2N6U
 
 | | Solution study of Astexin2-dC4 | | Descriptor: | Astexin2-dC4 | | Authors: | Link, A, Maksimov, M.O. | | Deposit date: | 2015-08-28 | | Release date: | 2015-11-11 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Elucidating the Specificity Determinants of the AtxE2 Lasso Peptide Isopeptidase.
J.Biol.Chem., 290, 2015
|
|
2N6V
 
 | | Solution study of Astexin3 | | Descriptor: | ASTEXIN3 | | Authors: | Link, A, Maksimov, M.O. | | Deposit date: | 2015-08-28 | | Release date: | 2015-11-11 | | Last modified: | 2024-11-06 | | Method: | SOLUTION NMR | | Cite: | Elucidating the Specificity Determinants of the AtxE2 Lasso Peptide Isopeptidase.
J.Biol.Chem., 290, 2015
|
|
2N6W
 
 | |
2N6X
 
 | |
2N6Y
 
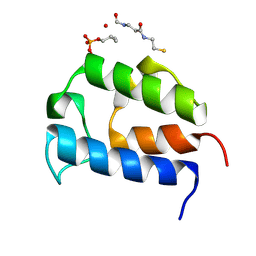 | |
2N6Z
 
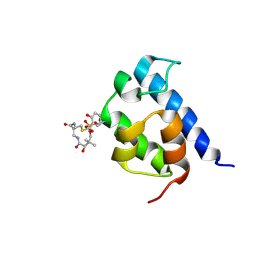 | | Solution structure of the salicylate-loaded ArCP from yersiniabactin synthetase | | Descriptor: | HMWP2 nonribosomal peptide synthetase, S-{2-[(N-{(2R)-2-hydroxy-3,3-dimethyl-4-[(trihydroxy-lambda~5~-phosphanyl)oxy]butanoyl}-beta-alanyl)amino]ethyl} 2-hydroxybenzene-1-carbothioate | | Authors: | Goodrich, A.C, Harden, B.J, Frueh, D.P. | | Deposit date: | 2015-08-31 | | Release date: | 2015-09-23 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of a Nonribosomal Peptide Synthetase Carrier Protein Loaded with Its Substrate Reveals Transient, Well-Defined Contacts.
J.Am.Chem.Soc., 137, 2015
|
|
2N70
 
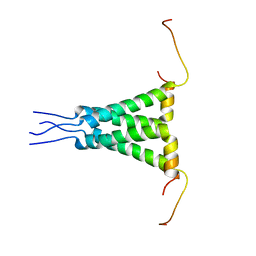 | | Two-fold symmetric structure of the 18-60 construct of S31N M2 from Influenza A in lipid bilayers | | Descriptor: | Matrix protein 2 | | Authors: | Andreas, L.B, Reese, M, Eddy, M.T, Gelev, V, Ni, Q, Miller, E.A, Emsley, L, Pintacuda, G, Chou, J.J, Griffin, R.G. | | Deposit date: | 2015-09-01 | | Release date: | 2015-09-23 | | Last modified: | 2024-05-15 | | Method: | SOLID-STATE NMR | | Cite: | Structure and Mechanism of the Influenza A M218-60 Dimer of Dimers.
J.Am.Chem.Soc., 137, 2015
|
|
2N71
 
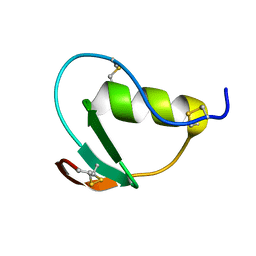 | | NMR structure of CmPI-II, a serin protease inhibitor isolated from mollusk Cenchitis muricatus | | Descriptor: | Protease inhibitor 2 | | Authors: | Cabrera-Munoz, A, Rojas, L, Alonso del Rivero Antigua, M, Pires, J. | | Deposit date: | 2015-09-01 | | Release date: | 2016-09-07 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | NMR structure of CmPI-II, a non-classical Kazal protease inhibitor: Understanding its conformational dynamics and subtilisin A inhibition.
J.Struct.Biol., 206, 2019
|
|
2N72
 
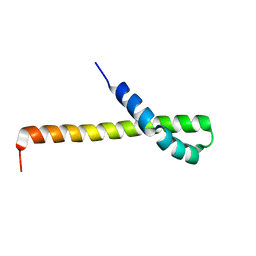 | | Solution structure of the Q domain from ACBD3 | | Descriptor: | Golgi resident protein GCP60 | | Authors: | Veverka, V, Hexnerova, R. | | Deposit date: | 2015-09-02 | | Release date: | 2016-07-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural insights and in vitro reconstitution of membrane targeting and activation of human PI4KB by the ACBD3 protein.
Sci Rep, 6, 2016
|
|
2N73
 
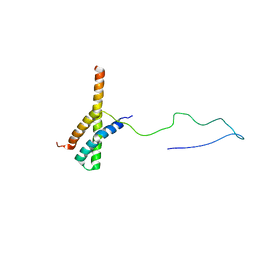 | | Solution structure of the ACBD3:PI4KB complex | | Descriptor: | Golgi resident protein GCP60, Phosphatidylinositol 4-kinase beta | | Authors: | Veverka, V, Hexnerova, R. | | Deposit date: | 2015-09-02 | | Release date: | 2016-07-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural insights and in vitro reconstitution of membrane targeting and activation of human PI4KB by the ACBD3 protein.
Sci Rep, 6, 2016
|
|
2N74
 
