[English] 日本語
 Yorodumi
Yorodumi- EMDB-25108: I53-50 nanoparticle core reconstructed from GPC-I53-50NP by focus... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | I53-50 nanoparticle core reconstructed from GPC-I53-50NP by focused refinement | |||||||||
 Map data Map data | I53-50 nanoparticle map reconstructed from GPC-I53-50 nanoparticle by focused refinement - Main map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | glycoprotein / Lassa / Josiah / vaccine design / nanoparticle / protein design / VIRAL PROTEIN | |||||||||
| Biological species | synthetic construct (others) | |||||||||
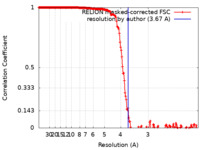
| Method | single particle reconstruction / cryo EM / Resolution: 3.67 Å | |||||||||
 Authors Authors | Antanasijevic A / Brouwer PJM | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Host Microbe / Year: 2022 Journal: Cell Host Microbe / Year: 2022Title: Lassa virus glycoprotein nanoparticles elicit neutralizing antibody responses and protection. Authors: Philip J M Brouwer / Aleksandar Antanasijevic / Adam J Ronk / Helena Müller-Kräuter / Yasunori Watanabe / Mathieu Claireaux / Hailee R Perrett / Tom P L Bijl / Marloes Grobben / Jeffrey C ...Authors: Philip J M Brouwer / Aleksandar Antanasijevic / Adam J Ronk / Helena Müller-Kräuter / Yasunori Watanabe / Mathieu Claireaux / Hailee R Perrett / Tom P L Bijl / Marloes Grobben / Jeffrey C Umotoy / Angela I Schriek / Judith A Burger / Khadija Tejjani / Nicole M Lloyd / Thijs H Steijaert / Marlies M van Haaren / Kwinten Sliepen / Steven W de Taeye / Marit J van Gils / Max Crispin / Thomas Strecker / Alexander Bukreyev / Andrew B Ward / Rogier W Sanders /     Abstract: The Lassa virus is endemic in parts of West Africa, and it causes hemorrhagic fever with high mortality. The development of a recombinant protein vaccine has been hampered by the instability of ...The Lassa virus is endemic in parts of West Africa, and it causes hemorrhagic fever with high mortality. The development of a recombinant protein vaccine has been hampered by the instability of soluble Lassa virus glycoprotein complex (GPC) trimers, which disassemble into monomeric subunits after expression. Here, we use two-component protein nanoparticles consisting of trimeric and pentameric subunits to stabilize GPC in a trimeric conformation. These GPC nanoparticles present twenty prefusion GPC trimers on the surface of an icosahedral particle. Cryo-EM studies of GPC nanoparticles demonstrated a well-ordered structure and yielded a high-resolution structure of an unliganded GPC. These nanoparticles induced potent humoral immune responses in rabbits and protective immunity against the lethal Lassa virus challenge in guinea pigs. Additionally, we isolated a neutralizing antibody that mapped to the putative receptor-binding site, revealing a previously undefined site of vulnerability. Collectively, these findings offer potential approaches to vaccine and therapeutic design for the Lassa virus. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25108.map.gz emd_25108.map.gz | 501.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25108-v30.xml emd-25108-v30.xml emd-25108.xml emd-25108.xml | 21.6 KB 21.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25108_fsc.xml emd_25108_fsc.xml | 18.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_25108.png emd_25108.png | 194.8 KB | ||
| Masks |  emd_25108_msk_1.map emd_25108_msk_1.map | 536.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-25108.cif.gz emd-25108.cif.gz | 6.9 KB | ||
| Others |  emd_25108_half_map_1.map.gz emd_25108_half_map_1.map.gz emd_25108_half_map_2.map.gz emd_25108_half_map_2.map.gz | 428.3 MB 428.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25108 http://ftp.pdbj.org/pub/emdb/structures/EMD-25108 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25108 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25108 | HTTPS FTP |
-Related structure data
| Related structure data |  7sgeMC  7sgdC  7sgfC C: citing same article ( M: atomic model generated by this map |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25108.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25108.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | I53-50 nanoparticle map reconstructed from GPC-I53-50 nanoparticle by focused refinement - Main map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25108_msk_1.map emd_25108_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: I53-50 nanoparticle map reconstructed from GPC-I53-50 nanoparticle by...
