-検索条件
-検索結果
検索 (著者・登録者: thomas & jm)の結果164件中、1から50件目までを表示しています

EMDB-18779: 
Structure of the non-mitochondrial citrate synthase from Ananas comosus

EMDB-18990: 
CryoEM map of tau PHF sarkosyl-extracted from a human AD patient (associated with in situ tomography)

EMDB-50148: 
Tau PHF subtomogram average relating to CS1 extended data Figure 9A

EMDB-50152: 
Tau PHF subtomogram average relating to CS2 Figure 3i-j.

EMDB-50153: 
Tau PHF subtomogram average relating to CS3 extended data Figure 9c

EMDB-50155: 
Tau PHF subtomogram average relating to CS4 extended data Figure 9d

EMDB-50156: 
Tau PHF subtomogram average relating to CS5 extended data Figure 9b

EMDB-50157: 
Tau PHF subtomogram average relating to CS6 extended data Figure 9e

EMDB-50159: 
Tau PHF subtomogram average relating to CS7 extended data Figure 9f

EMDB-50160: 
Tau PHF subtomogram average relating to LOL1_PHF Figure 4g-h

EMDB-50161: 
Tau SF subtomogram average relating to LOL1_SF Figure 4g-h

EMDB-50162: 
Tau SF subtomogram average relating to LOL2_SF Figure 4i-j

EMDB-41583: 
Actin 1 from T. gondii in filaments bound to MgADP

EMDB-41584: 
Actin 1 from T. gondii in filaments bound to MgADP and jasplakinolide

EMDB-19250: 
Pseudoatomic model of a second-order Sierpinski triangle formed by the citrate synthase from Synechococcus elongatus

EMDB-19251: 
Structure of a first order Sierpinski triangle formed by the H369R mutant of the citrate synthase from Synechococcus elongatus

EMDB-41374: 
Antibody N3-1 bound to RBDs in the up and down conformations
EMDB-41382: 
Antibody N3-1 bound to RBD in the up conformation
EMDB-41399: 
Antibody N3-1 bound to SARS-CoV-2 spike

EMDB-42124: 
Cryo-EM structure of human STEAP1 in complex with AMG 509 Fab

EMDB-41617: 
CryoEM structure of PI3Kalpha

PDB-8tu6: 
CryoEM structure of PI3Kalpha

EMDB-29281: 
Cryo-EM structure of STING oligomer bound to cGAMP and NVS-STG2

EMDB-29282: 
Cryo-EM structure of STING oligomer bound to cGAMP, NVS-STG2 and C53

EMDB-40789: 
BAP1/ASXL1 bound to the H2AK119Ub Nucleosome

EMDB-40790: 
Map focused on acidic patch BAP1/ASXL1 bound to the H2AK119Ub Nucleosome

EMDB-40791: 
Overall map of BAP1/ASXL1 bound to the H2AK119Ub Nucleosome

EMDB-29930: 
T. cruzi topoisomerase II alpha bound to dsDNA and the covalent inhibitor CT1

EMDB-28178: 
Structure of lineage IV Lassa virus glycoprotein complex (strain Josiah)

EMDB-28179: 
Structure of lineage II Lassa virus glycoprotein complex (strain NIG08-A41)

EMDB-28180: 
Structure of lineage V Lassa virus glycoprotein complex (strain Soromba-R)

EMDB-28181: 
Structure of lineage VII Lassa virus glycoprotein complex (strain Togo/2016/7082)

EMDB-28182: 
Lassa virus glycoprotein complex (Josiah) bound to 12.1F Fab

EMDB-28183: 
Lassa virus glycoprotein complex (Josiah) bound to 19.7E Fab

EMDB-28184: 
Lassa virus glycoprotein complex (Josiah) bound to S370.7 Fab
EMDB-14842: 
Cryo-EM structure of the canine distemper virus tetrameric attachment glycoprotein

EMDB-15636: 
Human 80S ribosome structure from pFIB-lamellae

EMDB-16185: 
80S human ribosome structure from PFIB lamellae of HeLa cells for assessing the extend and depth of the damage layer: 15 to 30 nm

EMDB-16186: 
80S human ribosome structure from PFIB lamellae of HeLa cells for assessing the extend and depth of the damage layer: above 30 nm matched control (for 15 to 30 nm)

EMDB-16192: 
80S human ribosome structure from PFIB lamellae of HeLa cells for assessing the extend and depth of the damage layer:30 to 45 nm

EMDB-16193: 
80S human ribosome structure from PFIB lamellae of HeLa cells for assessing the extend and depth of the damage layer: above 45 nm matched control (for 30 to 45 nm)

EMDB-16194: 
80S human ribosome structure from PFIB lamellae of HeLa cells for assessing the extend and depth of the damage layer:45 to 60 nm

EMDB-16195: 
80S human ribosome structure from PFIB lamellae of HeLa cells for assessing the extend and depth of the damage layer: above 60 nm matched control (for 45 to 60 nm)

EMDB-16196: 
80S human ribosome structure from PFIB lamellae of HeLa cells for assessing the extend and depth of the damage layer: 0 to 15 nm

EMDB-16199: 
80S human ribosome structure from PFIB lamellae of HeLa cells for assessing the extend and depth of the damage layer: above 15 nm matched control (for 0 to 15 nm)
EMDB-28523: 
Structure of interleukin receptor common gamma chain (IL2Rgamma) in complex with two antibodies
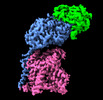
EMDB-15210: 
Human 4F2hc-LAT2 heterodimeric amino acid transporter in complex with anticalin D11vs

EMDB-25107: 
Lassa virus glycoprotein construct(Josiah GPCysR4) recovered from GPC-I53-50 nanoparticle by localized reconstruction

EMDB-25108: 
I53-50 nanoparticle core reconstructed from GPC-I53-50NP by focused refinement

EMDB-25109: 
Lassa virus glycoprotein construct (Josiah GPC-I53-50A) in complex with LAVA01 antibody
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
