-Search query
-Search result
Showing 1 - 50 of 97 items for (author: tian & lr)

EMDB-64823: 
PSI-LHCE supercomplex from Euglena gracilis
Method: single particle / : Bai TY, Mao ZY, Tian LR

EMDB-64824: 
PSI-LHCE supercomplex from Euglena gracilis.
Method: single particle / : Bai TY, Mao ZY, Tian LR

EMDB-72524: 
Structure of the Omicron Spike RBD bound by the monobody s19382 (local refinement from dimerized Spike protein ECDs)
Method: single particle / : Noland CL, Perez CP, Huang P

PDB-9y5y: 
Structure of the Omicron Spike RBD bound by the monobody s19382 (local refinement from dimerized Spike protein ECDs)
Method: single particle / : Noland CL, Perez CP, Huang P

EMDB-19692: 
Hexameric worm glutamate dehydrogenase (N-term. deletion 1-33)
Method: single particle / : Bohnacker S, Bohn S, Sattler M, Esser-von Bieren J

EMDB-19693: 
Hexameric worm glutamate dehydrogenase (C136S)
Method: single particle / : Bohnacker S, Bohn S, Sattler M, Esser-von Bieren J

EMDB-18456: 
CryoEM map of hexamer worm glutamate dehydrogenase
Method: single particle / : Bohnacker S, Bohn S, Sattler M, Esser-von Bieren J

EMDB-41156: 
HCMV Trimer in complex with CS2it1p2_F7K Fab and CS4tt1p1_E3K Fab
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-41157: 
Global reconstruction for HCMV Trimer in complex with CS2it1p2_F7K Fab and CS4tt1p1_E3K Fab
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-41158: 
CS2it1p2_F7K local refinement for HCMV Trimer in complex with CS2it1p2_F7K Fab and CS4tt1p1_E3K Fab
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-41160: 
CS4tt1p1_E3K local refinement for HCMV Trimer in complex with CS2it1p2_F7K Fab and CS4tt1p1_E3K Fab
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-41161: 
gH base local refinement for HCMV Trimer in complex with CS2it1p2_F7K Fab and CS4tt1p1_E3K Fab
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-41179: 
HCMV Pentamer in complex with CS2pt1p2_A10L Fab and CS3pt1p4_C1L Fab
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-41180: 
Global reconstruction for HCMV Pentamer in complex with CS2pt1p2_A10L Fab and CS3pt1p4_C1L Fab
Method: single particle / : Goldsmith JG, McLellan JS

PDB-8tco: 
HCMV Trimer in complex with CS2it1p2_F7K Fab and CS4tt1p1_E3K Fab
Method: single particle / : Goldsmith JA, McLellan JS

PDB-8tea: 
HCMV Pentamer in complex with CS2pt1p2_A10L Fab and CS3pt1p4_C1L Fab
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-16847: 
CryoEM structure of holo e4D2
Method: single particle / : Yadav KNS, Hutchins G, Berger Schaffitzel C, Anderson R

EMDB-15244: 
Tomogram of an Ebola VLP composed of GP, VP40, NP, VP24 and VP35 at pH 7.4 (Figure 1A-D)
Method: electron tomography / : Winter SL, Chlanda P

EMDB-15268: 
Tomogram of an Ebola VLP composed of VP40 at pH 4.5 (Figure 1J)
Method: electron tomography / : Winter SL, Chlanda P
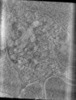
EMDB-15951: 
Tomogram of an EBOV-infected Huh7 cell showing a late endosome with internalized EBOV particles
Method: electron tomography / : Winter SL, Chlanda P

EMDB-15956: 
Tomogram of an extracellular EBOV particle adjacent to an EBOV-infected Huh7 cell
Method: electron tomography / : Winter SL, Chlanda P

EMDB-16010: 
Cryo-EM structure of SARS-CoV-2 spike (HexaPro variant) in complex with nanobody W25 (map 3, focus refinement on RBD, W25 and adjacent NTD)
Method: single particle / : Lauer S, Spahn CMT, Schwefel D

EMDB-16030: 
Cryo-EM structure of SARS-CoV-2 spike (Omicron BA.1 variant) in complex with nanobody W25 (map 5, focus refinement on RBD, W25 and adjacent NTD)
Method: single particle / : Modhiran N, Lauer S, Spahn CMT, Watterson D, Schwefel D

PDB-8bev: 
Cryo-EM structure of SARS-CoV-2 spike (HexaPro variant) in complex with nanobody W25 (map 3, focus refinement on RBD, W25 and adjacent NTD)
Method: single particle / : Lauer S, Spahn CMT, Schwefel D

PDB-8bgg: 
Cryo-EM structure of SARS-CoV-2 spike (Omicron BA.1 variant) in complex with nanobody W25 (map 5, focus refinement on RBD, W25 and adjacent NTD)
Method: single particle / : Modhiran N, Lauer S, Spahn CMT, Watterson D, Schwefel D

EMDB-33233: 
Cryo-EM structure of EDS1 and SAG101 with ATP-APDR
Method: single particle / : Huang SJ, Jia AL, Han ZF, Chai JJ

PDB-7xjp: 
Cryo-EM structure of EDS1 and SAG101 with ATP-APDR
Method: single particle / : Huang SJ, Jia AL, Han ZF, Chai JJ

EMDB-27095: 
Cryo-EM structure of BCL10 R58Q filament
Method: helical / : David L, Wu H

EMDB-27100: 
Cryo-EM structure of BCL10 CARD - MALT1 DD filament
Method: helical / : David L, Wu H

EMDB-32216: 
26S proteasome from the cell with TDP-25 inclusion
Method: subtomogram averaging / : Guo Q

EMDB-32217: 
tomogram of a rat primary neuron harboring TDP-25 inclusion
Method: electron tomography / : Guo Q

EMDB-13105: 
SARS-CoV-2 S (Spike) protein in complex with VHH Re5D06 (multi-body-refined RBD-Re5D06 body)
Method: single particle / : Guttler T, Aksu M, Dienemann C, Gorlich D

EMDB-13106: 
SARS-CoV-2 S (Spike) protein in complex with VHH Re5D06 (one RBD down)
Method: single particle / : Guttler T, Aksu M, Dienemann C, Gorlich D

EMDB-13107: 
SARS-CoV-2 S (Spike) protein in complex with VHH Re5D06 (all RBDs up)
Method: single particle / : Guttler T, Aksu M, Dienemann C, Gorlich D

EMDB-12007: 
Programmable icosahedral shell system for virus trapping: DNA-origami half octahedron
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12008: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=9 triangle (pentamer)
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12009: 
Programmable icosahedral shell system for virus trapping: DNA-origami octahedron triangle
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12010: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=1 triangle
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12011: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=3 triangle
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12012: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=4 triangle (isosceles)
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12013: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=4 triangle (equilateral)
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12014: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=9 triangle (hexamer 1)
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12015: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=9 triangle (hexamer 2)
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12016: 
Programmable icosahedral shell system for virus trapping: DNA-origami octahedron shell
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12019: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=3 shell
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12020: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=4 shell
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12021: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=1 shell
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H

EMDB-12022: 
Programmable icosahedral shell system for virus trapping: T=1 shell trapping a HBV core particle
Method: single particle / : Sigl C, Willner EM, Engelen W, Kretzmann JA, Sachenbacher K, Liedl A, Kolbe F, Wilsch F, Aghvami SA, Protzer U, Hagan MF, Fraden S, Dietz H
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