| File | emd_25108_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | I53-50 nanoparticle map reconstructed from GPC-I53-50 nanoparticle by focused refinement - Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: I53-50 nanoparticle map reconstructed from GPC-I53-50 nanoparticle by...
| File | emd_25108_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | I53-50 nanoparticle map reconstructed from GPC-I53-50 nanoparticle by focused refinement - Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : I53-50 nanoparticle core recovered from GPC-I53-50 nanoparticle
| Entire | Name: I53-50 nanoparticle core recovered from GPC-I53-50 nanoparticle |
|---|---|
| Components |
|
-Supramolecule #1: I53-50 nanoparticle core recovered from GPC-I53-50 nanoparticle
| Supramolecule | Name: I53-50 nanoparticle core recovered from GPC-I53-50 nanoparticle type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Nanoparticle consists of 60 monomeric I53-50A and 60 monomeric I53-50B building blocks. Nanoparticle was assembled by combining equimolar amounts of GPC-I53-50A and I53-50B. |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Macromolecule #1: Josiah GPCysR4-I53-50A - nanoparticle component
| Macromolecule | Name: Josiah GPCysR4-I53-50A - nanoparticle component / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 73.84557 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGQIVTFFQE VPHVIEEVMN IVLIALSVLA VLKGLYNFAT CGLVGLVTFL LLCGRSCTTS LYKGVYELQT LELNMETLNM TMPLSCTKN NSHHYIMVGN ETGLELTLTN TSIINHKFCN LSDAHKKNLY DHALMSIIST FHLSIPNFNQ YEAMSCDFNG G KISVQYNL ...String: MGQIVTFFQE VPHVIEEVMN IVLIALSVLA VLKGLYNFAT CGLVGLVTFL LLCGRSCTTS LYKGVYELQT LELNMETLNM TMPLSCTKN NSHHYIMVGN ETGLELTLTN TSIINHKFCN LSDAHKKNLY DHALMSIIST FHLSIPNFNQ YEAMSCDFNG G KISVQYNL SHSYAGDAAN HCGTVANGVL QTFMRMAWGG SYIALDSGCG NWDCIMTSYQ YLIIQNTTWE DHCQFSRPSP IG YLGLLSQ RTRDIYISRR RRGTFTWTLS DSEGKDTPGG YCLTRWMLIE AELKCFGNTA VAKCNEKHDE EFCDMLRLFD FNK QAIQRL KAPAQMSIQL INKAVNALIN DQLIMKNHLR DIMCIPYCNY SKYWYLNHTT TGRTSLPKCW LVSNGSYLNE THFS DDIEQ QADNMITEML QKEYMERQGG SGGSGGSGGS GGSEKAAKAE EAARKMEELF KKHKIVAVLR ANSVEEAIEK AVAVF AGGV HLIEITFTVP DADTVIKALS VLKEKGAIIG AGTVTSVEQC RKAVESGAEF IVSPHLDEEI SQFCKEKGVF YMPGVM TPT ELVKAMKLGH DILKLFPGEV VGPEFVKAMK GPFPNVKFVP TGGVDLDNVC EWFDAGVLAV GVGDALVEGD PDEVREK AK EFVEKIRGCT EGSLEWSHPQ FEK |
-Macromolecule #2: I53-50B component
| Macromolecule | Name: I53-50B component / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 18.236877 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNQHSHKDHE TVRIAVVRAR WHAEIVDACV SAFEAAMRDI GGDRFAVDVF DVPGAYEIPL HARTLAETGR YGAVLGTAFV VNGGIYRHE FVASAVINGM MNVQLNTGVP VLSAVLTPHN YDKSKAHTLL FLALFAVKGM EAARACVEIL AAREKIAAGS L EHHHHHH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7 Component:
Details: TBS | |||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 7 sec. / Pretreatment - Atmosphere: OTHER | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV / Details: Blot time 4s, Wait time 10s, Blot force 0. | |||||||||
| Details | I53-50 nanoparticle core recovered from GPC-I53-50 nanoparticle by focused refinement of the core. Nanoparticle was assembled by combining equimolar amounts of GPC-I53-50A and I53-50B. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 2 / Number real images: 2337 / Average exposure time: 10.5 sec. / Average electron dose: 50.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 29000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)