-検索条件
-検索結果
検索 (著者・登録者: yap & m)の結果248件中、1から50件目までを表示しています

EMDB-41433: 
Escherichia coli RNA polymerase unwinding intermediate (I1a) at the lambda PR promoter

EMDB-41437: 
Escherichia coli RNA polymerase unwinding intermediate (I1d) at the lambda PR promoter

EMDB-41439: 
Escherichia coli RNA polymerase unwinding intermediate (I1b) at the lambda PR promoter

EMDB-41448: 
Escherichia coli RNA polymerase unwinding intermediate (I1c) at the lambda PR promoter

EMDB-41456: 
Escherichia coli RNA polymerase closed complex intermediate at the lambda PR promoter

PDB-8to1: 
Escherichia coli RNA polymerase unwinding intermediate (I1a) at the lambda PR promoter

PDB-8to6: 
Escherichia coli RNA polymerase unwinding intermediate (I1d) at the lambda PR promoter

PDB-8to8: 
Escherichia coli RNA polymerase unwinding intermediate (I1b) at the lambda PR promoter

PDB-8toe: 
Escherichia coli RNA polymerase unwinding intermediate (I1c) at the lambda PR promoter

PDB-8tom: 
Escherichia coli RNA polymerase closed complex intermediate at the lambda PR promoter

EMDB-50218: 
Negative staining EM map for Mis18 core complex

EMDB-50219: 
Negative staining EM map for Mis18 core complex

EMDB-50220: 
Negative staining EM map for Mis18 core complex

EMDB-37320: 
CryoEM structure of NaDC1 with Citrate

EMDB-37321: 
CryoEM structure of NaDC1 in apo state

EMDB-37322: 
NaDC1 with inhibitor ACA

EMDB-37323: 
NaS1 with sulfate - IN/IN state

EMDB-37329: 
NaS1 with sulfate in IN/OUT state

EMDB-37330: 
NaS1 in IN/IN state
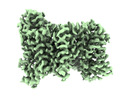
EMDB-37332: 
NaS1 in IN/OUT state

PDB-8w6c: 
CryoEM structure of NaDC1 with Citrate

PDB-8w6d: 
CryoEM structure of NaDC1 in apo state

PDB-8w6g: 
NaDC1 with inhibitor ACA

PDB-8w6h: 
NaS1 with sulfate - IN/IN state

PDB-8w6n: 
NaS1 with sulfate in IN/OUT state

PDB-8w6o: 
NaS1 in IN/IN state

PDB-8w6t: 
NaS1 in IN/OUT state

EMDB-29907: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v); consensus map with only Fab 1G01 resolved

EMDB-29908: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v), locally refined map

EMDB-29909: 
Structure of human NDS.3 Fab in complex with influenza virus neuraminidase from A/Darwin/09/2021 (H3N2)

PDB-8gat: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v), based on consensus cryo-EM map with only Fab 1G01 resolved

PDB-8gau: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v)

PDB-8gav: 
Structure of human NDS.3 Fab in complex with influenza virus neuraminidase from A/Darwin/09/2021 (H3N2)
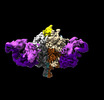
EMDB-41048: 
Lassa GPC Trimer in complex with Fab 8.11G and nanobody D5

PDB-8t5c: 
Lassa GPC Trimer in complex with Fab 8.11G and nanobody D5

EMDB-35254: 
ACE2-SIT1 complex bound with proline

EMDB-35255: 
ACE2-B0AT1 complex bound with glutamine
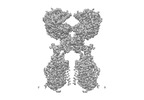
EMDB-35256: 
ACE2-B0AT1 complex bound with methionine

EMDB-35260: 
Cryo-EM map of the ACE2-SIT1 complex bound with proline, focused refined on extracellular region

EMDB-35261: 
cryo-EM map of the ACE2-SIT1 complex bound with proline, focused refined on transmembrane region

EMDB-35262: 
cryo-EM map of the ACE2-B0AT1 complex bound with glutamine, focused refined on extracellular region

EMDB-35265: 
cryo-EM map of the ACE2-B0AT1 complex bound with glutamine, focused refined on transmembrane region
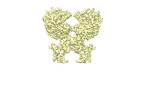
EMDB-35271: 
cryo-EM map of the ACE2-B0AT1 complex bound with methionine, focused refined on extracellular region
EMDB-35273: 
cryo-EM map of the ACE2-B0AT1 complex bound with methionine, focused refined on transmembrane region

PDB-8i91: 
ACE2-SIT1 complex bound with proline

PDB-8i92: 
ACE2-B0AT1 complex bound with glutamine

PDB-8i93: 
ACE2-B0AT1 complex bound with methionine

EMDB-41302: 
Lassa GPC trimer in complex with Fab GP23

EMDB-33650: 
SARS-CoV-2 spike glycoprotein trimer complexed with Fab fragment of anti-RBD antibody E7

EMDB-33651: 
SARS-CoV-2 spike glycoprotein trimer complexed with Fab fragment of anti-RBD antibody E7 (focused refinement on Fab-RBD interface)
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
