6R9Z
 | |
6RC7
 
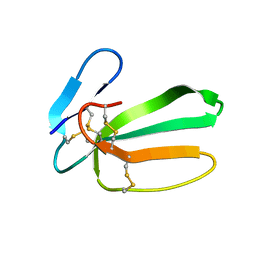 | | NMR structure of cytotoxin 3 from Naja kaouthia in solution, major form | | 分子名称: | Cytotoxin 3 | | 著者 | Dubinnyi, M.A, Dubovskii, P.V, Utkin, Y.N, Arseniev, A.S. | | 登録日 | 2019-04-11 | | 公開日 | 2020-05-13 | | 最終更新日 | 2024-10-09 | | 実験手法 | SOLUTION NMR | | 主引用文献 | The omega-loop of cobra cytotoxins tolerates multiple amino acid substitutions.
Biochem.Biophys.Res.Commun., 558, 2021
|
|
6RFM
 
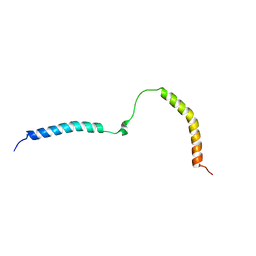 | |
6RH5
 
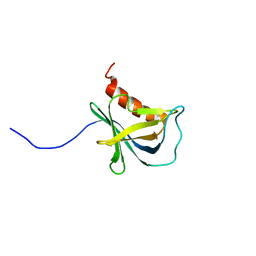 | | Solution structure and 1H, 13C and 15N chemical shift assignments for NECAP1 PHear domain | | 分子名称: | Adaptin ear-binding coat-associated protein 1 | | 著者 | Owen, D.J, Neuhaus, D, Yang, J.-C, Herrmann, T. | | 登録日 | 2019-04-18 | | 公開日 | 2019-09-04 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Temporal Ordering in Endocytic Clathrin-Coated Vesicle Formation via AP2 Phosphorylation.
Dev.Cell, 50, 2019
|
|
6RH6
 
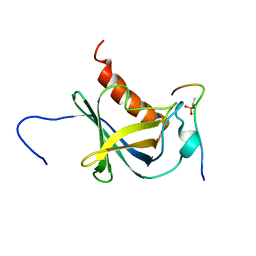 | | Solution structure and 1H, 13C and 15N chemical shift assignments for the complex of NECAP1 PHear domain with phosphorylated AP2 mu2 148-163 | | 分子名称: | AP-2 complex subunit mu, Adaptin ear-binding coat-associated protein 1 | | 著者 | Owen, D.J, Neuhaus, D, Yang, J.-C, Herrmann, T. | | 登録日 | 2019-04-18 | | 公開日 | 2019-09-04 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Temporal Ordering in Endocytic Clathrin-Coated Vesicle Formation via AP2 Phosphorylation.
Dev.Cell, 50, 2019
|
|
6RHY
 
 | |
6RIO
 
 | | Imidazole Polyamide-DNA complex NMR structure (5'-CGATGTACATCG-3') | | 分子名称: | 3-[3-[[4-[[4-[[4-[[4-[[(2~{R})-2-azaniumyl-4-[[1-methyl-4-[[1-methyl-4-[[1-methyl-4-[(1-methylimidazol-2-yl)carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]butanoyl]amino]-1-methyl-imidazol-2-yl]carbonylamino]-1-methyl-pyrrol-2-yl]carbonylamino]-1-methyl-pyrrol-2-yl]carbonylamino]-1-methyl-pyrrol-2-yl]carbonylamino]propanoylamino]propyl-dimethyl-azanium, DNA (5'-(*(DC5)P*GP*AP*TP*GP*TP*AP*CP*AP*TP*CP*(DG3))-3') | | 著者 | Padroni, G, Withers, J.M, Taladriz-Sender, A, Reichenbach, L.F, Parkinson, J.A, Burley, G.A. | | 登録日 | 2019-04-24 | | 公開日 | 2019-06-12 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Sequence-Selective Minor Groove Recognition of a DNA Duplex Containing Synthetic Genetic Components.
J.Am.Chem.Soc., 141, 2019
|
|
6RK3
 
 | | Solution structure of the ribosome Elongation Factor P (EF-P) from Staphylococcus aureus | | 分子名称: | Elongation factor P | | 著者 | Usachev, K, Fatkhullin, B, Gabdulkhakov, A, Khusainov, I, Golubev, A, Validov, S, Yusupova, G, Yusupov, M. | | 登録日 | 2019-04-30 | | 公開日 | 2020-03-11 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | NMR and crystallographic structural studies of the Elongation factor P from Staphylococcus aureus.
Eur.Biophys.J., 49, 2020
|
|
6RLS
 
 | | Concerted dynamics of metallo-base pairs in an A/B-form helical transition (apo species) | | 分子名称: | DNA (5'-D(*CP*GP*TP*CP*TP*CP*AP*TP*GP*AP*TP*AP*CP*G)-3')_apo | | 著者 | Schmidt, O.P, Jurt, S, Johannsen, S, Karimi, A, Sigel, R.K.O, Luedtke, N.W. | | 登録日 | 2019-05-02 | | 公開日 | 2019-10-09 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Concerted dynamics of metallo-base pairs in an A/B-form helical transition.
Nat Commun, 10, 2019
|
|
6RPV
 