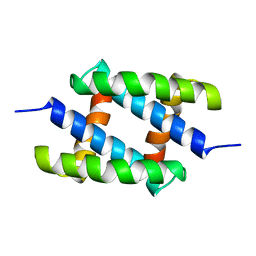 | | Solution Structure of the RNA-Binding domain of non-structural protein 1 from the 1918 H1N1 influenza virus | | Descriptor: | Non-structural protein 1 | | Authors: | Jureka, A.S, Kleinpeter, A.B, Cornilescu, G, Cornilescu, C.C, Schwieters, C.D, Petit, C.M. | | Deposit date: | 2015-09-03 | | Release date: | 2015-09-30 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for a Novel Interaction between the NS1 Protein Derived from the 1918 Influenza Virus and RIG-I.
Structure, 23, 2015
|
|
2N75
 
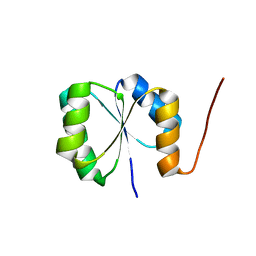 | | Solution NMR Structure of De novo designed protein, Rossmann2x2 Fold, Northeast Structural Genomics Consortium (NESG) Target OR446 | | Descriptor: | De novo designed protein | | Authors: | Liu, G, Lin, Y, Koga, N, Koga, R, Xiao, R, Janjua, H, Pederson, K, Acton, T.B, Kornhaber, G, Everett, J.K, Baker, D, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2015-09-03 | | Release date: | 2016-01-27 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of De novo designed protein, Rossmann2x2 Fold, Northeast Structural Genomics Consortium (NESG) Target OR446
To be Published
|
|
2N76
 
 | | Solution NMR Structure of De novo designed protein LFR1 1 with ferredoxin fold, Northeast Structural Genomics Consortium (NESG) Target OR414 | | Descriptor: | De novo designed protein LFR1 | | Authors: | Liu, G, Lin, Y, Koga, N, Koga, R, Xiao, R, Janjua, H, Pederson, K, Acton, T.B, Kornhaber, G, Everett, J.K, Baker, D, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2015-09-03 | | Release date: | 2016-01-27 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of De novo designed protein LFR1 1 with ferredoxin fold, Northeast Structural Genomics Consortium (NESG) Target OR414
To be Published
|
|
2N77
 
 | |
2N78
 
 | | NMR structure of IF1 from Pseudomonas aeruginosa | | Descriptor: | Translation initiation factor IF-1 | | Authors: | Zhang, Y. | | Deposit date: | 2015-09-04 | | Release date: | 2016-09-07 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | (1)H, (13)C and (15)N resonance assignments and secondary structure analysis of translation initiation factor 1 from Pseudomonas aeruginosa.
Biomol.Nmr Assign., 10, 2016
|
|
2N79
 
 | | The structural and functional effects of the Familial Hypertrophic Cardiomyopathy-linked cardiac troponin C mutation, L29Q | | Descriptor: | CALCIUM ION, Troponin C, slow skeletal and cardiac muscles | | Authors: | Robertson, I.M, Sevrieva, I, Li, M.X, Irving, M, Sun, Y, Sykes, B. | | Deposit date: | 2015-09-06 | | Release date: | 2015-10-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The structural and functional effects of the familial hypertrophic cardiomyopathy-linked cardiac troponin C mutation, L29Q.
J.MOL.CELL.CARDIOL., 87, 2015
|
|
2N7A
 
 | | Solution structure of the human Siglec-8 lectin domain | | Descriptor: | Sialic acid-binding Ig-like lectin 8 | | Authors: | Proepster, J.M, Yang, F, Rabbani, S, Ernst, B, Allain, F.H.-T, Schubert, M. | | Deposit date: | 2015-09-07 | | Release date: | 2016-07-06 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural basis for sulfation-dependent self-glycan recognition by the human immune-inhibitory receptor Siglec-8.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
2N7B
 
 | | Solution structure of the human Siglec-8 lectin domain in complex with 6'sulfo sialyl Lewisx | | Descriptor: | 3-aminopropan-1-ol, N-acetyl-alpha-neuraminic acid-(2-3)-6-O-sulfo-beta-D-galactopyranose-(1-4)-[alpha-L-fucopyranose-(1-3)]2-acetamido-2-deoxy-beta-D-glucopyranose, Sialic acid-binding Ig-like lectin 8 | | Authors: | Proepster, J.M, Yang, F, Rabbani, S, Ernst, B, Allain, F.H.-T, Schubert, M. | | Deposit date: | 2015-09-07 | | Release date: | 2016-07-06 | | Last modified: | 2020-07-29 | | Method: | SOLUTION NMR | | Cite: | Structural basis for sulfation-dependent self-glycan recognition by the human immune-inhibitory receptor Siglec-8.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
2N7C
 
 | |
2N7D
 
 | |
2N7E
 
 | |
2N7F
 
 | | NMR solution structure of muO-conotoxin MfVIA | | Descriptor: | muO-conotoxin MfVIA | | Authors: | Schroeder, C.I, Mobli, M. | | Deposit date: | 2015-09-09 | | Release date: | 2016-04-06 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | Development of a mu O-Conotoxin Analogue with Improved Lipid Membrane Interactions and Potency for the Analgesic Sodium Channel NaV1.8.
J.Biol.Chem., 291, 2016
|
|
2N7G
 
 | |
2N7H
 
 | | Hybrid structure of the Type 1 Pilus of Uropathogenic E.coli | | Descriptor: | FimA | | Authors: | Habenstein, B, Loquet, A, Giller, K, Vasa, S, Becker, S, Habeck, M, Lange, A. | | Deposit date: | 2015-09-11 | | Release date: | 2015-09-23 | | Last modified: | 2015-10-14 | | Method: | SOLID-STATE NMR | | Cite: | Hybrid Structure of the Type 1 Pilus of Uropathogenic Escherichia coli.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
2N7I
 
 | |
