl)oxyphenyl]ethyl]-3-(trifluoromethoxy)benzamide, 2.190A, P1211, Rfree=27.7% | | Descriptor: | Epithelial discoidin domain-containing receptor 1, IODIDE ION, N-(2-{3-[(2-aminopyrimidin-5-yl)oxy]phenyl}ethyl)-3-(trifluoromethoxy)benzamide | | Authors: | Stihle, M, Richter, H, Benz, J, Hochstrasser, R, Rudolph, M.G. | | Deposit date: | 2021-06-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Crystal Structure of a DDR1 complex
To be published
|
|
5SB1
 
 | | DDR1, 4-chloro-N-[(3S,4R)-4-phenylpyrrolidin-3-yl]-3-(1H-pyrrolo[2,3-b]pyridin-5-yloxymethyl)benzamide, 1.530A, P212121, Rfree=21.4% | | Descriptor: | 4-chloro-N-[(3S,4R)-4-phenylpyrrolidin-3-yl]-3-{[(1H-pyrrolo[2,3-b]pyridin-5-yl)oxy]methyl}benzamide, Epithelial discoidin domain-containing receptor 1, SULFATE ION | | Authors: | Stihle, M, Richter, H, Benz, J, Hochstrasser, R, Rudolph, M.G. | | Deposit date: | 2021-06-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Crystal Structure of a DDR1 complex
To be published
|
|
5SB2
 
 | | DDR1, 3-chloro-N-[(1R,2S)-2-phenylcyclopropyl]-5-(1H-pyrrolo[2,3-b]pyridin-5-yloxymethyl)benzamide, 1.600A, P212121, Rfree=23.2% | | Descriptor: | 3-chloro-N-[(1R,2S)-2-phenylcyclopropyl]-5-{[(1H-pyrrolo[2,3-b]pyridin-5-yl)oxy]methyl}benzamide, Epithelial discoidin domain-containing receptor 1, IODIDE ION | | Authors: | Stihle, M, Richter, H, Benz, J, Kocer, B, Hochstrasser, R, Rudolph, M.G. | | Deposit date: | 2021-06-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal Structure of a DDR1 complex
To be published
|
|
5SAZ
 
 | | DDR1, 3-chloro-N-[4-chloro-3-(1H-pyrrolo[2,3-b]pyridin-5-ylcarbamoyl)phenyl]-4-(2-hydroxyethylamino)benzamide, 1.802A, P212121, Rfree=22.2% | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-chloro-5-{3-chloro-4-[(2-hydroxyethyl)amino]benzamido}-N-(1H-pyrrolo[2,3-b]pyridin-5-yl)benzamide, CHLORIDE ION, ... | | Authors: | Stihle, M, Richter, H, Benz, J, Hochstrasser, R, Rudolph, M.G. | | Deposit date: | 2021-06-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of a DDR1 complex
To be published
|
|
5SAU
 
 | | DDR1, 3-[2-(6-aminopyridin-3-yl)ethynyl]-N-[3-(trifluoromethyl)phenyl]benzamide, 1.800A, P212121, Rfree=23.1% | | Descriptor: | 3-[(6-aminopyridin-3-yl)ethynyl]-N-[3-(trifluoromethyl)phenyl]benzamide, Epithelial discoidin domain-containing receptor 1, MALONATE ION | | Authors: | Stihle, M, Richter, H, Benz, J, Hochstrasser, R, Rudolph, M.G. | | Deposit date: | 2021-06-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of a DDR1 complex
To be published
|
|
5SAW
 
 | | DDR1, 2-[3-(2-pyridin-3-ylethynyl)phenyl]-N-[3-(trifluoromethyl)phenyl]acetamide, 1.601A, P212121, Rfree=22.6% | | Descriptor: | 2-{3-[(pyridin-3-yl)ethynyl]phenyl}-N-[3-(trifluoromethyl)phenyl]acetamide, Epithelial discoidin domain-containing receptor 1, IODIDE ION | | Authors: | Stihle, M, Richter, H, Benz, J, Hochstrasser, R, Rudolph, M.G. | | Deposit date: | 2021-06-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.601 Å) | | Cite: | Crystal Structure of a DDR1 complex
To be published
|
|
5SAX
 
 | | DDR1, 2-[3-(2-pyridin-3-ylethynyl)phenyl]-N-[3-(trifluoromethyl)phenyl]acetamide, 1.902A, second P212121 form, Rfree=25.4%, second form | | Descriptor: | 2-{3-[(pyridin-3-yl)ethynyl]phenyl}-N-[3-(trifluoromethyl)phenyl]acetamide, Epithelial discoidin domain-containing receptor 1, IODIDE ION | | Authors: | Stihle, M, Richter, H, Benz, J, Hochstrasser, R, Rudolph, M.G. | | Deposit date: | 2021-06-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.902 Å) | | Cite: | Crystal Structure of a DDR1 complex
To be published
|
|
5SB0
 
 | | DDR1, N-[[2-(2-pyridin-3-yloxyethyl)cyclohexyl]methyl]-3-(trifluoromethoxy)benzamide, 1.970A, P212121, Rfree=25.6% | | Descriptor: | CHLORIDE ION, Epithelial discoidin domain-containing receptor 1, IODIDE ION, ... | | Authors: | Stihle, M, Richter, H, Benz, J, Hochstrasser, R, Rudolph, M.G. | | Deposit date: | 2021-06-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Crystal Structure of a DDR1 complex
To be published
|
|
2R63
 
 | | STRUCTURAL ROLE OF A BURIED SALT BRIDGE IN THE 434 REPRESSOR DNA-BINDING DOMAIN, NMR, 20 STRUCTURES | | Descriptor: | REPRESSOR PROTEIN FROM BACTERIOPHAGE 434 | | Authors: | Pervushin, K.V, Billeter, M, Siegal, G, Wuthrich, K. | | Deposit date: | 1996-11-13 | | Release date: | 1997-06-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural role of a buried salt bridge in the 434 repressor DNA-binding domain.
J.Mol.Biol., 264, 1996
|
|
