-Search query
-Search result
Showing 1 - 50 of 55 items for (author: shen & yq)
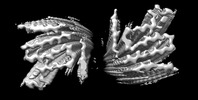
EMDB-36202: 
Cryo-EM structure of alpha-synuclein gS87 fibril

EMDB-36203: 
Cryo-EM structure of alpha-synuclein pS87 fibril

EMDB-35630: 
Cryo-EM structure of hMRS2-Mg

EMDB-35631: 
Cryo-EM structure of hMRS-highEDTA

EMDB-35632: 
Cryo-EM structure of hMRS2-lowEDTA

EMDB-35633: 
Cryo-EM structure of hMRS2-rest

EMDB-35522: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on receptor)

EMDB-35523: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on Giq-scFV16 complex)

EMDB-35524: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on receptor)

EMDB-35525: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on Gil-scFV16 complex)

EMDB-35529: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex (consensus map)

EMDB-35533: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi complex(consensus map)

EMDB-33884: 
Cryo-EM structure of Apo-alpha-syn fibril

EMDB-33890: 
Cryo-EM structure of dLAG3-alpha-syn fibril

EMDB-35356: 
Cryo-EM structure of the 9-hydroxystearic acid bound GPR120-Gi complex

EMDB-35357: 
Cryo-EM structure of the linoleic acid bound GPR120-Gi complex

EMDB-35358: 
Cryo-EM structure of the oleic acid bound GPR120-Gi complex

EMDB-35359: 
Cryo-EM structure of the TUG891 bound GPR120-Gi complex

EMDB-35360: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi complex

EMDB-29736: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex

EMDB-33934: 
Cryo-EM structure of in vitro PHF fibril

EMDB-33248: 
Cryo-EM structure of DHEA-ADGRG2-BT-Gs complex

EMDB-33249: 
Cryo-EM structure of DHEA-ADGRG2-FL-Gs complex

EMDB-33250: 
Cryo-EM structure of DHEA-ADGRG2-BT-Gs complex at lower state
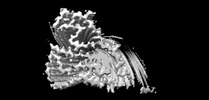
EMDB-33054: 
Cryo-EM structure of the TMEM106B fibril from normal elder

EMDB-33055: 
Cryo-EM structure of the TMEM106B fibril from Parkinson's disease dementia

EMDB-31232: 
Structural basis for the tethered peptide activation of adhesion GPCRs

EMDB-31254: 
GPR114-Gs-scFv16 complex

EMDB-30830: 
CALHM1 open state with disordered CTH

EMDB-30831: 
CALHM1 close state with disordered CTH

EMDB-30832: 
CALHM1 close state with ordered CTH

EMDB-31542: 
SARS-COV-2 Spike RBDMACSp6 binding to hACE2

EMDB-31543: 
SARS-COV-2 Spike RBDMACSp25 binding to hACE2

EMDB-31544: 
SARS-COV-2 Spike RBDMACSp36 binding to hACE2

EMDB-31546: 
SARS-COV-2 Spike RBDMACSp36 binding to mACE2

EMDB-30644: 
Structure of Calcium-Sensing Receptor in complex with NPS-2143

EMDB-30645: 
Structure of Ca2+/L-Trp-bonnd Calcium-Sensing Receptor in active state

EMDB-30646: 
Structure of Calcium-Sensing Receptor in an inactive state

EMDB-30647: 
Structure of Calcium-Sensing Receptor in complex with Evocalcet

EMDB-30503: 
Complex of SARS-CoV-2 spike trimer with its neutralizing antibody HB27

EMDB-30628: 
SARS-CoV-2 spike in complex with the CA521 neutralizing antibody (focused map)

EMDB-30629: 
SARS-CoV-2 spike in complex with the CA521 neutralizing antibody (consensus map)

EMDB-30950: 
SARS-CoV-2 spike in complex with the CA521 neutralizing antibody Fab (focused refinement on Fab-RBD)

EMDB-30951: 
SARS-CoV-2 spike in complex with the CA521 neutralizing antibody Fab (consensus map)

EMDB-30335: 
Cryo-EM structure of the flagellar LP ring from Salmonella

EMDB-30359: 
Cryo-EM structure of the flagellar motor-hook complex from Salmonella

EMDB-30602: 
Cryo-EM structure of the beclomethasone-bound adhesion receptor GPR97-Go complex

EMDB-30603: 
Cryo-EM structure of the cortisol-bound adhesion receptor GPR97-Go complex

EMDB-0983: 
Luminal ring of the Xenopus laevis nuclear pore complex

EMDB-0984: 
Luminal ring bumper-7 conformation of the Xenopus laevis nuclear pore complex
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