 | | Extremely stable monomeric variant of human cystatin C with single amino acid substitution | | 分子名称: | Cystatin-C | | 著者 | Zhukov, I, Rodziewicz-Motowidlo, S, Maszota-Zieleniak, M, Jurczak, P, Kozak, M. | | 登録日 | 2019-05-14 | | 公開日 | 2019-07-31 | | 最終更新日 | 2023-06-14 | | 実験手法 | SOLUTION NMR, SOLUTION SCATTERING | | 主引用文献 | NMR and crystallographic structural studies of the extremely stable monomeric variant of human cystatin C with single amino acid substitution.
Febs J., 287, 2020
|
|
6RQS
 | | RW16 peptide | | 分子名称: | ARG-ARG-TRP-ARG-ARG-TRP-TRP-ARG-ARG-TRP-TRP-ARG-ARG-TRP-ARG-ARG | | 著者 | Jobin, M.-L, Grelard, A, Alves, I, Mackereth, C.D. | | 登録日 | 2019-05-16 | | 公開日 | 2019-10-23 | | 最終更新日 | 2024-06-19 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Biophysical Insight on the Membrane Insertion of an Arginine-Rich Cell-Penetrating Peptide.
Int J Mol Sci, 20, 2019
|
|
6RRL
 
 | |
6RRO
 
 | |
6RS3
 | |
6RSF
 | |
6RSG
 | |
6RSM
 | | Solution NMR structure of the peptide 12530 from medicinal leech Hirudo medicinalis in dodecylphosphocholine micelles | | 分子名称: | peptide 12530 | | 著者 | Talyzina, I.A, Nadezhdin, K.D, Grafskaia, E.N, Arseniev, A.S, Lazarev, V.N. | | 登録日 | 2019-05-21 | | 公開日 | 2019-07-24 | | 最終更新日 | 2024-06-19 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Medicinal leech antimicrobial peptides lacking toxicity represent a promising alternative strategy to combat antibiotic-resistant pathogens.
Eur.J.Med.Chem., 180, 2019
|
|
6RSS
 
 | | Solution structure of the fourth WW domain of WWP2 with GB1-tag | | 分子名称: | NEDD4-like E3 ubiquitin-protein ligase WWP2 | | 著者 | Wahl, L.C, Watt, J.E, Tolchard, J, Blumenschein, T.M.A, Chantry, A. | | 登録日 | 2019-05-22 | | 公開日 | 2019-10-09 | | 最終更新日 | 2024-06-19 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Smad7 Binds Differently to Individual and Tandem WW3 and WW4 Domains of WWP2 Ubiquitin Ligase Isoforms.
Int J Mol Sci, 20, 2019
|
|
6RVA
 
 | | STRUCTURE OF [ASP58]-IGF-I ANALOGUE | | 分子名称: | Insulin-like growth factor I | | 著者 | Jiracek, J, Zakova, L, Socha, O. | | 登録日 | 2019-05-31 | | 公開日 | 2019-10-02 | | 最終更新日 | 2023-06-14 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Mutations at hypothetical binding site 2 in insulin and insulin-like growth factors 1 and 2 result in receptor- and hormone-specific responses.
J.Biol.Chem., 294, 2019
|
|
6RWG
 
 | | Structure of HIV-1 CAcSP1NC mutant(W41A,M42A) interacting with maturation inhibitor EP39 | | 分子名称: | Gag polyprotein, ZINC ION | | 著者 | Chen, X, Coric, P, Larue, V, Nonin-Lecomte, S, Bouaziz, S, Structural Genomics Consortium (SGC) | | 登録日 | 2019-06-05 | | 公開日 | 2020-05-20 | | 最終更新日 | 2024-06-19 | | 実験手法 | SOLUTION NMR | | 主引用文献 | The HIV-1 maturation inhibitor, EP39, interferes with the dynamic helix-coil equilibrium of the CA-SP1 junction of Gag.
Eur.J.Med.Chem., 204, 2020
|
|
6RY9
 | |
6RYQ
 
 | |
6RZ1
 
 | |
6RZC
 
 | |
6S0N
 
 | | A9 peptide derived from Herceptin fab binding region | | 分子名称: | GLN-ASP-VAL-ASN-THR-ALA-VAL-ALA-TRP | | 著者 | De Luca, S, Verdoliva, V, Saviano, M, Fattorusso, R, Diana, D. | | 登録日 | 2019-06-17 | | 公開日 | 2019-11-06 | | 最終更新日 | 2024-06-19 | | 実験手法 | SOLUTION NMR | | 主引用文献 | SPR and NMR characterization of the molecular interaction between A9 peptide and a model system of HER2 receptor: A fragment approach for selecting peptide structures specific for their target.
J.Pept.Sci., 26, 2020
|
|
