-Search query
-Search result
Showing 1 - 50 of 252 items for (author: rosenthal & p)

EMDB-17309: 
In situ cryoEM structure of Prototype Foamy Virus Env trimer

EMDB-17311: 
In situ cryoEM structure of Prototype Foamy Virus Env dimer of trimers

EMDB-17312: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, icosahedral map

EMDB-17313: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, pentamer localised reconstruction

EMDB-17314: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 1 localised reconstruction

EMDB-17315: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 2 localised reconstruction

EMDB-17316: 
In situ subtomogram average of Prototype Foamy Virus Env trimer

EMDB-17317: 
In situ subtomogram average of Prototype Foamy Virus Env pentamer of trimers

EMDB-17318: 
In situ subtomogram average of Prototype Foamy Virus Env hexamer of trimers

EMDB-17319: 
In situ subtomogram average of the Prototype Foamy Virus capsid, wild-type Gag

EMDB-17320: 
In situ subtomogram average of the Prototype Foamy Virus capsid, p68 Gag

EMDB-17321: 
Cryotomogram of Prototype Foamy Virus particles, wild-type Gag
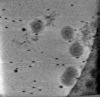
EMDB-17322: 
Cryotomogram of Prototype Foamy Virus particles, p68 Gag

PDB-8ozh: 
In situ cryoEM structure of Prototype Foamy Virus Env trimer

PDB-8ozj: 
In situ cryoEM structure of Prototype Foamy Virus Env dimer of trimers

PDB-8ozk: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, icosahedral map

PDB-8ozl: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, pentamer localised reconstruction

PDB-8ozm: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 1 localised reconstruction

PDB-8ozn: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 2 localised reconstruction

PDB-8ozp: 
In situ subtomogram average of Prototype Foamy Virus Env pentamer of trimers

PDB-8ozq: 
In situ subtomogram average of Prototype Foamy Virus Env hexamer of trimers

EMDB-17611: 
In-situ structure of the heptameric HEF trimers from influenza C viral particles

EMDB-17612: 
In-situ structure of the pentameric HEF trimers from influenza C viral particles

EMDB-18729: 
Cryo-EM structure of tetrameric human SAMHD1 with dApNHpp

EMDB-18730: 
Cryo-EM structure of tetrameric human SAMHD1 State I - Tense

EMDB-18731: 
Cryo-EM structure of tetrameric human SAMHD1 State II - Hemi-relaxed

EMDB-18732: 
Cryo-EM structure of tetrameric human SAMHD1 State III - Relaxed

EMDB-18733: 
Cryo-EM structure of tetrameric human SAMHD1 State IV - Depleted relaxed

EMDB-18734: 
Cryo-EM structure of tetrameric human SAMHD1 State V - Depleted relaxed

PDB-8qxj: 
Cryo-EM structure of tetrameric human SAMHD1 with dApNHpp

PDB-8qxk: 
Cryo-EM structure of tetrameric human SAMHD1 State I - Tense

PDB-8qxl: 
Cryo-EM structure of tetrameric human SAMHD1 State II - Hemi-relaxed

PDB-8qxm: 
Cryo-EM structure of tetrameric human SAMHD1 State III - Relaxed

PDB-8qxn: 
Cryo-EM structure of tetrameric human SAMHD1 State IV - Depleted relaxed

PDB-8qxo: 
Cryo-EM structure of tetrameric human SAMHD1 State V - Depleted relaxed
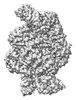
EMDB-18963: 
Structure of the SFTSV L protein in a transcription-priming state without capped RNA [TRANSCRIPTION-PRIMING (in vitro)]
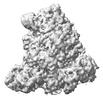
EMDB-18967: 
Structure of the SFTSV L protein in a transcription-priming state with bound capped RNA [TRANSCRIPTION-PRIMING]

EMDB-18969: 
Structure of the SFTSV L protein stalled in a transcription-specific early elongation state with bound capped RNA [TRANSCRIPTION-EARLY-ELONGATION]

PDB-8r6u: 
Structure of the SFTSV L protein in a transcription-priming state without capped RNA [TRANSCRIPTION-PRIMING (in vitro)]

PDB-8r6w: 
Structure of the SFTSV L protein in a transcription-priming state with bound capped RNA [TRANSCRIPTION-PRIMING]

PDB-8r6y: 
Structure of the SFTSV L protein stalled in a transcription-specific early elongation state with bound capped RNA [TRANSCRIPTION-EARLY-ELONGATION]

EMDB-17001: 
In-situ structure of the hexameric HEF trimers from influenza C viral particles
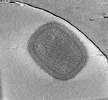
EMDB-18916: 
Cryotomogram of mature Vaccinia virus (WR) virion

EMDB-18917: 
Subtomogram average of the Vaccinia virus (WR) portal complex in mature virions

EMDB-18918: 
Subtomogram average of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions

PDB-8r5i: 
In situ structure of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions by flexible fitting into a cryoET map

EMDB-16258: 
Negative stain EM structure of Toxoplasma gondii glideosome-associated connector (subdomain coil 1-3)

EMDB-16259: 
Negative stain EM structure of Toxoplasma gondii glideosome-associated connector subdomain coil 3

EMDB-16260: 
Negative stain EM structure of Toxoplasma gondii glideosome-associated connector (subdomain coil 1-2)

EMDB-16257: 
Cryo-EM structure of Toxoplasma gondii glideosome-associated connector
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
