-Search query
-Search result
Showing all 20 items for (author: shah & nr)

EMDB-43385: 
Cryo-EM map of close dodecameric CaMKII beta holoenzyme T287A T306A T307A

EMDB-40873: 
Cryo-EM structure of tetradecameric hub domain of CaMKII alpha

EMDB-41070: 
Cryo-EM structure of tetradecameric CaMKII beta holoenzyme T287A T306A T307A
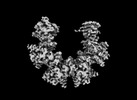
EMDB-41077: 
Cryo-EM structure of dodecameric CaMKII beta holoenzyme T287A T306A T307A

PDB-8syg: 
Cryo-EM structure of tetradecameric hub domain of CaMKII alpha

PDB-8t6k: 
Cryo-EM structure of tetradecameric CaMKII beta holoenzyme T287A T306A T307A

PDB-8t6q: 
Cryo-EM structure of dodecameric CaMKII beta holoenzyme T287A T306A T307A

EMDB-40955: 
Cryo-EM structure of dodecameric hub domain of CaMKII alpha

EMDB-40956: 
Cryo-EM structure of tetradecameric hub domain of CaMKII beta

EMDB-40957: 
Cryo-EM structure of dodecameric hub domain of CaMKII beta

PDB-8t15: 
Cryo-EM structure of dodecameric hub domain of CaMKII alpha

PDB-8t17: 
Cryo-EM structure of tetradecameric hub domain of CaMKII beta

PDB-8t18: 
Cryo-EM structure of dodecameric hub domain of CaMKII beta

EMDB-11172: 
Disulfide-locked early prepore intermedilysin-CD59

EMDB-0270: 
Stringent response control by a bifunctional RelA enzyme in the presence and absence of the ribosome

PDB-6htq: 
Stringent response control by a bifunctional RelA enzyme in the presence and absence of the ribosome

EMDB-0071: 
GAPDH-CP12-PRK complex

EMDB-3656: 
Structure of the Bacillus subtilis hibernating 100S ribosome reveals the basis for 70S dimerization.

PDB-5njt: 
Structure of the Bacillus subtilis hibernating 100S ribosome reveals the basis for 70S dimerization.

EMDB-3664: 
Structure of the Bacillus subtilis hibernating 100S ribosome reveals the basis for 70S dimerization.
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
