[English] 日本語
 Yorodumi
Yorodumi- EMDB-10205: Molecular structure of mouse apoferritin resolved at 2.7 Angstrom... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10205 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Molecular structure of mouse apoferritin resolved at 2.7 Angstroms with the Glacios cryo-microscope | |||||||||
 Map data Map data | Post-processed map of apoferritin at 200 kV resolved with the FEI Glacios microscope at 2.7 A | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Apoferritin / iron binding / iron storing / complex / METAL BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationIron uptake and transport / Golgi Associated Vesicle Biogenesis / ferroxidase / autolysosome / negative regulation of ferroptosis / ferroxidase activity / negative regulation of fibroblast proliferation / endocytic vesicle lumen / Neutrophil degranulation / ferric iron binding ...Iron uptake and transport / Golgi Associated Vesicle Biogenesis / ferroxidase / autolysosome / negative regulation of ferroptosis / ferroxidase activity / negative regulation of fibroblast proliferation / endocytic vesicle lumen / Neutrophil degranulation / ferric iron binding / autophagosome / iron ion transport / ferrous iron binding / intracellular iron ion homeostasis / immune response / iron ion binding / negative regulation of cell population proliferation / mitochondrion / extracellular region / membrane / identical protein binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.73 Å | |||||||||
 Authors Authors | Hamdi F / Tueting C | |||||||||
 Citation Citation |  Journal: PLoS One / Year: 2020 Journal: PLoS One / Year: 2020Title: 2.7 Å cryo-EM structure of vitrified M. musculus H-chain apoferritin from a compact 200 keV cryo-microscope. Authors: Farzad Hamdi / Christian Tüting / Dmitry A Semchonok / Koen M Visscher / Fotis L Kyrilis / Annette Meister / Ioannis Skalidis / Lisa Schmidt / Christoph Parthier / Milton T Stubbs / Panagiotis L Kastritis /   Abstract: Here we present the structure of mouse H-chain apoferritin at 2.7 Å (FSC = 0.143) solved by single particle cryogenic electron microscopy (cryo-EM) using a 200 kV device, the Thermo Fisher Glacios®. ...Here we present the structure of mouse H-chain apoferritin at 2.7 Å (FSC = 0.143) solved by single particle cryogenic electron microscopy (cryo-EM) using a 200 kV device, the Thermo Fisher Glacios®. This is a compact, two-lens illumination system with a constant power objective lens, without any energy filters or aberration correctors, often thought of as a "screening cryo-microscope". Coulomb potential maps reveal clear densities for main chain carbonyl oxygens, residue side chains (including alternative conformations) and bound solvent molecules. We used a quasi-crystallographic reciprocal space approach to fit model coordinates to the experimental cryo-EM map. We argue that the advantages offered by (a) the high electronic and mechanical stability of the microscope, (b) the high emission stability and low beam energy spread of the high brightness Field Emission Gun (X-FEG), (c) direct electron detection technology and (d) particle-based Contrast Transfer Function (CTF) refinement have contributed to achieving high resolution. Overall, we show that basic electron optical settings for automated cryo-electron microscopy imaging can be used to determine structures approaching atomic resolution. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10205.map.gz emd_10205.map.gz | 8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10205-v30.xml emd-10205-v30.xml emd-10205.xml emd-10205.xml | 25.8 KB 25.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10205_fsc.xml emd_10205_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_10205.png emd_10205.png | 103.4 KB | ||
| Masks |  emd_10205_msk_1.map emd_10205_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-10205.cif.gz emd-10205.cif.gz | 6.2 KB | ||
| Others |  emd_10205_additional.map.gz emd_10205_additional.map.gz emd_10205_half_map_1.map.gz emd_10205_half_map_1.map.gz emd_10205_half_map_2.map.gz emd_10205_half_map_2.map.gz | 59.8 MB 45.9 MB 45.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10205 http://ftp.pdbj.org/pub/emdb/structures/EMD-10205 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10205 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10205 | HTTPS FTP |
-Related structure data
| Related structure data |  6shtMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10205.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10205.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed map of apoferritin at 200 kV resolved with the FEI Glacios microscope at 2.7 A | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.96 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10205_msk_1.map emd_10205_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
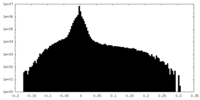
| Density Histograms |
-Additional map: Post-processed (unmasked) map of apoferritin at 200 kV...
| File | emd_10205_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed (unmasked) map of apoferritin at 200 kV resolved with the FEI Glacios microscope at 2.7 A | ||||||||||||
| Projections & Slices |
| ||||||||||||
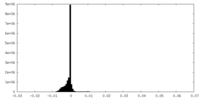
| Density Histograms |
-Half map: Refined half-map of apoferritin at 200 kV resolved...
| File | emd_10205_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Refined half-map of apoferritin at 200 kV resolved with the FEI Glacios microscope | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Refined half-map of apoferritin at 200 kV resolved...
| File | emd_10205_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Refined half-map of apoferritin at 200 kV resolved with the FEI Glacios microscope | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Mouse apoferritin
| Entire | Name: Mouse apoferritin |
|---|---|
| Components |
|
-Supramolecule #1: Mouse apoferritin
| Supramolecule | Name: Mouse apoferritin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 480 KDa |
-Macromolecule #1: Ferritin heavy chain
| Macromolecule | Name: Ferritin heavy chain / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: ferroxidase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21.097631 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTTASPSQVR QNYHQDAEAA INRQINLELY ASYVYLSMSC YFDRDDVALK NFAKYFLHQS HEEREHAEKL MKLQNQRGGR IFLQDIKKP DRDDWESGLN AMECALHLEK SVNQSLLELH KLATDKNDPH LCDFIETYYL SEQVKSIKEL GDHVTNLRKM G APEAGMAE YLFDKHTLGH GDES UniProtKB: Ferritin heavy chain |
-Macromolecule #2: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 2 / Number of copies: 1 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 72 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4.0 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Details | microscope model is Thermofisher Glacios 200 kV |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 300 / Average exposure time: 30.0 sec. / Average electron dose: 28.0 e/Å2 / Details: pixel size was 0.96 Angstroms |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 1.6 µm / Calibrated defocus min: 0.8 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)