[English] 日本語
 Yorodumi
Yorodumi- PDB-1qnh: Plasmodium falciparum Cyclophilin (double mutant) complexed with ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qnh | ||||||
|---|---|---|---|---|---|---|---|
| Title | Plasmodium falciparum Cyclophilin (double mutant) complexed with Cyclosporin A | ||||||
 Components Components |
| ||||||
 Keywords Keywords | ISOMERASE/IMMUNOSUPPRESSANT / ISOMERASE-IMMUNOSUPPRESSANT COMPLEX / CYCLOPHILIN-CYCLOSPORIN COMPLEX / CYCLOSPORIN A / IMMUNOSUPPRESSANT / CYCLOPHILIN | ||||||
| Function / homology |  Function and homology information Function and homology informationcyclosporin A binding / peptidylprolyl isomerase / peptidyl-prolyl cis-trans isomerase activity / protein folding / cytoplasm Similarity search - Function | ||||||
| Biological species |   TOLYPOCLADIUM INFLATUM (fungus) TOLYPOCLADIUM INFLATUM (fungus) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.1 Å MOLECULAR REPLACEMENT / Resolution: 2.1 Å | ||||||
 Authors Authors | Peterson, M.R. / Hall, D.R. / Hunter, W.N. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2000 Journal: J.Mol.Biol. / Year: 2000Title: The Three-Dimensional Structure of a Plasmodium Falciparum Cyclophilin in Complex with the Potent Anti-Malarial Cyclosporin A Authors: Peterson, M.R. / Hall, D.R. / Berriman, M. / Leonard, G.A. / Fairlamb, A.H. / Hunter, W.N. #1: Journal: Biochem.J. / Year: 1998 Title: Detailed Characterization of a Cyclophilin from the Human Malaria Parasite Plasmodium Falciparum Authors: Berriman, M. / Fairlamb, A.H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qnh.cif.gz 1qnh.cif.gz | 88.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qnh.ent.gz pdb1qnh.ent.gz | 67.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qnh.json.gz 1qnh.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qn/1qnh https://data.pdbj.org/pub/pdb/validation_reports/qn/1qnh ftp://data.pdbj.org/pub/pdb/validation_reports/qn/1qnh ftp://data.pdbj.org/pub/pdb/validation_reports/qn/1qnh | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1qngC 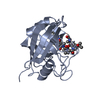 1cwaS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (-0.16335, -0.98606, 0.03182), Vector: |
- Components
Components
| #1: Protein | Mass: 18884.408 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Production host:  #2: Protein/peptide | 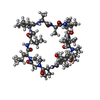  Type: Cyclic peptide / Class: Immunosuppressant / Mass: 1220.625 Da / Num. of mol.: 2 / Source method: obtained synthetically Type: Cyclic peptide / Class: Immunosuppressant / Mass: 1220.625 Da / Num. of mol.: 2 / Source method: obtained syntheticallyDetails: CYCLOSPORIN IS A CYCLIC UNDECAPEPTIDE. CYCLIZATION IS ACHIEVED BY LINKING THE N- AND THE C- TERMINI. Source: (synth.)  TOLYPOCLADIUM INFLATUM (fungus) / References: NOR: NOR00033, Cyclosporin A TOLYPOCLADIUM INFLATUM (fungus) / References: NOR: NOR00033, Cyclosporin A#3: Water | ChemComp-HOH / | Compound details | CYCLOSPORIN IS A CYCLIC UNDECAPEPTIDE. HERE, CYCLOSPORIN A IS REPRESENTED BY THE SEQUENCE (SEQRES) ...CYCLOSPORI | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 4 X-RAY DIFFRACTION / Number of used crystals: 4 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.74 Å3/Da / Density % sol: 55 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 7.7 / Details: pH 7.70 | |||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 277 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: BM14 / Wavelength: 0.98 / Beamline: BM14 / Wavelength: 0.98 |
| Detector | Type: FUJI BAS-2000 IMAGE PLATE / Detector: IMAGE PLATE |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.98 Å / Relative weight: 1 |
| Reflection | Resolution: 2.1→24 Å / Num. obs: 24606 / % possible obs: 98.2 % / Redundancy: 5 % / Biso Wilson estimate: 34.2 Å2 / Rmerge(I) obs: 0.076 / Net I/σ(I): 17.8 |
| Reflection shell | Resolution: 2.1→2.15 Å / Redundancy: 2 % / Rmerge(I) obs: 0.213 / Mean I/σ(I) obs: 3.7 / % possible all: 92.7 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1CWA Resolution: 2.1→24 Å / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 Details: REFINEMENT AT EARLY STAGES UTILISED X-PLOR AND REFMAC AND NCS RESTRAINTS WERE RELEASED AT THE FINAL STAGES OF REFINEMENT THE SIDE CHAINS OF THE FOLLOWING RESIDUES ARE DISORDERED ASN A 15, ...Details: REFINEMENT AT EARLY STAGES UTILISED X-PLOR AND REFMAC AND NCS RESTRAINTS WERE RELEASED AT THE FINAL STAGES OF REFINEMENT THE SIDE CHAINS OF THE FOLLOWING RESIDUES ARE DISORDERED ASN A 15, LYS A 51, LYS A 155, ARG B 49, LYS B 51, ARG B 88, LYS B 155, LYS B 160 SER A 2 IS DISORDERED AND NOT OBSERVED
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 26.6 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→24 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.1→2.11 Å / Total num. of bins used: 49
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller






































 PDBj
PDBj


